Abstract
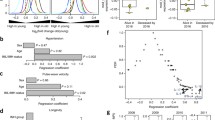
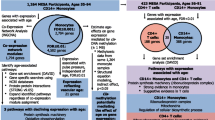
Translating our knowledge of the biological aging from animal models to humans may give rise to novel approaches of targeting multiple aging-related diseases simultaneously and increasing health span. Here, for the first time, we use transcriptomic signatures of monocytes to identify biological aging pathways underlying multiple aging-related diseases in humans. The ordinal logistic regression was used to cross-sectionally investigate transcriptomics of the comorbidity index in 1264 community-based Multi-Ethnic Study of Atherosclerosis (MESA) adults, 47% Caucasian, 32% Hispanic, 21% African American, and 51% female, aged 55–94 years. The comorbidity index was defined as the number of prevalent aging-related diseases including cardiovascular disease, type-2 diabetes, hypertension, cancer, dementia, chronic kidney disease, chronic obstructive pulmonary disease, and hip fracture. We identified 708 gene transcripts associated with the comorbidity index (FDR < 0.05) after adjusting for age, sex, ethnicity, and study site. In a weighted gene co-expression network analysis, as postulated, aging-related declines in apoptosis/autophagy (OR = 1.21 per SD increment, p = 0.0006) and ribosome/mitochondrion (OR = 0.90 per SD increment, p = 0.05) were positively associated with the comorbidity index. After adjusting for multiple comparisons, we identified 10 comorbidity-associated modules (FDR < 0.05), including the module of apoptosis/autophagy. There were three inter-correlated modules of these 10 involved in the complement subcomponent C1q, Fc gamma receptor I, and Fc gamma receptor III of the immune system, respectively. Aging-related upregulation of these three modules was positively associated with the comorbidity index. The odds of comorbidity increased with more of these modules acting together in a dose–response fashion. In conclusion, transcriptomic analysis of human immune cells may identify biomarker panels indicative of comprehensive biological mechanisms, especially immune signaling pathways, contributing to health aging.



Similar content being viewed by others
References
Tinetti ME, Fried TR, Boyd CM. Designing health care for the most common chronic condition–multimorbidity. JAMA. 2012;307(23):2493–4.
Pearson-Stuttard J, Ezzati M, Gregg EW. Multimorbidity-a defining challenge for health systems. Lancet Public Health. 2019;4(12):e599–600.
Harrison DE, Strong R, Allison DB, Ames BN, Astle CM, Atamna H, et al. Acarbose, 17-alpha-estradiol, and nordihydroguaiaretic acid extend mouse lifespan preferentially in males. Aging Cell. 2014;13(2):273–82.
Martin-Montalvo A, Mercken EM, Mitchell SJ, Palacios HH, Mote PL, Scheibye-Knudsen M, et al. Metformin improves healthspan and lifespan in mice. Nat Commun. 2013;4:2192.
Miller RA, Harrison DE, Astle CM, Baur JA, Boyd AR, de Cabo R, et al. Rapamycin, but not resveratrol or simvastatin, extends life span of genetically heterogeneous mice. J Gerontol A Biol Sci Med Sci. 2011;66(2):191–201.
Satoh A, Brace CS, Rensing N, Cliften P, Wozniak DF, Herzog ED, et al. Sirt1 extends life span and delays aging in mice through the regulation of Nk2 homeobox 1 in the DMH and LH. Cell Metab. 2013;18(3):416–30.
Hwangbo DS, Lee HY, Abozaid LS, Min KJ. Mechanisms of lifespan regulation by calorie restriction and intermittent fasting in model organisms. Nutrients. 2020;12(4):1194.
Justice JN, Ferrucci L, Newman AB, Aroda VR, Bahnson JL, Divers J, et al. A framework for selection of blood-based biomarkers for geroscience-guided clinical trials: report from the TAME Biomarkers Workgroup. Geroscience. 2018;40(5–6):419–36.
Karlmark KR, Tacke F, Dunay IR. Monocytes in health and disease - minireview. Eur J Microbiol Immunol (Bp). 2012;2(2):97–102.
Reynolds LM, Ding J, Taylor JR, Lohman K, Soranzo N, de la Fuente A, et al. Transcriptomic profiles of aging in purified human immune cells. BMC Genomics. 2015;16(1):333.
Rubinsztein DC, Marino G, Kroemer G. Autophagy and aging. Cell. 2011;146(5):682–95.
Choi AM, Ryter SW, Levine B. Autophagy in human health and disease. N Engl J Med. 2013;368(7):651–62.
Haas RH. Mitochondrial dysfunction in aging and diseases of aging. Biology (Basel). 2019;8(2):48.
Bild DE, Bluemke DA, Burke GL, Detrano R, Diez Roux AV, Folsom AR, et al. Multi-ethnic study of atherosclerosis: objectives and design. Am J Epidemiol. 2002;156(9):871–81.
Ding J, Reynolds LM, Zeller T, Muller C, Lohman K, Nicklas BJ, et al. Alterations of a cellular cholesterol metabolism network are a molecular feature of cbesity-related type 2 diabetes and cardiovascular disease. Diabetes. 2015;64(10):3464–74.
Espeland MA, Crimmins EM, Grossardt BR, Crandall JP, Gelfond JA, Harris TB, et al. Clinical trials targeting aging and age-related multimorbidity. J Gerontol A Biol Sci Med Sci. 2017;72(3):355–61.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinforma. 2008;9:559.
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4(5):3.
Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proc Natl Acad Sci USA. 2003;100(16):9440–5.
Tower J. Programmed cell death in aging. Ageing Res Rev. 2015;23(Pt A):90–100.
Leidal AM, Levine B, Debnath J. Autophagy and the cell biology of age-related disease. Nat Cell Biol. 2018;20(12):1338–48.
Marino G, Niso-Santano M, Baehrecke EH, Kroemer G. Self-consumption: the interplay of autophagy and apoptosis. Nat Rev Mol Cell Biol. 2014;15(2):81–94.
de Magalhaes JP, Curado J, Church GM. Meta-analysis of age-related gene expression profiles identifies common signatures of aging. Bioinformatics. 2009;25(7):875–81.
Peters MJ, Joehanes R, Pilling LC, Schurmann C, Conneely KN, Powell J, et al. The transcriptional landscape of age in human peripheral blood. Nat Commun. 2015;6:8570.
Ben Mkaddem S, Benhamou M, Monteiro RC. Understanding Fc receptor involvement in inflammatory diseases: from mechanisms to new therapeutic tools. Front Immunol. 2019;10:811.
Son M, Diamond B, Santiago-Schwarz F. Fundamental role of C1q in autoimmunity and inflammation. Immunol Res. 2015;63(1–3):101–6.
Wong KL, Yeap WH, Tai JJ, Ong SM, Dang TM, Wong SC. The three human monocyte subsets: implications for health and disease. Immunol Res. 2012;53(1–3):41–57.
Aiello A, Farzaneh F, Candore G, Caruso C, Davinelli S, Gambino CM, et al. Immunosenescence and its hallmarks: how to oppose aging strategically? A review of potential options for therapeutic intervention. Front Immunol. 2019;10:2247.
Ding J, Kritchevsky SB, Newman AB, Taaffe DR, Nicklas BJ, Visser M, et al. Effects of birth cohort and age on body composition in a sample of community-based elderly. Am J Clin Nutr. 2007;85(2):405–10.
Yi SJ, Kim K. New insights into the role of histone changes in aging. Int J Mol Sci. 2020;21(21):8241.
Zhong H, Yang X, Kaplan LM, Molony C, Schadt EE. Integrating pathway analysis and genetics of gene expression for genome-wide association studies. Am J Hum Genet. 2010;86(4):581–91.
Dimas AS, Deutsch S, Stranger BE, Montgomery SB, Borel C, Attar-Cohen H, et al. Common regulatory variation impacts gene expression in a cell type-dependent manner. Science. 2009;325(5945):1246–50.
Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013;153(6):1194–217.
Lorusso JS, Sviderskiy OA, Labunskyy VM. Emerging omics approaches in aging research. Antioxid Redox Signal. 2018;29(10):985–1002.
Frenk S, Houseley J. Gene expression hallmarks of cellular ageing. Biogerontology. 2018;19(6):547–66.
Acknowledgements
The authors thank the other MEA investigators, the staff, and the participants of the MESA study for their valuable contributions. A full list of participating MESA investigators and institutions can be found at http://www.mesa-nhlbi.org.
Funding
The MESA project is conducted and supported by the National Heart, Lung, and Blood Institute (NHLBI) in collaboration with MESA investigators. Support for MESA is provided by contracts 75N92020D00001, HHSN268201500003I, N01-HC-95159, 75N92020D00005, N01-HC-95160, 75N92020D00002, N01-HC-95161, 75N92020D00003, N01-HC-95162, 75N92020D00006, N01-HC-95163, 75N92020D00004, N01-HC-95164, 75N92020D00007, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169, UL1-TR-000040, UL1-TR-001079, UL1-TR-001420, UL1TR001881, DK063491, and R01HL105756. The MESA Transcriptomics Studies were funded by NIH grants R01HL101250, R01HL119962, R01DK101921, R01HL135009, RF1AG054474, and U01AG060897. This work was partially supported by P30 AG21332.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Ding, J., Lohman, K., Molina, A. et al. The association between aging-related monocyte transcriptional networks and comorbidity burden: the Multi-Ethnic Study of Atherosclerosis (MESA). GeroScience 45, 197–207 (2023). https://doi.org/10.1007/s11357-022-00608-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-022-00608-1