Abstract
The genes ECERIFERUM1 (CER1) and ECERIFERUM3 (CER3) encode the biosynthesis of alkane waxes, a key component of the plant cuticle. To study the evolution of CER1 and CER3 in a highly diverse group of eucalypts, we performed a genome-wide survey using recently released genome assemblies of 28 Myrtaceae species, with 22 species from the main eucalypt lineage and 6 non-eucalypt Myrtaceae tree species. We manually annotated 250 genes and pseudogenes, identifying a near-ubiquitous single copy of CER3 and 2 to 10 CER1 gene copies per Myrtaceae species. Phylogenetic analysis suggested that copy number variation in eucalypts is due to multiple tandem duplication events, both ancient (shared by all Myrtaceae species studied) and relatively recent (present only in eucalypts). Inter-chromosomal translocations were discovered for both CER1 and CER3, along with recurrent loss of often the same CER1 introns in the WAX2 domain, the domain that is essential for wax production. Despite the varied environments occupied by the eucalypt species in this study, we did not find statistically significant associations between intra-genic structural changes or CER1 copy number and aspects of the environment they occupy (including aridity). The challenge is now to explain the species-specific evolutionary histories that contributed to the observed variation in CER1 and the extent to which it may contribute to the adaptability of eucalypts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants need to retain water to survive in dehydrating habitats (Yeats and Rose 2013). They achieve this through the plant cuticle, a hydrophobic membrane that covers most of their aerial organs (Liu et al. 2021). While the cuticle’s primary function is to restrict non-stomatal water loss and control the exchange of solutes and gases between plants and the atmosphere, it also protects against environmental stressors, such as high temperature, UV radiation, microorganisms and insects (Domínguez et al. 2011). As such, it is one of the most important adaptations that allowed plants to transition from aquatic to terrestrial environments (Bhanot et al. 2021). The cuticle is chemically composed of an insoluble polymer called cutin and a wax mixture that includes hydrophobic lipids consisting primarily of very long-chain fatty acids and their derivatives, such as aldehydes, alkanes, primary and secondary alcohols, ketones and wax esters (Lee and Suh 2015; Domínguez et al. 2017). The quantity and chemical composition of cuticular waxes varies across species, organs, tissues and time (Nawrath 2006) and can change in response to biotic or abiotic stress (Shepherd and Wynne Griffiths 2006; Lewandowska et al. 2020).
The biosynthesis of cuticular waxes in flowering plants is a complex process controlled by a large number of genes (Xue et al. 2017; Ai et al. 2022). Among these, the ECERIFERUM (CER) gene family plays an important role in the biosynthesis of very long-chain fatty acids, which are a main component of cuticular waxes (Hannoufa et al. 1993; Jenks et al. 1995). The most studied member of this family is the gene ECERIFERUM1 (CER1), which encodes the biosynthesis of alkanes through decarboxylation of fatty acid metabolites and is implicated in drought tolerance (Aarts et al. 1995; Bourdenx et al. 2011; He et al. 2022). Alkanes are usually omnipresent in the cuticle of most species and organs, where they frequently accumulate to high concentrations (Samuels et al. 2008), increasing drought tolerance by reducing the permeability of the plant surface (Li et al. 2019). Transgenic experiments also showed that another member of the CER gene family, ECERIFERUM3 (CER3), interacts with CER1 to produce alkanes in Arabidopsis (Bernard et al. 2012). CER1 and CER3 share a common ancestry and their proteins are structurally similar (Wang et al. 2019), having evolved from the fusion of ERG3/FAH and WAX2 domains, which are their N-terminus and C-terminus, respectively (Chaudhary et al. 2021). The WAX2 domain is crucial for wax synthesis as its mutations can drastically affect the total amount of wax (Rowland et al. 2007). Schematic representations of their exon–intron structure show 10 exons for CER1, while CER3 presents 11 exons in Arabidopsis (Rowland et al. 2007; Sakuradani et al. 2013). There are also two CER1 homologues identified in Arabidopsis. One of these, CER1-LIKE1, is known to interact with CER3 to produce alkanes of different chain lengths compared to those produced by CER1 and is expressed in different organs and tissues (Pascal et al. 2018).
Phylogenetic analysis with diverse species of Archaeplastida (i.e., land plants, green and red algae and glaucophytes) has shown that CER1 and CER3 copy numbers tend to increase as plants evolved. Thus, ancient terrestrial lineages such as bryophytes, lycophytes and ferns present a low number of CER1 and CER3 genes, while seed plants usually have more copies. Similarly, gymnosperms often show lower number of copies of these genes than angiosperms, which can themselves vary within genera (Chaudhary et al. 2021). For instance, a phylogenetic study revealed that Quercus mongolica has a much-expanded number of CER1 copies compared with other species of this genus, with both tandem and dispersed duplicates detected (Ai et al. 2022). The authors suggested that this expansion could contribute to the adaptability of Quercus species to drought. Increasing the copy number has been suggested as an adaptation to environmental variation in several polyploid species, such as tobacco or wheat (Limin and Fowler 1989; Deng et al. 2012). In addition, an effect of copy number on function has also been reported with other genes, such as GhDREB1B in cotton, with increasing chilling tolerance with higher copy number (Wang et al. 2021). Less studied has been the effect of gene structural variation, such as intron loss, on function and environmental adaptation. One study showed that intron-poor members of the CIPK gene family are more highly expressed in response to drought stress than the intron-rich genes in soybeans (Zhu et al. 2016). No study has examined the relationship between variation in CER gene structure or copy number with environmental stressors such as drought.
Eucalypts are a group of trees and shrubs from the Myrtaceae family encompassed by the genera Eucalyptus L’Her. (~ 750 species), Corymbia K.D. Hill and L.A.S. Johnson (~ 100 species) and Angophora Cav. (10 species) that are naturally distributed in Australia and Malesia (Nicolle 2022a). Like the majority of Myrtaceae species, eucalypts are diploid with 2n = 22 (Grattapaglia et al. 2012). Eucalypts comprise a mixture of diverse and depauperate lineages, which are adapted to nearly every Australian environment (Thornhill et al. 2019; Slee et al. 2020). The adaptability and high growth rate of some species make eucalypts the most economically important hardwood trees worldwide for the production of timber, fibre and energy (Turnbull 1999). A key trait that is taxonomically and ecologically important in eucalypt species is the presence of glaucous waxy leaves (Barber 1955; Hallam and Chambers 1970). These waxes not only protect eucalypts from water loss (Hoffmann et al. 2013) but also other environmental stressors such as frost (Keller et al. 2013), high radiation (Close et al. 2007) as well as insects (Edwards 1982; Jones et al. 2002) and pathogens (Santos et al. 2019). The chemical composition and amount of cuticular waxes in eucalypts are influenced by both environmental and genetic factors (Koch et al. 2006; Gosney et al. 2016) and can be extremely variable between eucalypt species, with alkanes ranging from 0.6 to 74.3% of the total wax load (Li et al. 1997). Previous research showed quantitative trait loci (QTL) for wax yield (Gosney et al. 2016) and drought damage (Gosney et al. 2016; Ammitzboll et al. 2020) co-located with CER candidate genes in E. globulus. In E. grandis, a single copy of the gene CER3 was identified, while several copies of CER1 were detected (Chaudhary et al. 2021). Although their architecture is currently unstudied, gene families in eucalypts with multiple copies are often arranged in tandem duplicate arrays (Myburg et al. 2014; Li et al. 2015a; Healey et al. 2021), which may be the case here.
In this study, we performed a genome-wide survey for CER1 and CER3 genes across the main eucalypt lineages and compared them to those of other tree species of the Myrtaceae family, as well as Arabidopsis. We characterized their position in the genome, copy number, exon–intron structure and their phylogenetic relationships to better understand their evolution in eucalypts. We aim to determine (i) whether eucalypt CER gene duplication is ubiquitous across the genera, (ii) the extent to which CER genes exhibit structural variation, and (iii) if copy number or structural variation is associated with the species home-range environmental variation.
Materials and methods
Acquisition of CER sequences
To have a good representation of the taxonomic diversity of eucalypts, we used genome assemblies from 22 eucalypt species spanning the genera Angophora (1 species), Corymbia (2 species) and Eucalyptus (19 species), which cover all the subgenera and sections of the group that have a genome currently available. The genomes of another 6 tree species of the Myrtaceae family were included as outgroup (Melaleuca alternifolia) or sister taxa (Psidium guajava, Rhodamnia argentea, Metrosideros polymorpha, Syzygium aromaticum, Leptospermum scoparium) for comparison with the eucalypts (Thornhill et al. 2015), giving a total of 28 tree species studied (Supplementary Table S1). All the genomes were obtained from public repositories (see Myburg et al. 2014; Izuno et al. 2016; Thrimawithana et al. 2019; Wang et al. 2020; Ahrens et al. 2021; Healey et al. 2021; Voelker et al. 2021; Ferguson et al. 2023) except for E. globulus, which was obtained from Agriculture Victoria, Australia and Leptospermum scoparium which was obtained from the Aotearoa Genomic Data Repository (https://www.genomics-aotearoa.org.nz/data). The E. grandis and C. citriodora genomes were obtained from Phytozome 13 (https://phytozome-next.jgi.doe.gov), whereas the remaining genomes were obtained from GenBank (https://www.ncbi.nlm.nih.gov/genbank/). Most of these Genbank genomes were de novo assembled into very large contigs and scaffolded into chromosomes using the E. grandis assembly as a reference (Ferguson et al. 2023). Melaleuca alternifolia and E. pauciflora were not assembled into chromosomes, while E. grandis, E. globulus, C. citriodora, Psidium guajava, Rhodamnia argentea, Metrosideros polymorpha and Syzygium aromaticum were assembled to chromosome level de novo. To check that the chromosome assignment and orientation of Psidium guajava, Rhodamnia argentea, Metrosideros polymorpha and Syzygium aromaticum matched the most syntenic ones in eucalypts, the genomes of these species were aligned against the E. grandis genome using minimap2 (Li 2018) and their chromosomes were oriented and renamed if needed (Supplementary Figure S1). The chromosomes of all other species were numbered and oriented following E. grandis as is the convention in eucalypts.
The peptide sequences of CER1 (AT1G02205) and CER3 (AT5G57800) were obtained from the Arabidopsis genome (TAIR) via keyword search in Phytozome 13. We also obtained the peptide sequence of the two CER1 homologues in Arabidopsis (AT1G02190 and AT2G37700), which we named CER1a and CER1b, respectively, for inclusion in later analyses. The sequences of CER1 and CER3 were used in a tBLASTn search (Altschul et al. 1990) of all selected Myrtaceae genomes to find genomic regions that likely contain CER genes (e-value < 1e−03). The synteny of the genomic regions containing CER genes was examined through pairwise whole genome alignment using MUMmer (Marçais et al. 2018) using the same parameters as Ferguson et al. (2023), namely the tool nucmer (--maxmatch -l 40 -b 500 -c 200), and summarised using syri (Goel et al. 2019). Some alignments were able to be sourced from Ferguson et al. (2023).
Identification of genes, pseudogenes and exon–intron structure
To determine gene coordinates and exon–intron borders of putative CER genes, we first obtained from GenBank or Phytozome the nucleotide sequences of the genomic region (± 1 kb from hit) identified above and then used GeneWise (Birney et al. 2004) through the online platform EMBL-EBI (https://www.ebi.ac.uk/Tools/psa/genewise) (Madeira et al. 2022) using the nucleotide and peptide sequences of the genes CER1 and CER3 from Arabidopsis. Putative genes with reading frame shifts or insertions/deletions leading to premature stop codons were classified as pseudogenes. The chromosome number and position within a chromosome of each CER1 and CER3 gene and pseudogene were identified in each Myrtaceae species. The relative disposition of CER genes and pseudogenes with other non-CER transcripts was checked in the genomes of E. grandis, E. globulus and C. citriodora, which are the only eucalypt species that currently have annotations. To do that, we used the tools JBrowse (https://phytozome-next.jgi.doe.gov/jbrowse/index.html) for E. grandis and C. citriodora and the Integrative Genomics Viewer (https://igv.org/) for E. globulus.
Conserved domains were identified on each gene using the online tool Batch CD-Search (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi) at NCBI’s Conserved Domain Database (Marchler-Bauer et al. 2010). The Gene Structure Display Server (GSDS v2, http://gsds.gao-lab.org/index.php) was used to visualise the exon–intron structure and position of conserved domains within genes (Hu et al. 2014).
Phylogenetic analysis
Multiple alignments of amino acid sequences of all Myrtaceae CER1 and CER3 genes as well as the Arabidopsis CER1, CER3, CER1a and CER1b were performed in MUSCLE (Edgar 2004). To mitigate potential errors, a visual inspection of the multiple sequence alignment was performed, assessing place gaps and potentially poorly aligned positions, but no trimming was necessary as we could not find any inaccuracy. Using these aligned peptide sequences, a phylogenetic tree was generated in IQ-TREE v1.6.12 with 1000 ultrafast bootstrap replicates (Nguyen et al. 2015). The software FigTree v1.4.4 was used to visualise the phylogenetic tree (http://tree.bio.ed.ac.uk/software/Figtree).
Association of copy number and exon–intron structure with environmental variation
Values of ten environmental variables were obtained from the Atlas of Living Australia (https://www.ala.org.au/) for the natural distribution of 21 eucalypt species in Australia, with the exception of E. camaldulensis which was excluded from the analysis because of its extremely wide distribution in Australia (Slee et al. 2020). The environmental variables included elevation (ELE) and nine climate variables, which were mean annual temperature (MAT), mean temperature of the coldest quarter (MTCQ, 3-month period), mean temperature of the warmest quarter (MTWQ), a thermic index of continentality (TIC) calculated as the difference between MTWQ and MTCQ (Tuhkanen 1980), mean precipitation of the wettest quarter (MPWQ), mean annual precipitation (MAP), annual mean radiation (RAD) and two moisture indices used to describe overall aridity. These indices were the annual heat–moisture index (AHM), which was calculated using the equation AHM = (10 + MAT)/(1000−1 MAP), and a modified summer heat–moisture index (SHM), that was calculated using the equation SHM = MTWQ/(1000−1 MAP) (Wang et al. 2006), replacing the mean warmest month temperature of the original equation by MTWQ. Records outside a species’ natural distribution were identified using a modified z-score outlier test based on the median absolute deviation of the mean annual temperature, mean annual precipitation and elevation (Iglewicz and Hoaglin 1993). Records with a z-score > 3.5 were checked manually against the known natural species distribution and deleted when appropriate (Jordan et al. 2016). For each of the 21 species, the average of each environmental variable was then calculated from the retained distributional records. To explore the association between the copy number of CER genes and the adaptation of eucalypts to environmental variation, Pearson’s correlations were calculated between the number of functional genes per eucalypt species and the average of each environmental variable for the species distribution. To explore the effect of the gene structure on environmental adaptation in eucalypts, we split the eucalypt species into two contrasting groups according to the results of the exon–intron structure analysis. We then used Welch’s t-test to compare the means of the two groups for each environmental variable.
Results
Copy number variation across all Myrtaceae
The genome-wide search on the 28 Myrtaceae tree species revealed 250 genomic regions containing sequences with high similarity to Arabidopsis CER1 or CER3. 162 sequences were annotated as genes, from which 135 were CER1 and 27 were CER3 genes. A total of 88 sequences were identified as pseudogenes, of which 87 were CER1 pseudogenes and only one was a CER3 pseudogene (Supplementary Table S2). While there was only one copy of the gene CER3 per species in almost all species (the exception was Leptospermum scoparium with no CER3 detected), multiple copies of CER1 were found in all the Myrtaceae tree species (Table 1). Notably, the copy number of CER1 genes ranged from 2 to 10 in eucalypts, whereas it varied from 1 to 4 for the other Myrtaceae tree species. Similarly, the number of CER1 pseudogenes was also greater in eucalypts than the other Myrtaceae species. Within eucalypts, the copy number of CER1 ranged from 3 to 10 for the genus Eucalyptus and from 2 to 7 for its sister genus Corymbia and related Angophora, with a similar pattern detected in the pseudogenes. Within the genus Eucalyptus, multiple species were represented from the two most important subgenera—Symphyomyrtus and Eucalyptus—and considerable variation in the number of CER1 copies was detected within these subgenera. The Symphyomyrtus species ranged from 3 to 10 copies of CER1 and species from the subgenus Eucalyptus had between 5 and 7 (Table 1).
Conserved architecture of CER genes in Myrtaceae
All the Myrtaceae species with their genomes assembled at the chromosome level showed the majority of their CER1 genes and pseudogenes on chromosome 4 (Table 1). CER1 genes (and occasionally pseudogenes) were also present on chromosome 8 in most species of the genus Eucalyptus, except for six of the ten species representing subgenus Symphyomyrtus. CER1 genes were also absent from chromosome 8 in Angophora and Corymbia and the other Myrtaceae species, suggesting a recent evolutionary origin within only Eucalyptus for CER1 on this chromosome. Multiple copies of CER1 were present for C. citriodora on chromosome 9, but absent in Corymbia maculata (a eucalypt species not included in our dataset since its subgenus and section were already covered by the closely related C. citriodora) when a tBLASTn search of arabidopsis CER1 was performed (data not shown). Eucalyptus curtisii, E. melliodora and E. marginata, which belong to different Eucalyptus subgenera, showed CER1 genes and/or pseudogenes on chromosome 11, which appear to be independent translocations based on the lack of synteny between these loci (Fig. 1). The CER1 on chromosome 11 of E. curtisii and E. marginata are related to copies on chromosome 4, while the CER1 of E. melliodora on chromosome 11 is more related to CER1 on chromosome 8. Only one translocation of CER1 on chromosome 6 was registered for E. globulus. Most of the species showed CER3 genes and pseudogenes on chromosome 3 (Table 1), but translocations were also observed from this chromosome to chromosomes 1 and 8 for E. guilfoylei and E. cladocalyx, respectively. The chromosomal position of some CER genes or pseudogenes was undetermined (Un, Table 1) because some genomes were not assembled to chromosome level, or in other cases, CER genes/pseudogenes were identified within the unassembled component of data in Genbank.
Pairwise alignment of chromosomes 4, 8 and 11 for E. curtisii, E. marginata and E. melliodora. Black bars indicate the locations of CER1 genes/pseudogenes on the only three species with copies on chromosome 11. Grey lines indicate matching syntenic regions between the three eucalypt species. Green lines indicate translocations. Note the synteny of the region surrounding the CER1 loci on chromosomes 4 and 8 but the lack of synteny for the region surrounding the CER1 loci on chromosome 11. None of the CER1 genes on chromosome 11 share a common ancestry
The relative position of CER1 genes and pseudogenes within chromosome 4 shows the copies to be tandem repeats in a region that is syntenic across all studied species, spanning a region up to 1.6 Mbp (~ 4% of chromosome 4), except for Eucalyptus erythrocorys and Psidium guajava, which spanned regions of 4.6 and 7.4 Mbp, respectively (Supplementary Table S2; Supplementary Figure S2). While this region is in a more-or-less central position on chromosome 4 in most species, in E. regnans and E. tenuipes, in particular, it is shifted towards the chromosome end. A closer look at this region in E. grandis and E. globulus (the only Eucalyptus with annotations to date) reveals that CER1 genes and pseudogenes occur as interspersed repeats, since E. grandis has 40 non-CER1 transcripts dispersed among the eight CER1 genes and pseudogenes, while E. globulus has 46 non-CER1 transcripts dispersed among the 12 CER1 genes and pseudogenes (data not shown). In the case of C. citriodora (the only other eucalypt with an available annotation), there is only one CER1 gene copy and no pseudogene on chromosome 4. The tandem arrangement of CER1 sequences was also observed in chromosome 8 for some species of Eucalyptus, spanning a region up to 0.14 Mbp (Supplementary Table S2; Supplementary Figure S3), again in a syntenic region.
Gene structural variation across multiple Myrtaceae lineages
Most of the orthologues of CER1 in arabidopsis showed 10 exons and 9 introns in Myrtaceae. The ERG3 domain occupied exons 3 and 4 completely and exon 5 partially; while the WAX2 domain occupied exon 7 partially and exons 8, 9 and 10 completely, 10 being the last exon next to the 3′ end position. In some cases, a partial occupation of exon 3 by the ERG3 domain was observed (Fig. 2). We detected the loss of the 7th or 8th intron in the WAX2 domain for 24 CER1 genes from 15 species that included Metrosideros polymorpha, C. citriodora and Eucalyptus from diverse subgenera (Fig. 2, Supplementary Figure S4). Gene size of CER1 ranged from 5.9 to 18.9 kbp, with most of the genes around 7 kbp. Likewise, the exon–intron structure of CER3 in the Myrtaceae species followed the exon–intron structure of arabidopsis CER3, with 11 exons and 10 introns (Supplementary Figure S5). Corymbia calophylla was the only species exhibiting structural variation in CER3, with the loss of the first exon and intron. The ERG3 domain of CER3 occupied the same exons and showed the same variations described above for CER1, while the WAX2 domain occupied exon 8 partially and exons 9, 10 and 11 completely next to the 3′ end position. CER3 gene size was relatively uniform among the different Myrtaceae species, averaging 4.5 kbp (Supplementary Figure S5).
Range of structural variation of CER1. Genes are shown in the following order: Arabidopsis thaliana CER1; typical E. globulus CER1 with ERG3 domain occupying the exons 3 and 4 completely and the exon 5 partially; typical E. viminalis CER1 with ERG3 domain occupying the exon 4 completely and the exons 3 and 5 partially; typical C. calophylla CER1 with the 7th intron lost in the WAX2 domain; typical E. brandiana CER1 with the 8th intron lost in the WAX2 domain; and atypically long E. tenuipes CER1. Eucalypt sequences were named using species followed by chromosome number, number of exons (e9 or e10) and copy number on the chromosome if there were more than one
Phylogenetic relationships between CER genes
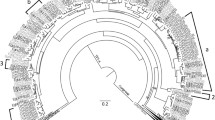
The phylogenetic analysis showed three main clades of CER genes for the Myrtaceae species (Fig. 3). Clades 1 and 2 were phylogenetically related to Arabidopsis CER1, whereas Clade 3 was related to Arabidopsis CER3. In addition, the Arabidopsis genes CER1a and CER1b (homologues of Arabidopsis CER1), were not closely related to Myrtaceae CER1 or CER3, thus the duplication of CER1 observed in Myrtaceae is independent of that in Arabidopsis. The duplication of CER1 into Clades 1 and 2 is relatively old as both of these clades include almost all eucalypt species and many of the other Myrtaceae species (Fig. 4a and b). Clade 2 includes 5 of the 6 other Myrtaceae species and 21 of the 22 eucalypts while Clade 1 includes all eucalypt species but only two of the other Myrtaceae species and, importantly, does not include Melaleuca alternifolia which is from a more ancient divergence within Myrtaceae compared to eucalypts as shown in the phylogeny of CER3 (Fig. 4c). Therefore, the duplication of CER1 into Clades 1 and 2 most likely occurred within Myrtaceae. The internal structure of Clades 1 to 3 generally reflected the species taxonomy (Fig. 4a, b and c) and phylogenies such as the one of Thornhill et al. (2015).
Phylogeny of CER1 and CER3 genes for eucalypts, other Myrtaceae tree species and arabidopsis. The scale represents amino acid substitution per site. Bootstrap values are displayed on branches. Arabidopsis sequences are shown using chromosome number and eceriferum gene (CER1, homologues CER1a and CER1b, and CER3). CER1 is shown in Clades 1 and 2 in Myrtaceae including eucalypts, while CER3 is only present in Clade 3
Details of the phylogeny of CER1 and CER3 for eucalypts and other Myrtaceae tree species based on peptide sequences. a Clade 1 and b Clade 2 of the phylogeny which were comprised only of CER1 genes and c Clade 3 of the phylogeny which comprised only CER3 genes. The legend differentiates non-eucalypt Myrtaceae, subgenus within eucalypts or genus when subgenus is inexistent for the taxon. Sequences were named using species or genus followed by chromosome number, number of exons (e9 or e10) and copy number on the chromosome if there were more than one. I, II and III are subclades that group sequences with intron loss. IV is a subclade that groups sequences from chromosome 8. Asterisk (*) indicates identical sequences for the same species. Asterisks (**) indicates identical sequences for different species. The scale represents amino acid substitution per site
The CER1 phylogeny grouped eucalypt sequences with intron loss into three small subclades (subclades I, II and III; Fig. 4a), reflecting the subgeneric structure. These three subclades only included Eucalyptus species. Note that the phylogeny was not affected by the intron–exon structure as it was based on the peptide sequences. The CER1 genes present on chromosome 8 were grouped into one single subclade (subclade IV; Fig. 4a) and included all 8 subgenera of Eucalyptus, but not the other two genera of eucalypts (Corymbia and Angophora). Thus, this translocation to chromosome 8 appears to be specific to the genus Eucalyptus. Exact copies of CER1 were found within the same species for E. globulus and E. brandiana (Fig. 4a and b), suggesting they are the result of recent tandem duplications. Exact copies were also found for the taxonomically close species E. globulus and E. viminalis, suggesting the presence of this gene in a common ancestor (Fig. 4a). As the phylogeny of CER3 includes only one copy per species, the position of sequences more clearly followed the species taxonomy (Fig. 4c).
Copy number and structural variation association with species home-environment
Correlations between the number of copies of CER1 genes per species and the average of each environmental variable for the eucalypt home range were not statistically significant (p > 0.05, Table 2), suggesting no association between the number of CER1 copies and environmental variation. Based on the exon–intron structure of the CER1 genes, eucalypt species were classified into two groups as follows: (i) species that present a complete gene structure (i.e., no intron loss, n = 8), which included A. floribunda, C. citriodora, E. erythrocorys, E. guilfoylei, E. virginea, E. leucophloia, E. grandis and E. melliodora; and (ii) species in which at least one CER1 gene showed intron loss (n = 13), which included C. calophylla, E. curtisii, E. tenuipes, E. cloeziana, E. marginata, E. pauciflora, E. regnans, E. microcorys, E. pumila, E. globulus, E. viminalis, E. cladocalyx and E. brandiana. No statistically significant differences were found for the environmental variables when the two groups of eucalypts were compared with a Welch’s t-test (p > 0.05, Table 2), suggesting that no association between gene structure and environmental variation exists. Since no major changes in copy number or gene structure were noticed in CER3, no such tests were performed.
Discussion
Most of the basic knowledge on eceriferum genes comes from research on the model plant species Arabidopsis (Koornneef et al. 1989; Aarts et al. 1995; Jenks et al. 1995; Rowland et al. 2007), but the development of high-throughput sequencing and the availability of new complete genomes have recently allowed the study of eceriferum homologues at the genome-wide level in other plant species. These species include non-woody species such as sunflower, tomato, wheat, passion fruit (Ahmad et al. 2021; He et al. 2022; Rizwan et al. 2022; Wu et al. 2022), as well as woody species such as jujube, oak, Chinese chestnut (Li et al. 2021; Ai et al. 2022; Zhao et al. 2022) and in the present case, the eucalypts, sister taxa and the outgroup. By manually annotating the CER1 and CER3 genes across the 28 Myrtaceae genomes, we showed that CER3 copy number was conserved across the Myrtaceae, while CER1 was highly variable, with multiple lineage-specific tandem duplications in eucalypts, along with a translocation event conserved in many subgenera. In addition, we identified variation in the exon–intron structure of CER1, detecting the loss of intron 7 or 8 in different lineages of Myrtaceae. We did not find evidence to link variation in gene structure and copy number with the capacity of eucalypts to adapt to different environments. The information presented in this study highlights the variability of the CER1 genes, presumably because of the instability induced by tandem arrays, which is in contrast to the stability of the single copy CER3 in eucalypts.
Eucalypts had a higher number of copies of CER1 genes than the Myrtaceae outgroup and sister species, and especially higher than arabidopsis. Most often these copies were grouped in the phylogeny indicating lineage-specific duplication events. These duplications were most common for a localised region of chromosome 4 which harboured between 1 and 18 genes/pseudogenes depending on the species, suggesting that tandem duplication was the mechanism responsible for these patterns. Similar gene expansion has been observed in other gene families of E. grandis (Li et al. 2015b), E. globulus (Külheim et al. 2015) and C. citriodora (Butler et al. 2018) and is a feature of eucalypts which has a high proportion of tandemly duplicated genes (Myburg et al. 2014; Healey et al. 2021). Expansion of the CER1 gene family specifically has also been observed in oaks, with Ai et al. (2022) suggesting this may confer greater adaptability to drought, potentially through a dosage effect from increased gene product (Kondrashov 2012; Kuzmin et al. 2022). This seemed to be a plausible explanation for the highly variable copy number observed across the eucalypts, especially considering the diverse environments and stresses these eucalypts are exposed. However, we found no obvious association between CER1 copy number and aridity (measured as moisture indexes) or any other environmental variable measured for the species’ current distributions. This lack of association is not entirely unexpected, given the disparate CER1 copy number among species occupying similar environmental niches in our study. For instance, E. globulus and E. viminalis, which share a large area of their distribution and are phylogenetically close, showed dissimilar copy numbers of the CER1 gene in our study yet when they co-occur show similar susceptibility to drought (Kirkpatrick and Marks 1985). A more detailed investigation of this issue is needed, and an investigation of CER1 expression levels will be required to determine if dosage amplification of CER1 contributes to the environmental adaptability of eucalypts.
Chromosome 3 appears to be the ancestral position for CER3 in Myrtaceae, given the conservation of the position of this gene across the phylogeny. Similarly, CER1 was primarily located in a syntenic region on chromosome 4 in the Myrtaceae species studied, which implies that this is the ancestral position for Myrtaceae. The tandem duplication of CER1 on chromosome 4 which gave rise to two large clades of CER1 (Clades 1 and 2) is likely to be Myrtaceae specific since Clade 1 did not contain sequences from the outgroup Melaleuca alternifolia, which is the most basal Myrtaceae divergence sampled in our study (Thornhill et al. 2015). A large proportion of the Eucalyptus species examined also had CER1 genes or pseudogenes on chromosome 8, which was not the case in the related Angophora and Corymbia or other Myrtaceae, suggesting a more recent duplication and translocation event was responsible for this novel locus. There were also species-specific duplications and translocations. For CER1, this included the positions on chromosome 11 observed in E. curtisii, E. marginata, E. melliodora and the position on chromosome 9 in C. citriodora. For CER3, this included the position on chromosome 8 in E. cladocalyx and the position on chromosome 1 in E. guilfoylei. These species-specific inter-chromosomal translocations may be the result of specific evolutionary events for these species, although assembly errors cannot be discounted (Wang et al. 2020). Indeed, the translocation of CER1 to chromosome 9 in C. citriodora may be an assembly error as there are no copies present on this chromosome in the related C. maculata (data not shown).
Intron loss was observed in only one of the two clades of CER1 (Clade 1), where it occurred in multiple lineages including the genus Corymbia, diverse subgenera of Eucalyptus as well as Metrosideros polymorpha, a genus thought to have diverged from the eucalypts around 65–70 million years ago (Thornhill et al. 2015). Our study suggests introns of the CER1 genes were independently lost several times in the evolution of Myrtaceae as the cases of intron loss were dispersed in different subclades within Clade 1 (Fig. 4a). This pattern has been observed previously, with intron loss and gain reported in different members of the eceriferum gene family for sunflower (Ahmad et al. 2021). In addition, Rizwan et al. (2022) suggested that the gene family has gone through several rounds of intron loss and gain during its evolution. However, none of this research reported intron loss in eceriferum genes as a frequent event as we do. Similar recurrent intron loss for other genes has been found in other evolutionary lineages in both plants (Wang et al. 2014; Milia et al. 2015) and animals (Cho et al. 2004; Coulombe-Huntington and Majewski 2007), but this appears to be the first observation of this phenomenon in eucalypts. Research on animals, plants and malaria has shown that intron loss was more frequent in highly duplicated genes (Castillo-Davis et al. 2004; Lin et al. 2006; Roy and Penny 2007), which accords with our observed differences between CER1 and CER3 in eucalypts. Other research in yeasts, mice, and arabidopsis showed that stress genes with fast-changing expression levels had significantly lower intron densities possibly to avoid delays in transcript production and energetic costs associated with increased transcript length (Jeffares et al. 2008). This hypothesis suggests that selection may favour intron loss during evolution in such cases. This hypothesis may be applicable to the observed intron loss in CER1, which is known to increase its expression level quickly in response to drought stress in Arabidopsis (Bourdenx et al. 2011) and several other plant species (He et al. 2022; Wu et al. 2022; Gao et al. 2023). However, we did not find that intron losses in CER1 were linked to the species’ home-environmental variation, including measurements of aridity. Species-specific evolutionary pathways could have contributed to the loss of introns at any point in their evolutionary history, which could have been independent of the environment, supporting Penny et al. (2009), who consider the attribute of intron loss to a particular environmental condition to be speculative.
Intron loss was restricted to introns 7 and 8 of CER1, which may be explained by the intron loss mechanisms or by the features of these introns. Intron loss can be due to stochastic processes such as direct genomic deletion or nonhomologous end joining during repair of DNA double-strand breaks (Cohen et al. 2011; Fawcett et al. 2011). However, a third mechanism, the reverse transcription model (Roy and Gilbert 2005), may offer an explanation, particularly as only specific introns are lost in our study (i.e., always introns 7 or 8). According to this mechanism, introns are lost through gene conversion by a retrotransposed copy of a spliced transcript of the gene often spanning multiple intron positions. Given that the process of reverse transcription initiates at the 3′ end of genes and the enzyme reverse transcriptase frequently disassociates from the template prematurely, introns closer to the 3′ end, like introns 7 and 8 in CER1, are more prone to be lost. In our study, the adjacent introns 7 and 8 were never lost simultaneously. The absence of this concurrent loss in our study is an argument against the reverse transcriptase model, which is reported to favour the concurrent loss of neighbouring introns (William Roy and Gilbert 2006; Ma et al. 2015). Specific features of these introns may also help explain their loss. For example, small introns, like intron 8, are more likely to be lost than larger introns (Coulombe-Huntington and Majewski 2007; Loh et al. 2007). Although the reason for this size effect is still not completely understood, one potential explanation is the lower likelihood of regulatory modules occurring in short introns thus their loss is less deleterious (Wang et al. 2014). However, it is notable that introns 7 and 8 interrupt the conserved functional WAX2 domain of CER1. Functional domains uninterrupted by introns can provide a selective advantage due to a greater capacity for exon shuffling as hypothesised by Liu and Grigoriev (2004). However, changes in exon position were not observed in our study arguing against this hypothesis. Intron loss remains a subject of great interest for molecular biologists because of its importance in the evolution of life (Rodríguez-Trelles et al. 2006), but at this stage remains poorly understood (Rogozin et al. 2012; Milia et al. 2015).
In conclusion, our study of CER1 and CER3 genes across the main eucalypt lineages and other Myrtaceae species showed that gene structure and copy number varied markedly among species for CER1, but was highly conserved for CER3. Several evolutionary events were specific to eucalypts, such as a high level of tandem duplications or specific inter-chromosomal translocations in Eucalyptus. Although no association was found between CER1 gene structure or copy number and the environment of origin of the studied species, more research is needed to investigate the link between the observed variation in the eceriferum genes and its possible contribution to adaptability.
Data availability
Genomes used in this study are available at GenBank (https://www.ncbi.nlm.nih.gov/genbank/), except for Eucalyptus globulus and Leptospermum scoparium that were provided by Agriculture Victoria and Genomics Aotearoa, respectively.
References
Aarts MG, Keijzer CJ, Stiekema WJ, Pereira A (1995) Molecular characterization of the CER1 gene of arabidopsis involved in epicuticular wax biosynthesis and pollen fertility. Plant Cell 7:2115–2127. https://doi.org/10.1105/tpc.7.12.2115
Ahmad HM, Wang X, Fiaz S, Mahmood Ur R, Azhar Nadeem M, Aslam Khan S, Ahmar S, Azeem F, Shaheen T, Mora-Poblete F (2021) Comprehensive genomics and expression analysis of eceriferum (CER) genes in sunflower (Helianthus annuus). Saudi J Biol Sci 28:6884–6896. https://doi.org/10.1016/j.sjbs.2021.07.077
Ahrens CW, Murray K, Mazanec RA, Ferguson S, Bragg J, Jones A, Tissue DT, Byrne M, Borevitz JO, Rymer PD (2021) Genomic constraints to drought adaptation. bioRxiv 11:3044. https://doi.org/10.1101/2021.08.07.455511
Ai W, Liu Y, Mei M, Zhang X, Tan E, Liu H, Han X, Zhan H, Lu X (2022) A chromosome-scale genome assembly of the Mongolian oak (Quercus mongolica). Mol Ecol Resour 22:2396–2410. https://doi.org/10.1111/1755-0998.13616
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Ammitzboll H, Vaillancourt RE, Potts BM, Harrison PA, Brodribb T, Sussmilch FC, Freeman JS (2020) Independent genetic control of drought resistance, recovery, and growth of Eucalyptus globulus seedlings. Plant, Cell Environ 43:103–115. https://doi.org/10.1111/pce.13649
Barber HN (1955) Adaptive gene substitutions in Tasmanian eucalypts: I. Genes controlling the development of glaucousness. Evolution 9:1–15
Bernard A, Domergue F, Pascal S, Jetter R, Renne C, Faure J-D, Haslam RP, Napier JA, Lessire R, Joubès J (2012) Reconstitution of plant alkane biosynthesis in yeast demonstrates that Arabidopsis ECERIFERUM1 and ECERIFERUM3 are core components of a very-long-chain alkane synthesis complex. Plant Cell 24:3106–3118. https://doi.org/10.1105/tpc.112.099796
Bhanot V, Fadanavis SV, Panwar J (2021) Revisiting the architecture, biosynthesis and functional aspects of the plant cuticle: there is more scope. Environ Exp Bot 183:104364. https://doi.org/10.1016/j.envexpbot.2020.104364
Birney E, Clamp M, Durbin R (2004) GeneWise and genomewise. Genome Res 14:988–995. https://doi.org/10.1101/gr.1865504
Bourdenx B, Bernard A, Domergue F, Pascal S, Léger A, Roby D, Pervent M, Vile D, Haslam RP, Napier JA, Lessire R, Joubès J (2011) Overexpression of Arabidopsis ECERIFERUM1 promotes wax very-long-chain alkane biosynthesis and influences plant response to biotic and abiotic stresses. Plant Physiol 156:29–45. https://doi.org/10.1104/pp.111.172320
Butler JB, Freeman JS, Potts BM, Vaillancourt RE, Grattapaglia D, Silva-Junior OB, Simmons BA, Healey AL, Schmutz J, Barry KW, Lee DJ, Henry RJ, King GJ, Baten A et al (2018) Annotation of the Corymbia terpene synthase gene family shows broad conservation but dynamic evolution of physical clusters relative to Eucalyptus. Heredity 121:87–104. https://doi.org/10.1038/s41437-018-0058-1
Castillo-Davis CI, Bedford TBC, Hartl DL (2004) Accelerated rates of intron gain/loss and protein evolution in duplicate genes in human and mouse malaria parasites. Mol Biol Evol 21:1422–1427. https://doi.org/10.1093/molbev/msh143
Chaudhary K, Geeta R, Panjabi P (2021) Origin and diversification of ECERIFERUM1 (CER1) and ECERIFERUM3 (CER3) genes in land plants and phylogenetic evidence that the ancestral CER1/3 gene resulted from the fusion of pre-existing domains. Mol Phylogen Evol 159:107101. https://doi.org/10.1016/j.ympev.2021.107101
Cho S, Jin S-W, Cohen A, Ellis RE (2004) A phylogeny of Caenorhabditis reveals frequent loss of introns during nematode evolution. Genome Res 14:1207–1220. https://doi.org/10.1101/gr.2639304
Close DC, Davidson NJ, Shields CB, Wiltshire R (2007) Reflectance and phenolics of green and glaucous leaves of Eucalyptus urnigera. Aust J Bot 55:561–567. https://doi.org/10.1071/bt06037
Cohen NE, Shen R, Carmel L (2011) The role of reverse transcriptase in intron gain and loss mechanisms. Mol Biol Evol 29:179–186. https://doi.org/10.1093/molbev/msr192
Coulombe-Huntington J, Majewski J (2007) Characterization of intron loss events in mammals. Genome Res 17:23–32. https://doi.org/10.1101/gr.5703406
Deng B, Du W, Liu C, Sun W, Tian S, Dong H (2012) Antioxidant response to drought, cold and nutrient stress in two ploidy levels of tobacco plants: low resource requirement confers polytolerance in polyploids? Plant Growth Regul 66:37–47. https://doi.org/10.1007/s10725-011-9626-6
Domínguez E, Heredia-Guerrero JA, Heredia A (2011) The biophysical design of plant cuticles: an overview. New Phytol 189:938–949. https://doi.org/10.1111/j.1469-8137.2010.03553.x
Domínguez E, Heredia-Guerrero JA, Heredia A (2017) The plant cuticle: old challenges, new perspectives. J Exp Bot 68:5251–5255. https://doi.org/10.1093/jxb/erx389
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Edwards PB (1982) Do waxes on Eucalyptus leaves provide protection from grazing insects? Aust J Ecol 7:347–352. https://doi.org/10.1111/j.1442-9993.1982.tb01309.x
Fawcett JA, Rouzé P, Van de Peer Y (2011) Higher intron loss rate in Arabidopsis thaliana than A. lyrata is consistent with stronger selection for a smaller genome. Mol Biol Evol 29:849–859. https://doi.org/10.1093/molbev/msr254
Ferguson S, Jones A, Murray K, Andrew R, Schwessinger B, Borevitz J (2023) Plant genome evolution in the genus Eucalyptus driven by structural rearrangements that promote sequence divergence. bioRxiv 2(8). https://doi.org/10.1101/2023.04.19.537464
Gao Y, Zhang Z, Cheng J, Xian X, Li C, Wang Y (2023) Genome-wide identification of the CER1 gene family in apple and response of MdCER1-1 to drought stress. Funct Integr Genomics 23:17. https://doi.org/10.1007/s10142-022-00940-x
Goel M, Sun H, Jiao W-B, Schneeberger K (2019) SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol 20:277. https://doi.org/10.1186/s13059-019-1911-0
Gosney BJ, Potts BM, O’Reilly-Wapstra JM, Vaillancourt RE, Fitzgerald H, Davies NW, Freeman JS (2016) Genetic control of cuticular wax compounds in Eucalyptus globulus. New Phytol 209:202–215. https://doi.org/10.1111/nph.13600
Govaerts R, Sobral M, Ashton P, Barrie F, Holst BK, Landrum LR, Matsumoto K, Mazine FF, Nic Lughadha E, Proença C, Soares-Silva LH, Wilson PG, Lucas EJ (2008) World checklist of Myrtaceae. Kew Publishing, London
Grattapaglia D, Vaillancourt RE, Shepherd M, Thumma BR, Foley W, Kuelheim C, Potts BM, Myburg AA (2012) Progress in Myrtaceae genetics and genomics: Eucalyptus as the pivotal genus. Tree Genet Genom 8:463–508. https://doi.org/10.1007/s11295-012-0491-x
Hallam N, Chambers T (1970) Leaf waxes of genus Eucalyptus l’Héritier. Aust J Bot 18:335–386. https://doi.org/10.1071/BT9700335
Hannoufa A, McNevin J, Lemieux B (1993) Epicuticular waxes of eceriferum mutants of Arabidopsis thaliana. Phytochemistry 33:851–855. https://doi.org/10.1016/0031-9422(93)85289-4
He J, Li C, Hu N, Zhu Y, He Z, Sun Y, Wang Z, Wang Y (2022) ECERIFERUM1-6A is required for the synthesis of cuticular wax alkanes and promotes drought tolerance in wheat. Plant Physiol 190:1640–1657. https://doi.org/10.1093/plphys/kiac394
Healey AL, Shepherd M, King GJ, Butler JB, Freeman JS, Lee DJ, Potts BM, Silva-Junior OB, Baten A, Jenkins J, Shu S, Lovell JT, Sreedasyam A, Grimwood J et al (2021) Pests, diseases, and aridity have shaped the genome of Corymbia citriodora. Commun Biol 4:537. https://doi.org/10.1038/s42003-021-02009-0
Hoffmann B, Kahmen A, Cernusak LA, Arndt SK, Sachse D (2013) Abundance and distribution of leaf wax n-alkanes in leaves of Acacia and Eucalyptus trees along a strong humidity gradient in northern Australia. Org Geochem 62:62–67. https://doi.org/10.1016/j.orggeochem.2013.07.003
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2014) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Iglewicz B, Hoaglin D (1993) How to detect and handle outliers. ASQC Quality Press, Milwaukee
Izuno A, Hatakeyama M, Nishiyama T, Tamaki I, Shimizu-Inatsugi R, Sasaki R, Shimizu KK, Isagi Y (2016) Genome sequencing of Metrosideros polymorpha (Myrtaceae), a dominant species in various habitats in the Hawaiian Islands with remarkable phenotypic variations. J Plant Res 129:727–736. https://doi.org/10.1007/s10265-016-0822-3
Jeffares DC, Penkett CJ, Bähler J (2008) Rapidly regulated genes are intron poor. Trends Genet 24:375–378. https://doi.org/10.1016/j.tig.2008.05.006
Jenks MA, Tuttle HA, Eigenbrode SD, Feldmann KA (1995) Leaf epicuticular waxes of the eceriferum mutants in Arabidopsis. Plant Physiol 108:369–377. https://doi.org/10.1104/pp.108.1.369
Jones TH, Potts BM, Vaillancourt RE, Davies NW (2002) Genetic resistance of Eucalyptus globulus to autumn gum moth defoliation and the role of cuticular waxes. Can J for Res 32:1961–1969. https://doi.org/10.1139/x02-118
Jordan GJ, Harrison PA, Worth JRP, Williamson GJ, Kirkpatrick JB (2016) Palaeoendemic plants provide evidence for persistence of open, well-watered vegetation since the Cretaceous. Global Ecol Biogeogr 25:127–140. https://doi.org/10.1111/geb.12389
Keller G, Cao PB, San Clemente H, El Kayal W, Marque C, Teulières C (2013) Transcript profiling combined with functional annotation of 2,662 ESTs provides a molecular picture of Eucalyptus gunnii cold acclimation. Trees 27:1713–1735. https://doi.org/10.1007/s00468-013-0918-5
Kirkpatrick JB, Marks F (1985) Observations on drought damage to some native plant species in eucalypt forests and woodlands near Hobart, Tasmania. Pap Proc R Soc Tasman 119:15–21
Koch K, Hartmann KD, Schreiber L, Barthlott W, Neinhuis C (2006) Influences of air humidity during the cultivation of plants on wax chemical composition, morphology and leaf surface wettability. Environ Exp Bot 56:1–9. https://doi.org/10.1016/j.envexpbot.2004.09.013
Kondrashov FA (2012) Gene duplication as a mechanism of genomic adaptation to a changing environment. Proc Royal Soc B 279:5048–5057. https://doi.org/10.1098/rspb.2012.1108
Koornneef M, Hanhart CJ, Thiel F (1989) A genetic and phenotypic description of eceriferum (cer) mutants in Arabidopsis thaliana. J Hered 80:118–122. https://doi.org/10.1093/oxfordjournals.jhered.a110808
Külheim C, Padovan A, Hefer C, Krause ST, Köllner TG, Myburg AA, Degenhardt J, Foley WJ (2015) The Eucalyptus terpene synthase gene family. BMC Genom 16:450. https://doi.org/10.1186/s12864-015-1598-x
Kuzmin E, Taylor JS, Boone C (2022) Retention of duplicated genes in evolution. Trends Genet 38:59–72. https://doi.org/10.1016/j.tig.2021.06.016
Lee SB, Suh MC (2015) Advances in the understanding of cuticular waxes in Arabidopsis thaliana and crop species. Plant Cell Rep 34:557–572. https://doi.org/10.1007/s00299-015-1772-2
Lewandowska M, Keyl A, Feussner I (2020) Wax biosynthesis in response to danger: its regulation upon abiotic and biotic stress. New Phytol 227:698–713. https://doi.org/10.1111/nph.16571
Li H (2018) Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34:3094–3100. https://doi.org/10.1093/bioinformatics/bty191
Li H, Madden JL, Potts BM (1997) Variation in leaf waxes of the Tasmanian Eucalyptus species—I. Subgenus Symphyomyrtus. Biochem Syst Ecol 25:631–657. https://doi.org/10.1016/S0305-1978(97)00044-6
Li F, Zhou C, Weng Q, Li M, Yu X, Guo Y, Wang Y, Zhang X, Gan S (2015) Comparative genomics analyses reveal extensive chromosome colinearity and novel quantitative trait loci in Eucalyptus. PLoS ONE 10:e0145144. https://doi.org/10.1371/journal.pone.0145144
Li Q, Yu H, Cao PB, Fawal N, Mathé C, Azar S, Cassan-Wang H, Myburg AA, Grima-Pettenati J, Marque C, Teulières C, Dunand C (2015) Explosive tandem and segmental duplications of multigenic families in Eucalyptus grandis. Genome Biol Evol 7:1068–1081. https://doi.org/10.1093/gbe/evv048
Li T, Sun Y, Liu T, Wu H, An P, Shui Z, Wang J, Zhu Y, Li C, Wang Y, Jetter R, Wang Z (2019) TaCER1-1A is involved in cuticular wax alkane biosynthesis in hexaploid wheat and responds to plant abiotic stresses. Plant, Cell Environ 42:3077–3091. https://doi.org/10.1111/pce.13614
Li N, Li XZ, Song YQ, Yang ST, Li LL (2021) Genome-wide identification, characterization, and expression profiling of the ECERIFERUM (CER) gene family in Ziziphus jujube. Russ J Plant Physiol 68:828–837. https://doi.org/10.1134/S1021443721050101
Limin AE, Fowler DB (1989) The influence of cell size and chromosome dosage on cold-hardiness expression in the Triticeae. Genome 32:667–671. https://doi.org/10.1139/g89-496
Lin H, Zhu W, Silva JC, Gu X, Buell CR (2006) Intron gain and loss in segmentally duplicated genes in rice. Genome Biol 7:R41. https://doi.org/10.1186/gb-2006-7-5-r41
Liu M, Grigoriev A (2004) Protein domains correlate strongly with exons in multiple eukaryotic genomes – evidence of exon shuffling? Trends Genet 20:399–403. https://doi.org/10.1016/j.tig.2004.06.013
Liu X, Bourgault R, Galli M, Strable J, Chen Z, Feng F, Dong J, Molina I, Gallavotti A (2021) The FUSED LEAVES1-ADHERENT1 regulatory module is required for maize cuticle development and organ separation. New Phytol 229:388–402. https://doi.org/10.1111/nph.16837
Loh Y-H, Brenner S, Venkatesh B (2007) Investigation of loss and gain of introns in the compact genomes of pufferfishes (Fugu and Tetraodon). Mol Biol Evol 25:526–535. https://doi.org/10.1093/molbev/msm278
Ma M-Y, Che X-R, Porceddu A, Niu D-K (2015) Evaluation of the mechanisms of intron loss and gain in the social amoebae Dictyostelium. BMC Evol Biol 15:286. https://doi.org/10.1186/s12862-015-0567-y
Madeira F, Pearce M, Tivey ARN, Basutkar P, Lee J, Edbali O, Madhusoodanan N, Kolesnikov A, Lopez R (2022) Search and sequence analysis tools services from EMBL-EBI in 2022. Nucleic Acids Res 50:W276–W279. https://doi.org/10.1093/nar/gkac240
Marçais G, Delcher AL, Phillippy AM, Coston R, Salzberg SL, Zimin A (2018) MUMmer4: a fast and versatile genome alignment system. PLoS Comp Biol 14:e1005944. https://doi.org/10.1371/journal.pcbi.1005944
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z et al (2010) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229. https://doi.org/10.1093/nar/gkq1189
Milia G, Camiolo S, Avesani L, Porceddu A (2015) The dynamic loss and gain of introns during the evolution of the Brassicaceae. Plant J 82:915–924. https://doi.org/10.1111/tpj.12860
Myburg AA, Grattapaglia D, Tuskan GA, Hellsten U, Hayes RD, Grimwood J, Jenkins J, Lindquist E, Tice H, Bauer D, Goodstein DM, Dubchak I, Poliakov A, Mizrachi E et al (2014) The genome of Eucalyptus grandis. Nature 510:356–362. https://doi.org/10.1038/nature13308
Nawrath C (2006) Unraveling the complex network of cuticular structure and function. Curr Opin Plant Biol 9:281–287. https://doi.org/10.1016/j.pbi.2006.03.001
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Nicolle D (2022a) Native eucalypts of Victoria and Tasmania. Bloomings Books, Melbourne, South-eastern Australia
Nicolle D (2022b) Classification of the eucalypts (Angophora, Corymbia and Eucalyptus) Version 6. http://www.dn.com.au/Classification-Of-The-Eucalypts.pdf. Accessed 27 June 2023
Pascal S, Bernard A, Deslous P, Gronnier J, Fournier-Goss A, Domergue F, Rowland O, Joubès J (2018) Arabidopsis CER1-LIKE1 functions in a cuticular very-long-chain alkane-forming complex. Plant Physiol 179:415–432. https://doi.org/10.1104/pp.18.01075
Penny D, Hoeppner MP, Poole AM, Jeffares DC (2009) An overview of the introns-first theory. J Mol Evol 69:527–540. https://doi.org/10.1007/s00239-009-9279-5
Rizwan HM, Waheed A, Ma S, Li J, Arshad MB, Irshad M, Li B, Yang X, Ali A, Ahmed MAA, Shaheen N, Scholz SS, Oelmüller R, Lin Z et al (2022) Comprehensive genome-wide identification and expression profiling of eceriferum (CER) gene family in passion fruit (Passiflora edulis) under Fusarium kyushuense and drought stress conditions. Front Plant Sci 13:898307. https://doi.org/10.3389/fpls.2022.898307
Rodríguez-Trelles F, Tarrío R, Ayala FJ (2006) Origins and evolution of spliceosomal introns. Annu Rev Genet 40:47–76. https://doi.org/10.1146/annurev.genet.40.110405.090625
Rogozin IB, Carmel L, Csuros M, Koonin EV (2012) Origin and evolution of spliceosomal introns. Biol Direct 7:11. https://doi.org/10.1186/1745-6150-7-11
Rowland O, Lee R, Franke R, Schreiber L, Kunst L (2007) The CER3 wax biosynthetic gene from Arabidopsis thaliana is allelic to WAX2/YRE/FLP1. FEBS Lett 581:3538–3544. https://doi.org/10.1016/j.febslet.2007.06.065
Roy SW, Gilbert W (2005) The pattern of intron loss. Proc Natl Acad Sci USA 102:713–718. https://doi.org/10.1073/pnas.0408274102
Roy SW, Penny D (2007) On the incidence of intron loss and gain in paralogous gene families. Mol Biol Evol 24:1579–1581. https://doi.org/10.1093/molbev/msm082
Sakuradani E, Zhao L, Haslam TM, Kunst L (2013) The CER22 gene required for the synthesis of cuticular wax alkanes in Arabidopsis thaliana is allelic to CER1. Planta 237:731–738. https://doi.org/10.1007/s00425-012-1791-y
Samuels L, Kunst L, Jetter R (2008) Sealing plant surfaces: cuticular wax formation by epidermal cells. Annu Rev Plant Biol 59:683–707. https://doi.org/10.1146/annurev.arplant.59.103006.093219
Santos IBd, Lopes MdS, Bini AP, Tschoeke BAP, Verssani BAW, Figueredo EF, Cataldi TR, Marques JPR, Silva LD, Labate CA, Quecine MC (2019) The Eucalyptus cuticular waxes contribute in preformed defense against Austropuccinia psidii. Front Plant Sci 9:1978. https://doi.org/10.3389/fpls.2018.01978
Shepherd T, Wynne Griffiths D (2006) The effects of stress on plant cuticular waxes. New Phytol 171:469–499. https://doi.org/10.1111/j.1469-8137.2006.01826.x
Slee AV, Brooker H, Duffy M, West JG (2020) EUCLID. Eucalypts of Australia. CSIRO National Research Collections. https://apps.lucidcentral.org/euclid/text/intro/about.htm. Accessed 27 June 2023
Thornhill AH, Crisp MD, Külheim C, Lam KE, Nelson LA, Yeates DK, Miller JT (2019) A dated molecular perspective of eucalypt taxonomy, evolution and diversification. Aust Syst Bot 32:29–48. https://doi.org/10.1071/SB18015
Thornhill AH, Ho SYW, Külheim C, Crisp MD (2015) Interpreting the modern distribution of Myrtaceae using a dated molecular phylogeny. Mol Phylogen Evol 93. https://doi.org/10.1016/j.ympev.2015.07.007
Thrimawithana AH, Jones D, Hilario E, Grierson E, Ngo HM, Liachko I, Sullivan S, Bilton TP, Jacobs JME, Bicknell R, David C, Deng C, Nieuwenhuizen N, Lopez-Girona E et al (2019) A whole genome assembly of Leptospermum scoparium (Myrtaceae) for mānuka research. N Z J Crop Hortic Sci 47:233–260. https://doi.org/10.1080/01140671.2019.1657911
Tuhkanen S (1980) Climatic parameters and indices in plant geography. Almqvist & Wiksell, Uppsala
Turnbull J (1999) Eucalypt plantations. New for 17:37–52. https://doi.org/10.1023/A:1006524911242
Voelker J, Shepherd M, Mauleon R (2021) A high-quality draft genome for Melaleuca alternifolia (tea tree): a new platform for evolutionary genomics of myrtaceous terpene-rich species. Gigabyte https://doi.org/10.46471/gigabyte.28:28. https://doi.org/10.46471/gigabyte.28
Wang T, Hamann A, Yanchuk A, O’Neill GA, Aitken SN (2006) Use of response functions in selecting lodgepole pine populations for future climates. Global Change Biol 12:2404–2416. https://doi.org/10.1111/j.1365-2486.2006.01271.x
Wang H, Devos KM, Bennetzen JL (2014) Recurrent loss of specific introns during angiosperm evolution. PLoS Genet 10:e1004843. https://doi.org/10.1371/journal.pgen.1004843
Wang H, Ni X, Harris-Shultz K (2019) Molecular evolution of the plant ECERIFERUM1 and ECERIFERUM3 genes involved in aliphatic hydrocarbon production. Comput Biol Chem 80:1–9. https://doi.org/10.1016/j.compbiolchem.2019.02.009
Wang W, Das A, Kainer D, Schalamun M, Morales-Suarez A, Schwessinger B, Lanfear R (2020) The draft nuclear genome assembly of Eucalyptus pauciflora: a pipeline for comparing de novo assemblies. GigaScience 9:160. https://doi.org/10.1093/gigascience/giz160
Wang Y, Wang Y, Meng Z, Wei Y, Du X, Liang C, Zhang R (2021) Elevation of GhDREB1B transcription by a copy number variant significantly improves chilling tolerance in cotton. Planta 254:42. https://doi.org/10.1007/s00425-021-03686-1
William Roy S, Gilbert W (2006) The evolution of spliceosomal introns: patterns, puzzles and progress. Nat Rev Genet 7:211–221. https://doi.org/10.1038/nrg1807
Wu H, Liu L, Chen Y, Liu T, Jiang Q, Wei Z, Li C, Wang Z (2022) Tomato SlCER1–1 catalyzes the synthesis of wax alkanes, increasing drought tolerance and fruit storability. Hort Res 9:uhac004. https://doi.org/10.1093/hr/uhac004
Xue D, Zhang X, Lu X, Chen G, Chen Z-H (2017) Molecular and evolutionary mechanisms of cuticular wax for plant drought tolerance. Front Plant Sci 8:621. https://doi.org/10.3389/fpls.2017.00621
Yeats TH, Rose JK (2013) The formation and function of plant cuticles. Plant Physiol 163:5–20. https://doi.org/10.1104/pp.113.222737
Zhao S, Nie X, Liu X, Wang B, Liu S, Qin L, Xing Y (2022) Genome-wide identification of the CER gene family and significant features in climate adaptation of Castanea mollissima. Int J Mol Sci 23:16202. https://doi.org/10.3390/ijms232416202
Zhu K, Chen F, Liu J, Chen X, Hewezi T, Cheng Z-M (2016) Evolution of an intron-poor cluster of the CIPK gene family and expression in response to drought stress in soybean. Sci Rep 6:28225. https://doi.org/10.1038/srep28225
Acknowledgements
The authors thank Rangitāne O Manawatu, Genomics Aotearoa and the New Zealand eScience Infrastructure for providing access to the Leptospermum scoparium genome. We also thank Josquin Tibbits and Agriculture Victoria for providing access to the Eucalyptus globulus genome and Peter Harrison for providing R codes used for the study of environmental associations.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This research was supported by the Australian Research Council Discovery grant DP190102053 and the Instituto Nacional de Tecnología Agropecuaria (INTA), Argentina [MAH].
Author information
Authors and Affiliations
Contributions
M.A.H., J.B.B. and R.E.V. conceived the ideas and designed the methodology. M.A.H. led the analysis and writing of the manuscript under the supervision of J.B.B, R.E.V. and B.M.P. All authors contributed to the drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by L. Bianco.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hernández, M.A., Vaillancourt, R.E., Potts, B.M. et al. Insights into the evolution of the eucalypt CER1 and CER3 genes involved in the synthesis of alkane waxes. Tree Genetics & Genomes 20, 4 (2024). https://doi.org/10.1007/s11295-023-01637-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-023-01637-3