Abstract
Because of their protein cross-linking properties, transglutaminases are widely used in several industrial processes, including the food and pharmaceutical industries. Transglutaminases obtained from animal tissues and organs, the first sources of this enzyme, are being replaced by microbial sources, which are cheaper and easier to produce and purify. Since the discovery of microbial transglutaminase (mTGase), the enzyme has been produced for industrial applications by traditional fermentation process using the bacterium Streptomyces mobaraensis. Several studies have been carried out in this field to increase the enzyme industrial productivity. Researches on gene expression encoding transglutaminase biosynthesis were performed in Streptomyces lividans, Escherichia coli, Corynebacterium glutamicum, Yarrowia lipolytica, and Pichia pastoris. In the first part of this review, we presented an overview of the literature on the origins, types, mediated reactions, and general characterizations of these important enzymes, as well as the studies on recombinant microbial transglutaminases. In this second part, we focus on the application versatility of mTGase in three broad areas: food, pharmacological, and biotechnological industries. The use of mTGase is presented for several food groups, showing possibilities of applications and challenges to further improve the quality of the end-products. Some applications in the textile and leather industries are also reviewed, as well as special applications in the PEGylation reaction, in the production of antibody drug conjugates, and in regenerative medicine.
Graphic abstract

Similar content being viewed by others
Introduction
Starting in 1989, microbial transglutaminases (mTGase) have been produced for industrial applications through traditional fermentation process using Streptomyces mobaraensis bacterium. mTGase is produced by this microorganism as an extracellular enzyme, having a molecular mass of approximately 38 kDa. This enzyme acts in a wide range of pH and temperatures (pH 5.0 to 8.0, and active in between 40 and 70 °C). S. mobaraensis mTGase is Ca2+ independent, and its activation requires no special cofactors (Ando et al. 1989; Yokoyama et al. 2004).
Enzymatic modifications of proteins mediated by mTGase have been generally used as tools for improving the properties of a given target product. These enzymatic reactions provide high specificity, occurring under gentle reaction conditions, producing no toxic products (Fatima and Khare 2018). In recent years, researchers have intensified their search for application possibilities of the mTGase to obtain methods and products that can alter the technological and functional properties of final products, not only in the food industry, but also in several biochemical reactions.
In the first part of this review, we focused on general aspects of the origins, reactions, and characteristics of transglutaminases of mammals, non-mammals, and microbial sources. Some studies on recombinant transglutaminases were also covered. In this second-part of the review, we highlight the importance of mTGase in four major research areas showing potential of applications of this enzyme: the food, biotechnology, leather, and textile industries.
Food applications
Studies for the application of transglutaminases in food technology began in the nineties, after the discovery of mTGase in microorganisms such as Streptomyces mobaraensis (Streptoverticillium mobaraense) (Ando et al. 1989), Streptomyces cinnamoneum (Duran et al. 1998), and Bacillus subtilis (Suzuki et al. 2000). The first application of mTGase in food technology was reported by Gottmann and Sprössler in 1992 (1992), who reported that mTGase could be a cost-effective enzyme to be used in food applications. Two decades later mTGases are mainly used in the processing of meat, fish, dairy, and baking products (Strop 2014). mTGase modifies the functional properties of food proteins by incorporation of amines, crosslinking, deamidation, and bonding surfaces of foods. However, in protein-containing food systems, the cross-linking reaction proceeds prior to other reactions (Santhi et al. 2017). In Table 1 are shown relevant aspects of some investigations on the use of mTGase for the modification of properties in different foods, according to the nature of the protein substrate, the amount of enzyme used, and the conditions of enzymatic reactions. Understanding mTGase mechanisms of action in altering protein properties is of major importance for its industrial use, which has not been completely elucidated (Gaspar and de Góes-Favoni 2015).
The first industrial scale production of mTGase was performed by the Japanese company Ajinomoto Co., in collaboration with Amano Enzyme Co. (Nagoya, Japan). The interest of the scientific community in mTGases is demonstrated by approximately 615 published papers in the last five years (Fig. 1a) investigating their applicability, structural characteristics, and substrate specificities, whereas 346 of these papers deal with applications in the field of Food Science and Technology (Fig. 1b), the area showing the greatest interest in this enzyme, as shown in Fig. 1c (Web of Science: June 2019).
Meat and seafood products
Microbial tranglutaminases attracted initial interests of the food industry due to its ability to mold minced meat into a firm steak. The restructure of meat products ensures greater firmness causing little loss of quality during cooking (Lesiow et al. 2017). The cross-linking of proteins and other compounds of the gel system causes changes in the proteic fraction of food matrices, leading to improved texture and stability in terms of temperature denaturation, emulsifying properties, gelation, and increased water-binding capacity (Dondero et al. 2006). The mTGase yields a final product with retained organoleptic properties similar to conventional meat in terms of flavor, texture, appearance, and taste (Hong et al. 2016).
Several studies are reported on the use of mTGase in meat products. As shown in Table 1, the enzyme can be used in a wide range of temperatures, from 10 to 50 °C. Some of these studies also show that mTGase supplementation could increase the gel strength in meat products and cause positive effects on the development of meat proteins of pork, beef, chicken, and fish (Ahhmed et al. 2009a, b; Canto et al. 2014; Dondero et al. 2006; Feng et al. 2018; Hong and Chin 2010; Hong and Xiong 2012; Jira and Schwagele 2017; Monteiro et al. 2015; Sorapukdee and Tangwatcharin 2018; Wu et al. 2016).
Because meat products are highly proteic, the myofibrillar proteins have marked influence on the textural quality of these products. Actin and myosin, which constitute the majority of myofibrillar proteins, are important substrates of mTGase and can also be polymerized by its addition, thus improving the textural properties of structured meat products (Uran and Yilmaz 2018). The addition of mTGase also allows for the utilization of raw materials such as collagen and mechanically deboned meat in manufacturing meat products, with enhanced nutritive value by supplementation with amino acids otherwise deficient in these products (e.g. exogenous lysine) (Kieliszek and Misiewicz 2014).
Efforts to reduce the sodium content of meat products is an important issue concerning the health of people and to attend these demands, the meat industry is focusing on the development of techniques to reduce the use of salt in processed meat products, without impacting their quality (Atilgan and Kilic 2017). Strategies such as the use of mTGase can be applied in the manufacture of meat products with low salt content to avoid quality deterioration arising from this reduction, as suggested by Atilgan and Kilic (2017). These authors investigated the effects of mTGase, fibrin/thrombin (fibrimex), alginate, and their combinations on the quality of reduced-salt cooked meat. Their results indicated that the fibrimex/mTGase combination improved the texture properties of minced beef with low salt content.
However, for restructured meat in which fat can interfere with meat binding, it is essential to evaluate the grading level of formulated beef trimmings to enhance product quality or, at least, to avoid the minimum detrimental impact on product quality. The research of Sorapukdee and Tangwatcharin (2018) indicated that the most suitable raw beef for producing restructured steaks without detrimental effect on product quality, was beef trimmings containing up to 17% fat treated with 1% (weight fraction) of mTGase Activa TG-B. At this level of enzyme addition, both the sensory quality and increased tenderness were positively affected.
Dairy products
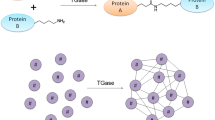
Improving the quality and functionality of the dairy products is been considered of paramount importance for better appreciation by people in a scenario of competitive dairy market. One of the most promising strategies to promote bio-functionality properties of dairy products is the cross-linking of milk proteins with transglutaminase. The use of mTGase can be a successful strategy to improve dairy products nutritional and technological characteristics, at the same time reducing the costs production by decreasing the amount of fat and stabilizer in the final product (Taghi Gharibzahedi et al. 2018). This enzyme has the ability to form intra- and intermolecular covalent crosslinks between two amino-acid residues in the structure of milk proteins. Both casein and whey α-lactalbumin and β-lactoglobulin are excellent acyl donors and/or acceptors substrates for transglutaminase, although some differences between them apply in relation to the crosslinking reaction (Færgemand and Qvist 1997; Oner et al. 2008; Rodriguez-Nogales 2006; Rossa et al. 2011). According to a study by Chen and Hsieh (2016), in a cascade reaction, mTGase catalyzes the cross-linking of κ-casein (κ-CN) and β-casein (β-CN) before it proceeds to cross-link the serum albumin (AS), α-lactalbumin (α-LA), αs1-casein (αs1-CN), αs2-casein (αs2-CN), and β-lactoglobulin (β-LG) moieties, as shown in Fig. 2. In this particular case, the caseins appear to be readily cross-linked because of their flexible, random-coil structures and the absence of any disulphide bonds in the αs1-CN and β-CN (Færgemand and Qvist 1997; Oner et al. 2008; Rodriguez-Nogales 2006; Rossa et al. 2011). Due to their compact globular structures, whey proteins tend to cross-link less efficiently. The β-LG is more susceptible and show a higher cross-linking rate than α-LA, but the β-LG is able to cross-link with the reduction of its disulphide bonds, whereas α-LA can be cross-linked without the reduction (Rodriguez-Nogales 2006; Rossa et al. 2011).

Adapted from Chen and Hsieh (2016)
Representation of the cross-linking reaction of milk proteins induced by mTGase. a The milk proteins without mTGase action are schematized; b the preference of mTGase for β-CN and κ-CN; c cross-linking occurs with all milk proteins with time.
The benefits brought about by the application of mTGase in dairy products include increased gel strength and improved storage stability and viscosity (Domagała et al. 2016). When mTGase is added to the system, it enhances heat-resistance and firmness of gel. Yogurt, a milk gel formed by acidic fermentation mediated by lactic-acid bacteria, has the disadvantage of serum separation upon change of temperature or physical impact. The addition of mTGase to yogurt can avoid this problem because mTGase improves the water holding capacity of the gel (Yokoyama et al. 2004). Ice creams treated with mTGase result in more consistent end-products, showing better aeration and foam stability. The mTGase also makes it possible to produce ice cream and cheese with low fat contents or reduced content of non-fat solids (Gaspar and de Góes-Favoni 2015; Yokoyama et al. 2004). The addition of mTGase during cheese preparation may increase the moisture content altering the palatability and the yield of different cheese products. In relation to surface texture of curds, ice-creams, milk and cheeses, it is noticed an improvement in the creaminess, homogeneity, smoothness, and consistency after mTGase is used in the production process (Wen-qiong et al. 2017).
Cereal based product
The first positive effects of mTGase application in baking was reported by Gottmann and Sprössler (1992). The applications of mTGase in cereal proteins, particularly wheat proteins (globulins, glutenins, gliadins, and prolamins), have attracted huge interest from the bakery industry (Mazzeo et al. 2013). The cross-links formed between the wheat proteins by the action of mTGase greatly influenced the characteristics of the products, determining the quality, functional and rheological properties of these systems, such as stability, elasticity, resilience, and water adsorption, with proper pore size along with adequate dough volume (Bonet et al. 2005; Gerrard et al. 1998; Gujral and Rosell 2004; Scarnato et al. 2017). The cross-linking reaction of mTGase promotes aggregation and polymerization, leading to the formation of polypeptide networks showing differentiated viscoelastic properties (Bonet et al. 2005; Gujral and Rosell 2004).
Gerrard et al. (1998) were the first researchers who used the mTGase in white bread. These authors suggested that the enzyme could have beneficial effects during the manufacture of bread, comparable to those produced by traditional chemical oxidant improvers. Scarnato et al. (2017) showed that using low amounts of mTGase, positive effects were observed on aspects of crumb and crust of bread, as well as in the rheological properties and physico-chemical properties of the dough. The release of some peptides from gluten obtained through the activity of mTGase can also influence the modulation of bread microbiota during the storage and consequently increase the final product shelf-life (Scarnato et al. 2017).
Another application of mTGase in the bakery industry is related to the production of pasta and instant noodles. In 1996, research by Sakamoto et al. (1996) showed that the treatment of noodles and pasta with mTGase prevented the deterioration of texture upon cooking and improved the strength of the products, even when low-grade flours were used in the manufacture, reducing the costs of production (Sakamoto et al. 1996; Yokoyama et al. 2004).
Soybean products
Soy protein isolate (IPS) is widely used as an important ingredient in Asiatic diets and in general processed foods due to its nutritional value and functional properties. IPS consists of glycinin (11S) and β-conglycinin (7S), which account for approximately 70% of its total protein content. These globulins are good substrates for mTGase activity (Qin et al. 2016; Song and Zhao 2014). Tang et al. (2005) investigated the use of mTGase on the properties and microstructures of IPS films molded with various plasticizers (glycerol, sorbitol, and 1:1 mixtures of glycerol and sorbitol). The cross-linking treatment by mTGase produced an effective method to improve the films cast properties of all tested plasticizers.
Tofu, a typical soybean curd product, is prepared by the coagulation of soybean proteins with the addition of Ca2+ and Mg2+ and/or glucono-δ-lactone. Coagulation or gelation of soymilk is the most important step in the production of tofu. A popular food in many countries, tofu shelf life is generally very short because its softness and smooth texture that prevents its sterilization. The introduction of mTGase in its processing produces an edge of texture control and enhances its quality, yielding a product with better consistency and silky texture and ability to tolerate temperature fluctuations (Chang et al. 2011).
Finally, proteins from sources other than soy can be covalently linked to soy protein by mTGase to produce combinations showing novel functionalities. For instance, conjugation of milk caseins or soybean globulins with ovomucin (an egg white glycoprotein), has shown to improve the emulsifying activity of the combined protein when compared to both isolated proteins (Kato et al. 1991; Yokoyama et al. 2004).
Food coating and edible films
The research to produce protein films as an alternative to petroleum-based polymeric materials has been receiving a great deal of attention in the food industry. Protein films can be used as coatings on fresh fruits and vegetables to increase the shelf life of these products. These films are non-toxic, natural, health safe, biodegradable, and might be edible. Protein edible films produced by the cross-linking action of mTGase present structural homogeneity, have a smooth surface, are mechanically resistant, and are gas-permeable (Porta et al. 2016). In the work to Rossi Marquez et al. (2017), apple weight losses during storage was significantly reduced, approximately 80%, after 10 days when the samples were coated with whey protein grafted film with pectin and transglutaminase. Similarly, this grafted film was able to prevent weight loss of potato and carrot samples until the 6th day of storage.
The research carried out by Fernandez-Bats et al. (2018) showed that it was possible to obtain mesoporous silica nanocomposite bioplastics prepared by using bitter vetch (Vicia ervilia) proteins crosslinked by mTGase, which showed improved gas and water vapor barrier properties. The prepared material showed antimicrobial and antifungal activities, possibly increased by nisin addition to the film-forming solutions, suggesting their potential application as an active bio-preservative packaging to improve the shelf life of a variety of different food products.
Health aspects concerning the use of transglutaminase in food industry
Because of increased applications of mTGase in food, important health concerns appeared, pressuring the need for regulations to inform people on the safety when consuming products containing this enzyme. In 1998, Motoki and Seguro (1998) showed that the only difference between food containing mTGase-modified proteins and native proteins was the number of links between glutamine and lysine residues (G–L). This chemical modification is also present when proteic foods are heated, for example, in cooking, generating the G–L bond. In this respect, humans have been ingesting foods rich in G–L residues since the discovery of fire and cooking. Although not scientifically demonstrated, the safety of the G–L modified linkage can be assumed by the long-term consumption of the G–L moiety in cooked foods (Motoki and Seguro 1998). On the other hand, Bernard et al. (1998) studied the mutagenesis and toxicity risk presented by the addition of mTGase in food preparations, tested in experimental animals. Their results suggest that the acute toxicity of the enzyme seems to be relatively low, since it was not observed any mortality, morbidity, or signs of toxicity at doses of 2 g/kg body weight.
There have been some evidences of increased nutritional properties of foods enzymatically modified by mTGase. According to studies conducted by Xing et al. (2016), the addition of mTGase to soy extract in preparation of tofu lead to modifications of proteins that increased the perception of satiety and reduced the allergenicity towards soy proteins. In another application concerning allergenicity, shrimp products processed using mTGase showed reduced allergenicity due to glycosylation of proteins catalyzed by this enzyme (Yuan et al. 2017).
Concerning bakery products, Zhou et al. (2017a) have shown that mTGase can effectively transamidate gliadin peptides and gluten proteins, thus concluding that mTGase lowered the allergenicity and immunogenicity caused by wheat flours. The resulting peptides are barred to cross intestinal mucosa where they initiate the celiac immunological activity. These results demonstrate a potential strategy to prevent cereal toxicity in celiac disease (Zhou et al. 2017b). However, multiple mTGase linked proteins are immunogenic in celiac disease patients. In the study conducted by Lerner and Matthias (2015), the authors indicate that the use of this enzyme can further increase antigenic load presented to the immune system and increase the risk for gluten-sensitive populations. In a recent research, Matthias et al. (Matthias et al. 2016) have suggested that mTGase increases immunogenicity in children with celiac disease because mTGase antibodies correlates to intestinal damage in the same degree as transglutaminase human tissue antibodies. Authors suggested that further investigation is necessary to elucidate the role of anti-mTGase antibodies in this disease.
Although scientific findings reported in the literature regarding the safety of the use of mTGase in foods can be classified as inconclusive, the FDA has approved the use of mTGase as a “Generally Recognized as Safe—GRAS” for food applications since 1998. This enzyme is considered an adjunct of technology and it is not regarded as an ingredient, and therefore does not need to be listed in the composition of ingredients of the commercial product (Romeih and Walker 2017; Taghi Gharibzahedi et al. 2018).
Biotechnology applications of mTGase
The biotechnological applications of transglutaminases are one of the fastest growing areas on mTGase research. Classical applications of transglutaminases in biomedical research include PEGylation, the production of antibody–drug conjugates, tissue engineering, regenerative medicine, and the production of microparticles for enteric delivery of substances of interest in the food and pharmaceutical industry. Finally, we will be briefly reviewing the use of transglutaminases in the treatment of textiles and leather.
Enzymes immobilization mediated by mTGase-catalyzed bioconjugation
Protein immobilization in solid supports has been used as a technique for biotechnological applications of enzymes, offering several advantages over the use of free forms, such as easing separation from reaction media and the possibility of reuse (Duarte et al. 2017; Mateo et al. 2007; Matte et al. 2014; Rodrigues et al. 2013). In general, proteins bound to functional groups on supports show high stability due to reduced protein loss to the medium. The immobilization of proteins through covalent bond formation has been routinely carried out using chemically-activated supports or chemical cross-linking reagents (Mateo et al. 2007). However, because the presence of multiple functional groups on protein surfaces, proteins are in general randomly attached onto supports, resulting in the reduction of total enzymatic activity. Several techniques have been devised to preserve the activity of biomolecules upon immobilization, among them the immobilization using mTGase as site-specific binding (Tominaga et al. 2004). The immobilization mediated by transglutaminase-catalyzed bioconjugations offers the advantage of improved selectivity and compatibility with sensitive biological systems relative to traditional chemical methodologies (Wang et al. 2019).
The mTGase is unique in catalyzing the acyl transfer reaction between a primary amine and the γ-carboxyamide group of glutamine (Gln) residues in peptides and proteins. When the ɛ-amino group of lysine (Lys) residues in protein acts as an acyl acceptor, cross-linking of proteins becomes possible through the ɛ-(γ-Gln)Lys bond, resulting in the formation of a new γ-glutaminyl covalent link (Kamiya et al. 2003; Li et al. 2018; Motoki and Seguro 1998; Wang et al. 2019). The mTGase displays broad acyl-acceptor substrate specificity, enabling the use of a variety of scaffolds with primary amine groups as solid supports, such as the polysaccharide chitosan and gelatin (Li et al. 2018).
The study of Tominaga et al. (2004) demonstrated site-specific immobilization via covalent attachment of recombinant alkaline phosphatase with a specific peptide linker by mTGase. To allow the mTGase-mediated site-specific immobilization, a solid support of casein-coated polyacrylic resin was designed to display mTGase recognition sites on its surface. It was found that this immobilization exhibited much higher specific activity, with higher stability upon repeated use than the biocatalyst prepared via chemical modification.
Wang et al. (2019) reported the controlled, site-specific and covalent cross-linking of an engineered enterokinase on amine-modified magnetic nanoparticles via mTGase-catalyzed bioconjugation for the development of the oriented-immobilized enzyme. A glutamyl (Gln-donor tag) was genetically incorporated into the C-terminus of enterokinase. An amide linkage was formed between the glutamyl group of Gln tag and the primary amines of the support via the covalent immobilization catalyzed by mTGase. Upon the site-specific immobilization, approximately 90% enterokinase activity was retained, and the biocatalyst exhibited more than 85% of initial enzymatic activity reusable stability over a month (Wang et al. 2019).
In the work of Synowiecki and Wołosowska (2006), a β-glucosidase from Sulfolobus shibatae was immobilized on silica gel modified with 3-aminopropyl-triethoxysilane using transglutaminase as a cross-linking factor, and the immobilization process did not influence the optimum pH and temperature of substrate hydrolysis (Synowiecki and Wołosowska 2006). On the other hand, Bechtold et al. reported the preparation of protein G-soybean peroxidase conjugate catalyzed by mTGase with yields calculated to be only about 0.1%, suggesting the difficulty in controlling the reaction when using native proteins (Bechtold et al. 2000).
PEGylation
At the end of the 1960s, Davis proposed the idea of conjugating PEG [poly (ethylene glycol)] to a protein, i.e., to “PEGylate” a protein (Davis 2002; Hoffman 2016). Since then, the PEGylation is considered one of the most successful methods to prolong the circulatory half-life and reduce the in vivo immunogenicity of therapeutic proteins, among many other applications in pharmacology (Pasut and Veronese 2012).
PEG is biocompatible, lacks immunogenicity and antigenicity, is soluble in water and other organic solvents, is readily cleared from the body, and has high mobility in solution and, more important, it is not toxic, making this the polymer of choice for bioconjugations. PEG use was approved by the FDA in the early 1990s (Bhattarai et al. 2005; Harris and Chess 2003; Mariniello et al. 2014; Roberts et al. 2002). Thus, a number of protein/PEG conjugates, are available in the market such as for the treatment of chronic hepatitis C (PEGinterferon α-2a and α-2b), for the treatment of acute lymphoblastic leukaemia (mPEG-L-Asparaginase), to treat severe combined immunodeficiency (SCID) disease (mPEG-Adenosine Deaminase), and to treat acromegaly (PEG-visomant) (Banerjee et al. 2012).
Chemical strategies used for the PEGylation of proteins produces random derivatives of lysine (Lys) residues, leading to heterogeneity and decreased bioactivity of the products (da Silva Freitas et al. 2013). Instead, the use of transglutaminase for the covalent attachment of PEG molecules to pharmaceutical proteins shows stringent substrate specificity, and site specific modification or PEGylation of the Gln residues bound to the proteins on the substrates can be obtained. (Fontana et al. 2008).
Because transglutaminases have partial selectivity to the carboxamide substrate, they are interesting options for the PEGylation of proteins. However, for the reaction to occur, the carboxamide must be in the flexible part of the protein molecule. (Dozier and Distefano 2015; Fontana et al. 2008). Consequently, mTGase has been intensively used to site-specifically incorporate mPEG–NH2 to the reactive Gln residue of proteins (da Silva Freitas et al. 2013). The reactive Gln residues modified by mTGase must locate at the disordered protein regions and satisfy its sequence requirement. As many target proteins lack the reactive Gln residues that can satisfy the structural and the sequence requirement of mTGase, its use is limited (da Silva Freitas et al. 2013; Mero et al. 2009). Several researches have been developed in this area and some of them are listed in Table 2.
So far, only a limited number of researches has been carried out on mTGase-mediated protein modification at the level of Lys residues. One of them is the work, of Zhou et al. (2016b), who linked carboxybenzyl-glutaminyl-glycine (CBZ-QG) to mPEG amine to form CBZ-QG-mPEG for the PEGilation of cytochrome C.
The hydroxyethyl starch (HES), which is a biodegradable derivative of starch, can be an alternative to PEG as blood plasma volume expander and in the design of drug delivery systems (Treib et al. 1999). It has been reported the use of HES conjugation using mTGase to produce fully biodegradable polymer–drug and polymer–protein conjugates (Besheer et al. 2009).
Antibody drug conjugates (ADCs)
Another promising technology is the use of mTGase to attach antibodies to diverse compounds in order to produce antibody–drug conjugates (ADC). ADC are emerging therapeutic agents in the treatment of cancer, using antibodies to selectively deliver a cytotoxic compound to tumor cells, thus improving the therapeutic index of chemotherapeutic agents, and showing better safety potential than nontargeted cytotoxics (Anami et al. 2017; Strop et al. 2013). One of the major challenges in the development of ADC is the application of suitable linkers to conjugate drugs to antibodies (Yao et al. 2016). The ADC have been largely manufactured by using chemical conjugation methods, generally resulting in heterogeneous mixtures of ADC having different physical and pharmacokinetic properties of the proposed ones (Axup et al. 2012; Dennler et al. 2014; Junutula et al. 2008; Okeley et al. 2013; Shen et al. 2012; Strop et al. 2013; Xiao et al. 2013; Zuberbühler et al. 2012).
An alternative strategy to the chemical modification of ADC is the use of mTGase because the enzyme will prevent the formation of these heterogeneous mixtures. Moreover, it is possible to introduce appropriate amine containing linkers making the mTGase able to conjugate structurally diverse probes and drugs (Ohtsuka et al. 2000). Strop et al. (2013) investigated how the conjugation site influences the stability, toxicity and efficacy of ADC obtained by mTGase reaction and whether these differences could be directly attributed to the binding position. By designing a “glutamine label”, 90 sites were tested to attach several compounds and 12 sites showing a high degree of conjugation were found.
A two-step chemo-enzymatic approach, where mTGase binds a spacer entity that is reactive to the antibody, and subsequently reacts with the antimitotic toxin monomethyl auristatin E (MMAE), produced the highly homogeneous trastuzumab-MMAE conjugate with DAR (Drug-Antibody Ratios) of 2 (Dennler et al. 2014). Some ADC currently in use in clinical development based on target antigens using tranglutaminase are: PF-06664178, Trop-2 ADC, RN927C (Phase I, for treatment of ovarian cancers, non-small cell lung cancer and breast cancer—site-specific transglutaminase tag, AcLys-VC-PABC linker) and PF-06647020, h6M24-vc0101, PTK7-targeted ADC (Phase I, for treatment of non-small-cell lung carcinoma, triple-negative breast cancer and ovarian cancers—transglutaminase tag (LLQGA) located at the C-terminus of the antibody heavy chain, cleavable VC-PABC-linker) (Damelin et al. 2017; Nejadmoghaddam et al. 2019; Sachdev et al. 2016; Strop et al. 2016).
Several other investigations have been reported on the production of monoclonal antibodies using mTGases and are well documented in recent works (Dennler et al. 2014; Farias et al. 2014; Grünberg et al. 2013; Jeger et al. 2010; Lhospice et al. 2015; Siegmund et al. 2015; Spidel et al. 2017; Strop et al. 2013).
Tissue engineering and regenerative medicine
The term Tissue Engineering (TE) was first introduced in 1993 by Langer and Vacanti (1993) to describe an interdisciplinary field encompassing cell biology, material science, chemistry, molecular biology, engineering, and medicine, with the objective of developing advanced biological tissues and organs. These engineered biological materials are intended to maintain, improve, or restore functionalities of natural tissues combining scaffolds, cells and/or bioactive molecules (Griffith and Swartz 2006; Langer and Vacanti 1993; Lee et al. 2014; O'Brien 2011). The potential applications are being investigated in the field of tissue engineering of bones, cartilage, cardiac system, pancreas, and the vascular system, among others (Zhu and Tramper 2008). The main bulk of research in this area has been focused in the development of biomaterials capable of mimicking the structure and composition of the extracellular matrix. Such biomaterials must present biocompatibility and biodegradability and should not be toxic. In addition, the production and processing of biomaterials must be easy and scalable. Because hydrogels have high plasticity and high moisture content they have been the most important biomaterials employed in tissue engineering (Polak 2010; Toh and Loh 2014). Hydrogels can be formed from gelatin, collagen, chitosan, hyaluronic acid, and sodium alginate, as well as synthetic materials such as polylactide, polylactic-co-glycolic acid copolymer, polyethylene glycol, polycaprolactone, and polyacrylamide (El-Sherbiny and Yacoub 2013). Gelatin is a protein derived from the hydrolysis of collagen with characteristics of biodegradability and cell adhesion capacity, considered as GRAS material by the FDA and it has a long history of safe use in food products, pharmaceuticals and cosmetics (Elzoghby et al. 2012). Unfortunately, owing to a lack of mechanical strength and sensitivity to in vivo enzymes, the biomedical applications of gelatin is limited being necessary to increase its physical performance and to strengthen its resistance against enzymes hydrolyses (Zhao et al. 2016). To achieve this goal, crosslinks are usually introduced in biomaterials such as collagen mediated by mTGase, replacing physical methods like dehydrothermal drying (DHT) and UV-irradiation, among others, and chemical crosslinking mediated by glutaraldehyde, formaldehyde, and 1-ethyl-3-(3-dimethylaminopropyl)-carbodiimide (EDC). The physical methods produce weak bonds that have a high risk of degradation, whereas the chemical methods use often toxic compounds that must be removed from hydrogels before being applied (Stachel et al. 2010; Yang et al. 2018). Therefore, the substitution of these methods by the enzymatic application of mTGases in order to generate hydrogels are among the most promising technologies to obtain biomaterials, since the mTGase-mediated process presents no risk of toxicity and eases the preparation of the materials, showing high mechanical stabilities (Milczek 2018). There is a plethora of research showing the use of transglutaminases to produce hydrogels, some of them listed in Table 3.
In 1986, Charles Hull described for the first time the technique of 3D bioprinting, which has since been used to produce a large variety of scaffolds in different tissue engineering areas. The term 3D bioprinting is used to describe the precise layering of cells, biologic scaffolds, and biologic factors with the goal of building a biological tissue (Bishop et al. 2017). The vascularization of 3D scaffolds is crucial for their functionalities, assuring the delivery of nutrients and oxygen to tissues, promoting cell proliferation and subsequent development of the new tissue (Castells-Sala et al. 2013). The main techniques used in tissue engineering by 3D bioprinting are stereolithography, extrusion-based, laser-assisted and inkjet-based printing (Derakhshanfar et al. 2018), and a sample of the investigations using 3D printing for tissue engineering with the use of transglutaminase are listed in the Table 3. In this technique, a great variety of polymers, hydrogels, bioceramics, among other biomaterials, have been used. Natural biological materials, such as cells, can also be employed in 3D bioprinting (Tappa and Jammalamadaka 2018).
Transglutaminase-crosslinked microparticles for enteric delivery
The microencapsulation technique is widely used in the fields of food, pharmaceuticals, and biotechnology in order to preserve physicochemical and sensorial attributes and to produce control-released compounds. Microencapsulation is obtained by the use of various techniques, such as spray drying, cooling, extrusion coating, fluidized bed coating, liposome retention, inclusion complexation, centrifugal extrusion, and rotational suspension separation, techniques that are chosen based on final product characteristics and costs (Desai and Park 2005). Recently, another technique has been described, the complex coacervation, which produces high encapsulation efficiencies, and has since been widely used in food and pharmaceutical industries, because it does not require harsh production conditions (temperature, pressure, pH, etc.) (Tello et al. 2016). This technique uses a combination of encapsulating agents of opposing charges to create electrostatic attraction between two molecules, and other interactions, such as hydrogen bonds and hydrophilic interactions, that also contribute to the formation of complexes (Ach et al. 2014). As the nature of these bonds is weak, there is the need to strengthen the interactions between polymers. For this purpose, mTGase has been used as a cross-linking agent showing promising results (Sanchez and Renard 2002). In Table 4 is presented a summary the microencapsulation and complex coacervation techniques found in literature involving the use of microbial transglutaminase.
Transglutaminase applications in textile industry
The textile finishing industry has been the focus of considerable criticism because it uses traditional chemical treatments in wool processing, which is perceived as highly damaging to the environment. Unfortunately, the alternative enzymatic processes using proteases can cause an excessive loss of fabric weight and yarn strength. Therefore, the use of transglutaminases in treatments of wool and leather fabrics has become extensively explored in order to develop appropriate technologies based on the use of this enzyme. It has been found that mTGase is capable of recovering properties of wool and silk treated with chemicals and enzymes used at different processing stages, such as cleaning, carding, bleaching, combing, drawing, spinning, and twisting (Tesfaw and Assefa 2014). Wool fabrics treated with Streptomyces hygroscopicus mTGase showed recovered fiber structures that were damaged during protease treatments (Du et al. 2007). The application of Guinea pig liver transglutaminase or the mTGase isolated from Streptoverticilium mobaraense in wool processing resulted in the reduction of the propensity of wool yarn or fabric to shrink, and to improve yarn resistance, suggesting that transglutaminases can remediate the negative effects of proteolytic processing of the wool (Cortez et al. 2004). Mojsov (2017) showed that the characteristics of wool fabric pretreated with proteolytic enzymes and transglutaminase is comparable to untreated wool fabric. The author points to the following benefits of treating wool with mTGase: improvement in fabric softness, increased absorption characteristics, and resistance to pilling and retraction of the felting (Mojsov 2017).
Wool garments industrialized using fabrics treated with mTGase are likely to have increased resistance to domestic washing. Biological detergents containing proteases can cause irreversible damage to the fiber, leading to loss of fabric strength, shape, and color fading (Cortez et al. 2005). However, combining the advantages of using both proteases and transglutaminases in a simultaneous enzymatic treatment of wool, resulted in the development of a bioprocess for machine washable wool with insignificant fiber damage (Gaffar Hossain et al. 2008). Casein incorporated to wool mediated by mTGase was used as a surface coating material for smoothing the texture of the wool fiber by coating or filling the damaged scales in wool yarn (Cui et al. 2011).
Finally, excellent antibacterial properties were obtained when Ɛ-Poly-l-lysine (Ɛ-PL), which is a natural biomacromolecule having a broad spectrum of antibacterial activity, was grafted onto the wool fiber via mTGase, showing 97% bacteriostasis to Escherichia coli (Wang et al. 2010).
Transglutaminase applications in leather processing
The process of filling, which is the introduction of materials into the voids between leather fibers in order to smooth surface irregularities is considered one of the most important steps in leather processing, used to increase material quality. Common materials used as fillers are glucose, flour, and gum, as well as enzyme-modified gelatin and casein, the last two being cross-linked with leather proteins by the action of mTGase (Zhu and Tramper 2008). Experimental results showed that fillers incorporated by mTGase were firmly bound to the leather and would not be easily removed during further processing (Taylor et al. 2006).
Finally, the use of gelatin-sodium caseinate modified by mTGase was investigated regarding subjective aspects of leather (visual aspects, touch, etc.), as well as for its mechanical and structural properties. The application of mTGase improved the subjective aspects, without significantly affecting the mechanical properties such as tensile strength and elongation at break (Liu et al. 2011).
Conclusion
We addressed the several uses of microbial transglutaminases in the food, pharmaceutical, and biotechnology industries. The applications of mTGase have important implications for the development of these industries, producing new products at low cost, improving the application and quality of food, pharmaceuticals, and other goods such as wool and leather, designed for improving human life in a more sustainable way. mTGases became crucial to produce processed meat and seafood products, dairy products, bread, noodle, soybean products, and to produce coating and edible films. In more sophisticated fields, mTGase has become relevant in PEGylation, antibody drug conjugates, tissue engineering, regenerative medicine, production of microparticles for enteric delivery, directly impacting health products and services. Due to its importance and value aggregation to final products, research on the applications of mTGases is ever growing, showing many possibilities to produce new materials and improving the quality of the existing ones. Further research should focus on the bioprocess technology to reduce production costs of mTGases and enhance their biochemical properties.
References
Ach D, Briançon S, Broze G, Puel F, Rivoire A, Galvan J-M, Chevalier Y (2014) Formation of microcapsules by complex coacervation. Can J Chem Eng 93:183–191. https://doi.org/10.1002/cjce.22086
Agyare KK, Addo K, Xiong YL (2009) Emulsifying and foaming properties of transglutaminase-treated wheat gluten hydrolysate as influenced by pH, temperature and salt. Food Hydrocoll 23:72–81. https://doi.org/10.1016/j.foodhyd.2007.11.012
Ahhmed AM, Kuroda R, Kawahara S, Ohta K, Nakade K, Aoki T, Muguruma M (2009a) Dependence of microbial transglutaminase on meat type in myofibrillar proteins cross-linking. Food Chem 112:354–361. https://doi.org/10.1016/j.foodchem.2008.05.078
Ahhmed AM, Nasu T, Huy DQ, Tomisaka Y, Kawahara S, Muguruma M (2009b) Effect of microbial transglutaminase on the natural actomyosin cross-linking in chicken and beef. Meat Sci 82:170–178. https://doi.org/10.1016/j.meatsci.2009.01.007
Ajay T, Scott Alexander I, Udi S, Priyadarshini M, Vrushali B, Subbu V (2018) Contact guidance for cardiac tissue engineering using 3D bioprinted gelatin patterned hydrogel. Biofabrication 10:025003. https://doi.org/10.1088/1758-5090/aaa15d
Alarake NZ, Frohberg P, Groth T, Pietzsch M (2017) Mechanical properties and biocompatibility of in situ enzymatically cross-linked gelatin hydrogels. Int J Artif Organs 40:159–168. https://doi.org/10.5301/ijao.5000553
Anami Y et al (2017) Enzymatic conjugation using branched linkers for constructing homogeneous antibody–drug conjugates with high potency. Org Biomol Chem 15:5635–5642. https://doi.org/10.1039/C7OB01027C
Ando H et al (1989) Purification and characteristics of a novel transglutaminase derived from microorganisms. Agric Biol Chem 53:2613–2617. https://doi.org/10.1271/bbb1961.53.2613
Atilgan E, Kilic B (2017) Effects of microbial transglutaminase, fibrimex and alginate on physicochemical properties of cooked ground meat with reduced salt level. J Food Sci Technol 54:303–312. https://doi.org/10.1007/s13197-016-2463-x
Axup JY et al (2012) Synthesis of site-specific antibody-drug conjugates using unnatural amino acids. Proc Natl Acad Sci 109:16101. https://doi.org/10.1073/pnas.1211023109
Babiker EE (2000) Effect of transglutaminase treatment on the functional properties of native and chymotrypsin-digested soy protein. Food Chem 70:139–145. https://doi.org/10.1016/S0308-8146(99)00231-9
Banerjee SS, Aher N, Patil R, Khandare J (2012) Poly(ethylene glycol)-prodrug conjugates: concept, design, and applications. J Drug Deliv 2012:103973. https://doi.org/10.1155/2012/103973
Bechtold U, Otterbach JT, Pasternack R, Fuchsbauer H-L (2000) Enzymic preparation of protein G-peroxidase conjugates catalysed by transglutaminase1. J Biochem 127:239–245. https://doi.org/10.1093/oxfordjournals.jbchem.a022600
Bernard BK, Tsubuku S, Shioya S (1998) Acute toxicity and genotoxicity studies of a microbial transglutaminase. Int J Toxicol 17:703–721. https://doi.org/10.1080/109158198225928
Besheer A, Hertel TC, Kressler J, Mader K, Pietzsch M (2009) Enzymatically catalyzed HES conjugation using microbial transglutaminase: proof of feasibility. J Pharm Sci 98:4420–4428. https://doi.org/10.1002/jps.21675
Bhattarai N, MatsenFrederick A, Zhang M (2005) PEG-grafted chitosan as an injectable thermoreversible hydrogel. Macromol Biosci 5:107–111. https://doi.org/10.1002/mabi.200400140
Bishop ES et al (2017) 3-D bioprinting technologies in tissue engineering and regenerative medicine: current and future trends. Genes Dis 4:185–195. https://doi.org/10.1016/j.gendis.2017.10.002
Bode F, da Silva MA, Drake AF, Ross-Murphy SB, Dreiss CA (2011) Enzymatically cross-linked tilapia gelatin hydrogels: physical, chemical, and hybrid networks. Biomacromol 12:3741–3752. https://doi.org/10.1021/bm2009894
Bode F, da Silva MA, Smith P, Lorenz CD, McCullen S, Stevens MM, Dreiss CA (2013) Hybrid processes in enzymatically gelled gelatin: impact on, macroscopic properties and cellular response. Soft Matter 9:6986–6999. https://doi.org/10.1039/c3sm00125c
Bonet A, Caballero PA, Gómez M, Rosell CM (2005) Microbial transglutaminase as a tool to restore the functionality of gluten from insect-damaged wheat. Cereal Chem 82:425–430. https://doi.org/10.1094/CC-82-0425
Broderick EP, O'Halloran DM, Rochev YA, Griffin M, Collighan RJ, Pandit AS (2004) Enzymatic stabilization of gelatin-based scaffolds. J Biomed Mater Res B 72B:37–42. https://doi.org/10.1002/jbm.b.30119
Canto AC et al (2014) Physico-chemical and sensory attributes of low-sodium restructured caiman steaks containing microbial transglutaminase and salt replacers. Meat Sci 96:623–632. https://doi.org/10.1016/j.meatsci.2013.08.003
Castells-Sala C et al (2013) Current applications of tissue engineering in biomedicine. J Biochip Tissue Chip. https://doi.org/10.4172/2153-0777.S2-004
Chang Y-H, Shiau S-Y, Chen F-B, Lin F-R (2011) Effect of microbial transglutaminase on the rheological and textural characteristics of black soybean packed tofu coagulating with agar. LWT: Food Sci Technol 44:1107–1112. https://doi.org/10.1016/j.lwt.2010.10.020
Chen CC, Hsieh JF (2016) Microwave-assisted cross-linking of milk proteins induced by microbial transglutaminase. Sci Rep 6:39040. https://doi.org/10.1038/srep39040
Chen T, Small DA, McDermott MK, Bentley WE, Payne GF (2003) Enzymatic methods for in situ cell entrapment and cell release. Biomacromol 4:1558–1563. https://doi.org/10.1021/bm034145k
Chen PY, Yang KC, Wu CC, Yu JH, Lin FH, Sun JS (2014a) Fabrication of large perfusable macroporous cell-laden hydrogel scaffolds using microbial transglutaminase. Acta Biomater 10:912–920. https://doi.org/10.1016/j.actbio.2013.11.009
Chen Z, Raj D, Victor P-L, Nancy K (2014b) PEGylation of lysine residues improves the proteolytic stability of fibronectin while retaining biological activity. Biotechnol J 9:1033–1043. https://doi.org/10.1002/biot.201400115
Cortez J, Bonner PLR, Griffin M (2004) Application of transglutaminases in the modification of wool textiles. Enzyme Microb Technol 34:64–72. https://doi.org/10.1016/j.enzmictec.2003.08.004
Cortez J, Bonner PLR, Griffin M (2005) Transglutaminase treatment of wool fabrics leads to resistance to detergent damage. J Biotechnol 116:379–386. https://doi.org/10.1016/j.jbiotec.2004.12.007
Cui L, Fan X, Wang P, Wang Q, Fu G (2011) Casein and transglutaminase-mediated modification of wool surface. Eng Life Sci 11:201–206. https://doi.org/10.1002/elsc.201000110
da Silva MA, Bode F, Drake AF, Goldoni S, Stevens MM, Dreiss CA (2014) Enzymatically cross-linked gelatin/chitosan hydrogels: tuning gel properties and cellular response. Macromol Biosci 14:817–830. https://doi.org/10.1002/mabi.201300472
da Silva MA, Bode F, Grillo I, Dreiss CA (2015) Exploring the kinetics of gelation and final architecture of enzymatically cross-linked chitosan/gelatin gels. Biomacromol 16:1401–1409. https://doi.org/10.1021/acs.biomac.5b00205
da Silva Freitas D, Mero A, Pasut G (2013) Chemical and enzymatic site specific PEGylation of hGH. Bioconjug Chem 24:456–463. https://doi.org/10.1021/bc300594y
Damelin M et al (2017) A PTK7-targeted antibody-drug conjugate reduces tumor-initiating cells and induces sustained tumor regressions. Sci Transl Med 9:2611. https://doi.org/10.1126/scitranslmed.aag2611
Davis FF (2002) The origin of PEGnology. Adv Drug Deliv Rev 54:457–458. https://doi.org/10.1016/S0169-409X(02)00021-2
Dennler P et al (2014) Transglutaminase-based chemo-enzymatic conjugation approach yields homogeneous antibody-drug conjugates. Bioconjug Chem 25:569–578. https://doi.org/10.1021/bc400574z
Derakhshanfar S, Mbeleck R, Xu K, Zhang X, Zhong W, Xing M (2018) 3D bioprinting for biomedical devices and tissue engineering: a review of recent trends and advances. Bioact Mater 3:144–156. https://doi.org/10.1016/j.bioactmat.2017.11.008
Desai KGH, Park HJ (2005) Recent developments in microencapsulation of food ingredients. Drying Technol 23:1361–1394. https://doi.org/10.1081/drt-200063478
Domagała J, Najgebauer-Lejko D, Wieteska-Śliwa I, Sady M, Wszołek M, Bonczar G, Filipczak-Fiutak M (2016) Influence of milk protein cross-linking by transglutaminase on the rennet coagulation time and the gel properties. J Sci Food Agric 96:3500–3507. https://doi.org/10.1002/jsfa.7534
Dondero M, Figueroa V, Morales X, Curotto E (2006) Transglutaminase effects on gelation capacity of thermally induced beef protein gels. Food Chem 99:546–554. https://doi.org/10.1016/j.foodchem.2005.08.022
Dozier JK, Distefano MD (2015) Site-specific PEGylation of therapeutic proteins. Int J Mol Sci 16:25831–25864. https://doi.org/10.3390/ijms161025831
Du G, Cui L, Zhu Y, Chen J (2007) Improvement of shrink-resistance and tensile strength of wool fabric treated with a novel microbial transglutaminase from Streptomyces hygroscopicus. Enzyme Microb Technol 40:1753–1757. https://doi.org/10.1016/j.enzmictec.2006.12.001
Duarte LS, Schöffer JdN, Lorenzoni ASG, Rodrigues RC, Rodrigues E, Hertz PF (2017) A new bioprocess for the production of prebiotic lactosucrose by an immobilized β-galactosidase. Process Biochem 55:96–103. https://doi.org/10.1016/j.procbio.2017.01.015
Duran R, Junqua M, Schmitter JM, Gancet C, Goulas P (1998) Purification, characterisation, and gene cloning of transglutaminase from Streptoverticillium cinnamoneum CBS 68368. Biochimie 80:313–319. https://doi.org/10.1016/S0300-9084(98)80073-4
El-Sherbiny IM, Yacoub MH (2013) Hydrogel scaffolds for tissue engineering: progress and challenges. Glob Cardiol Sci Pract 2013:316–342. https://doi.org/10.5339/gcsp.2013.38
Elzoghby AO, Samy WM, Elgindy NA (2012) Protein-based nanocarriers as promising drug and gene delivery systems. J Control Release 161:38–49. https://doi.org/10.1016/j.jconrel.2012.04.036
Færgemand M, Qvist KB (1997) Transglutaminase: effect on rheological properties, microstructure and permeability of set style acid skim milk gel. Food Hydrocoll 11:287–292. https://doi.org/10.1016/S0268-005X(97)80058-6
Fan Z, Zhang Y, Fang S, Xu C, Li X (2015) Bienzymatically crosslinked gelatin/hyaluronic acid interpenetrating network hydrogels: preparation and characterization. RSC Adv 5:1929–1936. https://doi.org/10.1039/C4RA12446D
Farias SE et al (2014) Mass spectrometric characterization of transglutaminase based site-specific antibody–drug conjugates. Bioconjug Chem 25:240–250. https://doi.org/10.1021/bc4003794
Fatima SW, Khare SK (2018) Current insight and futuristic vistas of microbial transglutaminase in nutraceutical industry. Microbiol Res 215:7–14. https://doi.org/10.1016/j.micres.2018.06.001
Feng J et al (2018) Effects of partial substitution of NaCl on gel properties of fish myofibrillar protein during heating treatment mediated by microbial transglutaminase. LWT 93:1–8. https://doi.org/10.1016/j.lwt.2018.03.018
Fernandez-Bats I, Di Pierro P, Villalonga-Santana R, Garcia-Almendarez B, Porta R (2018) Bioactive mesoporous silica nanocomposite films obtained from native and transglutaminase-crosslinked bitter vetch proteins. Food Hydrocoll 82:106–115. https://doi.org/10.1016/j.foodhyd.2018.03.041
Fontana A, Spolaore B, Mero A, Veronese FM (2008) Site-specific modification and PEGylation of pharmaceutical proteins mediated by transglutaminase. Adv Drug Deliv Rev 60:13–28. https://doi.org/10.1016/j.addr.2007.06.015
Gaffar Hossain KM, Juan AR, Tzanov T (2008) Simultaneous protease and transglutaminase treatment for shrink resistance of wool. Biocatal Biotransformation 26:405–411. https://doi.org/10.1080/10242420802364940
Garrigle MM, Mullen C, Haugh M, Voisin M, McNamara L (2016) Osteocyte differentiation and the formation of an interconnected cellular network in vitro. Eur Cell Mater 31:323–340. https://doi.org/10.22203/eCM
Gaspar ALC, de Góes-Favoni SP (2015) Action of microbial transglutaminase (MTGase) in the modification of food proteins: a review. Food Chem 171:315–322. https://doi.org/10.1016/j.foodchem.2014.09.019
Gerrard JA, Fayle SE, Wilson AJ, Newberry MP, Ross M, Kavale S (1998) Dough properties and crumb strength of white pan bread as affected by microbial transglutaminase. J Food Sci 63:472–475. https://doi.org/10.1111/j.1365-2621.1998.tb15766.x
Gottmann KD, Sprössler BD (1992) Baking agent or flour, and method of production of dough and bakery products. Publication number: EP0492406A1. Germany Patent
Griffith LG, Swartz MA (2006) Capturing complex 3D tissue physiology in vitro. Nat Rev Mol Cell Biol 7:211. https://doi.org/10.1038/nrm1858
Grigoletto A, Mero A, Zanusso I, Schiavon O, Pasut G (2015) Chemical and enzymatic site specific PEGylation of hGH: the stability and in vivo activity of PEG-N-terminal-hGH and PEG-Gln141-hGH conjugates. Macromol Biosci 16:50–56. https://doi.org/10.1002/mabi.201500282
Grigoletto A, Mero A, Yoshioka H, Schiavon O, Pasut G (2017) Covalent immobilisation of transglutaminase: stability and applications in protein PEGylation. J Drug Target 25:856–864. https://doi.org/10.1080/1061186X.2017.1363211
Grünberg J, Jeger S, Sarko D, Dennler P, Zimmermann K, Mier W, Schibli R (2013) DOTA-functionalized polylysine: a high number of DOTA chelates positively influences the biodistribution of enzymatic conjugated anti-tumor antibody chCE7agl. PLoS ONE 8:e60350. https://doi.org/10.1371/journal.pone.0060350
Gujral HS, Rosell CM (2004) Functionality of rice flour modified with a microbial transglutaminase. J Cereal Sci 39:225–230. https://doi.org/10.1016/j.jcs.2003.10.004
Harris JM, Chess RB (2003) Effect of pegylation on pharmaceuticals. Nat Rev Drug Discov 2:214. https://doi.org/10.1038/nrd1033
Hoffman AS (2016) The early days of PEG and PEGylation (1970s–1990s). Acta Biomater 40:1–5. https://doi.org/10.1016/j.actbio.2016.05.029
Hong GP, Chin KB (2010) Effects of microbial transglutaminase and sodium alginate on cold-set gelation of porcine myofibrillar protein with various salt levels. Food Hydrocoll 24:444–451. https://doi.org/10.1016/j.foodhyd.2009.11.011
Hong GP, Xiong YL (2012) Microbial transglutaminase-induced structural and rheological changes of cationic and anionic myofibrillar proteins. Meat Sci 91:36–42. https://doi.org/10.1016/j.meatsci.2011.12.002
Hong GP, Min SG, Chin KB (2012) Emulsion properties of pork myofibrillar protein in combination with microbial transglutaminase and calcium alginate under various pH conditions. Meat Sci 90:185–193. https://doi.org/10.1016/j.meatsci.2011.06.023
Hong PK, Ndagijimana M, Betti M (2016) Glucosamine-induced glycation of hydrolysed meat proteins in the presence or absence of transglutaminase: chemical modifications and taste-enhancing activity. Food Chem 197:1143–1152. https://doi.org/10.1016/j.foodchem.2015.11.096
Irvine SA et al (2015) Printing cell-laden gelatin constructs by free-form fabrication and enzymatic protein crosslinking. Biomed Microdevices 17:16. https://doi.org/10.1007/s10544-014-9915-8
Jeger S et al (2010) Site-specific and stoichiometric modification of antibodies by bacterial transglutaminase. Angew Chem Int Ed 49:9995–9997. https://doi.org/10.1002/anie.201004243
Jin I-H, Kim J-E, Seo J-H, Lee S-P (2013) Physicochemical properties of soy protein isolate gels emulsified with various oils using a microbial transglutaminase. Food Sci Biotechnol 22:129–136. https://doi.org/10.1007/s10068-013-0058-0
Jira W, Schwagele F (2017) A sensitive high performance liquid chromatography-tandem mass spectrometry method for the detection of microbial transglutaminase in different types of restructured meat. Food Chem 221:1970–1978. https://doi.org/10.1016/j.foodchem.2016.11.148
Junutula JR et al (2008) Site-specific conjugation of a cytotoxic drug to an antibody improves the therapeutic index. Nat Biotechnol 26:925. https://doi.org/10.1038/nbt.1480
Kamiya N, Takazawa T, Tanaka T, Ueda H, Nagamune T (2003) Site-specific cross-linking of functional proteins by transglutamination. Enzyme Microb Technol 33:492–496. https://doi.org/10.1016/S0141-0229(03)00154-6
Kato A, Wada T, Kobayashi K, Seguro K, Motoki M (1991) Ovomucin-food protein conjugates prepared through the transglutaminase reaction. Agric Biol Chem 55:1027–1031. https://doi.org/10.1271/bbb1961.55.1027
Khameneh B et al (2015) Preparation, characterization and molecular modeling of PEGylated human growth hormone with agonist activity. Int J Biol Macromol 80:400–409. https://doi.org/10.1016/j.ijbiomac.2015.06.037
Kieliszek M, Misiewicz A (2014) Microbial transglutaminase and its application in the food industry: a review. Folia Microbiol 59:241–250. https://doi.org/10.1007/s12223-013-0287-x
Kim Y, Kee JI, Lee S, Yoo S-H (2014) Quality improvement of rice noodle restructured with rice protein isolate and transglutaminase. Food Chem 145:409–416. https://doi.org/10.1016/j.foodchem.2013.08.078
Langer R, Vacanti JP (1993) Tissue engineering. Science 260:920. https://doi.org/10.1126/science.8493529
Lee EJ, Kasper FK, Mikos AG (2014) Biomaterials for tissue engineering. Ann Biomed Eng 42:323–337. https://doi.org/10.1007/s10439-013-0859-6
Lerner A, Matthias T (2015) Possible association between celiac disease and bacterial transglutaminase in food processing: a hypothesis. Nutr Rev 73:544–552. https://doi.org/10.1093/nutrit/nuv011
Lesiow T, Rentfrow GK, Xiong YL (2017) Polyphosphate and myofibrillar protein extract promote transglutaminase-mediated enhancements of rheological and textural properties of PSE pork meat batters. Meat Sci 128:40–46. https://doi.org/10.1016/j.meatsci.2017.02.002
Lhospice F et al (2015) Site-specific conjugation of monomethyl auristatin e to anti-CD30 antibodies improves their pharmacokinetics and therapeutic Index in rodent models. Mol Pharm 12:1863–1871. https://doi.org/10.1021/mp500666j
Li C, Wang C-L, Sun Y, Li A-L, Liu F, Meng X-C (2016) Microencapsulation of Lactobacillus rhamnosus GG by transglutaminase cross-linked soy protein isolate to improve survival in simulated gastrointestinal conditions and yoghurt. J Food Sci 81:M1726–M1734. https://doi.org/10.1111/1750-3841.13337
Li T, Li C, Quan DN, Bentley WE, Wang L-X (2018) Site-specific immobilization of endoglycosidases for streamlined chemoenzymatic glycan remodeling of antibodies. Carbohydr Res 458–459:77–84. https://doi.org/10.1016/j.carres.2018.02.007
Liu Q, Liu L, Li J, Zhang D, Sun J, Du G, Chen J (2011) Influence of microbial transglutaminase modified gelatin-sodium caseinate, as a filler, on the subjective mechanical and structural properties of leather. J Am Leather Chem Assoc 106:200–207. https://doi.org/10.1016/S0268-005X(01)00025-X
Liu L et al (2018) Preparation and characterization of chitosan–collagen peptide/oxidized konjac glucomannan hydrogel. Int J Biol Macromol 108:376–382. https://doi.org/10.1016/j.ijbiomac.2017.11.128
Mariniello L, Porta R, Sorrentino A, Giosafatto CV, Rossi Marquez G, Esposito M, Di Pierro P (2014) Transglutaminase-mediated macromolecular assembly: production of conjugates for food and pharmaceutical applications. Amino Acids 46:767–776. https://doi.org/10.1007/s00726-013-1561-6
Mateo C, Palomo JM, Fernandez-Lorente G, Guisan JM, Fernandez-Lafuente R (2007) Improvement of enzyme activity, stability and selectivity via immobilization techniques. Enzyme Microb Technol 40:1451–1463. https://doi.org/10.1016/j.enzmictec.2007.01.018
Matte CR, Bussamara R, Dupont J, Rodrigues RC, Hertz PF, Ayub MAZ (2014) Immobilization of Thermomyces lanuginosus lipase by different techniques on Immobead 150 support: characterization and applications. Appl Biochem Biotechnol 172:2507–2520. https://doi.org/10.1007/s12010-013-0702-4
Matthias T, Jeremias P, Neidhöfer S, Lerner A (2016) The industrial food additive, microbial transglutaminase, mimics tissue transglutaminase and is immunogenic in celiac disease patients. Autoimmun Rev 15:1111–1119. https://doi.org/10.1016/j.autrev.2016.09.011
Maullu C et al (2009) Site-directed enzymatic PEGylation of the human granulocyte colony-stimulating factor. FEBS J 276:6741–6750. https://doi.org/10.1111/j.1742-4658.2009.07387.x
Mazzeo MF, Bonavita R, Maurano F, Bergamo P, Siciliano RA, Rossi M (2013) Biochemical modifications of gliadins induced by microbial transglutaminase on wheat flour. Biochim Biophys Acta 1830:5166–5174. https://doi.org/10.1016/j.bbagen.2013.07.021
McDermott MK, Chen T, Williams CM, Markley KM, Payne GF (2004) Mechanical properties of biomimetic tissue adhesive based on the microbial transglutaminase-catalyzed crosslinking of gelatin. Biomacromol 5:1270–1279. https://doi.org/10.1021/bm034529a
Mero A, Spolaore B, Veronese FM, Fontana A (2009) Transglutaminase-mediated PEGylation of proteins: direct identification of the sites of protein modification by mass spectrometry using a novel monodisperse PEG. Bioconjug Chem 20:384–389. https://doi.org/10.1021/bc800427n
Mihalcea L et al (2018) Transglutaminase mediated microencapsulation of sea buckthorn supercritical CO2 extract in whey protein isolate and valorization in highly value added food products. Food Chem 262:30–38. https://doi.org/10.1016/j.foodchem.2018.04.067
Milczek EM (2018) Commercial applications for enzyme-mediated protein conjugation: new developments in enzymatic processes to deliver functionalized proteins on the commercial scale. Chem Rev 118:119–141. https://doi.org/10.1021/acs.chemrev.6b00832
Moghaddas Kia E, Alizadeh M, Esmaiili M (2018) Development and characterization of probiotic UF Feta cheese containing Lactobacillus paracasei microencapsulated by enzyme based gelation method. J Food Sci Technol 55:3657–3664. https://doi.org/10.1007/s13197-018-3294-8
Mojsov K (2017) Enzymatic treatment of wool fabrics: opportunity of the improvement on some physical and chemical properties of the fabrics. J Text Inst 108:1136–1143. https://doi.org/10.1080/00405000.2016.1222856
Monteiro ML, Marsico ET, Lazaro CA, da Silva Canto AC, Lima BR, da Cruz AG, Conte-Junior CA (2015) Effect of transglutaminase on quality characteristics of a value-added product tilapia wastes. J Food Sci Technol 52:2598–2609. https://doi.org/10.1007/s13197-014-1327-5
Motoki M, Seguro K (1998) Transglutaminase and its use for food processing. Trends Food Sci Technol 9:204–210. https://doi.org/10.1016/S0924-2244(98)00038-7
Nawong S, Oonsivilai R, Boonkerd N, Truelstrup Hansen L (2016) Entrapment in food-grade transglutaminase cross-linked gelatin–maltodextrin microspheres protects Lactobacillus spp. during exposure to simulated gastro-intestinal juices. Food Res Int 85:191–199. https://doi.org/10.1016/j.foodres.2016.04.041
Nejadmoghaddam M-R, Minai-Tehrani A, Ghahremanzadeh R, Mahmoudi M, Dinarvand R, Zarnani A-H (2019) Antibody-drug conjugates: possibilities and challenges. Avicenna J Med Biotechnol 11:3–23
O'Brien FJ (2011) Biomaterials & scaffolds for tissue engineering. Mater Today 14:88–95. https://doi.org/10.1016/S1369-7021(11)70058-X
Ohtsuka T, Sawa A, Kawabata R, Nio N, Motoki M (2000) Substrate specificities of microbial transglutaminase for primary amines. J Agric Food Chem 48:6230–6233. https://doi.org/10.1021/jf000302k
Okeley NM, Toki BE, Zhang X, Jeffrey SC, Burke PJ, Alley SC, Senter PD (2013) Metabolic engineering of monoclonal antibody carbohydrates for antibody–drug conjugation. Bioconjug Chem 24:1650–1655. https://doi.org/10.1021/bc4002695
Oner Z, Karahan A, Aydemir S, Sanlidere Aloglu H (2008) Effect of ransglutaminase on physicochemical properties of set-style yogurt. Int J Food Prop 11:196–205. https://doi.org/10.1080/10942910701286346
Pasut G, Veronese FM (2012) State of the art in PEGylation: the great versatility achieved after forty years of research. J Control Release 161:461–472. https://doi.org/10.1016/j.jconrel.2011.10.037
Polak DJ (2010) Regenerative medicine. Opportunities and challenges: a brief overview. J R Soc Interface 7:S777–S781. https://doi.org/10.1098/rsif.2010.0362.focus
Porta R, Di Pierro P, Sabbah M, Regalado-Gonzales C, Mariniello L, Kadivar M, Arabestani A (2016) Blend films of pectin and bitter vetch (Vicia ervilia) proteins: properties and effect of transglutaminase. Innov Food Sci Emerg Technol 36:245–251. https://doi.org/10.1016/j.ifset.2016.07.001
Prakasan V, Chawla SP, Sharma A (2015) Effect of transglutaminase treatment on functional properties of paneer. Int J Curr Microbiol Appl Sci 4:227–238
Qin XS, Luo SZ, Cai J, Zhong XY, Jiang ST, Zhao YY, Zheng Z (2016) Transglutaminase-induced gelation properties of soy protein isolate and wheat gluten mixtures with high intensity ultrasonic pretreatment. Ultrason Sonochem 31:590–597. https://doi.org/10.1016/j.ultsonch.2016.02.010
Roberts MJ, Bentley MD, Harris JM (2002) Chemistry for peptide and protein PEGylation. Adv Drug Deliv Rev 54:459–476. https://doi.org/10.1016/S0169-409X(02)00022-4
Rodrigues RC, Ortiz C, Berenguer-Murcia Á, Torres R, Fernández-Lafuente R (2013) Modifying enzyme activity and selectivity by immobilization. Chem Soc Rev 42:6290–6307. https://doi.org/10.1039/C2CS35231A
Rodriguez-Nogales JM (2006) Enhancement of transglutaminase-induced protein cross-linking by preheat treatment of cows’ milk: a statistical approach. Int Dairy J 16:26–32. https://doi.org/10.1016/j.idairyj.2005.01.003
Romeih E, Walker G (2017) Recent advances on microbial transglutaminase and dairy application. Trends Food Sci Technol 62:133–140. https://doi.org/10.1016/j.tifs.2017.02.015
Romero de Ávila MD, Ordóñez JA, de la Hoz L, Herrero AM, Cambero MI (2010) Microbial transglutaminase for cold-set binding of unsalted/salted pork models and restructured dry ham. Meat Sci 84:747–754. https://doi.org/10.1016/j.meatsci.2009.11.011
Rossa PN, de Sá EMF, Burin VM, Bordignon-Luiz MT (2011) Optimization of microbial transglutaminase activity in ice cream using response surface methodology. LWT: Food Sci Technol 44:29–34. https://doi.org/10.1016/j.lwt.2010.06.013
Rossi Marquez G, Di Pierro P, Mariniello L, Esposito M, Giosafatto CVL, Porta R (2017) Fresh-cut fruit and vegetable coatings by transglutaminase-crosslinked whey protein/pectin edible films. LWT 75:124–130. https://doi.org/10.1016/j.lwt.2016.08.017
Sachdev JC et al (2016) A phase 1 study of PF-06647020, an antibody-drug conjugate (ADC) targeting protein tyrosine kinase 7 (PTK7), in patients with advanced solid tumors including platinum resistant ovarian cancer (OVCA). Ann Oncol. https://doi.org/10.1093/annonc/mdw435.29
Sakamoto H, Yamazaki K, Kaga C, Yamamoto Y, Ito R, Kurosawa Y (1996) Strength enhancement by addition of microbial transglutaminase during Chinese noodle processing. J Jpn Soc Food Sci 43:598–602. https://doi.org/10.3136/nskkk.43.598
Salma HA, Nahid AA, ElShazali AM, Isam AMA, Elfadil EB (2010) Changes in the functional properties as a function of NaCl concentration of legumes protein isolate by transglutaminase cross linking. Food Res Int 17:817–824
Sanchez C, Renard D (2002) Stability and structure of protein-polysaccharide coacervates in the presence of protein aggregates. Int J Pharm 242:319–324. https://doi.org/10.1016/S0378-5173(02)00174-6
Santhi D, Kalaikannan A, Malairaj P, Arun Prabhu S (2017) Application of microbial transglutaminase in meat foods: a review. Crit Rev Food Sci Nutr 57:2071–2076. https://doi.org/10.1080/10408398.2014.945990
Scaramuzza S, Tonon G, Olianas A, Messana I, Schrepfer R, Orsini G, Caliceti P (2012) A new site-specific monoPEGylated filgrastim derivative prepared by enzymatic conjugation: production and physicochemical characterization. J Control Release 164:355–363. https://doi.org/10.1016/j.jconrel.2012.06.026
Scarnato L, Montanari C, Serrazanetti DI, Aloisi I, Balestra F, Del Duca S, Lanciotti R (2017) New bread formulation with improved rheological properties and longer shelf-life by the combined use of transglutaminase and sourdough. LWT: Food Sci Technol 81:101–110. https://doi.org/10.1016/j.lwt.2017.03.042
Selis F et al (2012) Enzymatic mono-pegylation of glucagon-like peptide 1 towards long lasting treatment of type 2 diabetes. Res Pharm Sci 2:58–65. https://doi.org/10.1016/j.rinphs.2012.09.001
Sharma R, Zakora M, Qvist KB (2002) Susceptibility of an industrial α-lactalbumin concentrate to cross-linking by microbial transglutaminase. Int Dairy J 12:1005–1012. https://doi.org/10.1016/S0958-6946(02)00122-X
Sharma A, Brand D, Fairbank J, Ye H, Lavy C, Czernuszka J (2017) A self-organising biomimetic collagen/nano-hydroxyapatite-glycosaminoglycan scaffold for spinal fusion. J Mater Sci 52:12574–12592. https://doi.org/10.1007/s10853-017-1229-9
Shen B-Q et al (2012) Conjugation site modulates the in vivo stability and therapeutic activity of antibody-drug conjugates. Nat Biotechnol 30:184. https://doi.org/10.1038/nbt.2108
Suzuki S, Izawa Y, Kobayashi K, Eto Y, Yamanaka S, Kubota K, Yokozeki K (2000) Purification and characterization of novel transglutaminase from Bacillus subtilis spores. Biosci Biotechnol Biochem 64:2344–2351. https://doi.org/10.1271/bbb.64.2344
Siegmund V et al (2015) Locked by design: a conformationally constrained transglutaminase tag enables efficient site-specific conjugation. Angew Chem Int Ed 54:13420–13424. https://doi.org/10.1002/anie.201504851
Song F, Zhang LM (2008) Enzyme-catalyzed formation and structure characteristics of a protein-based hydrogel. J Phys Chem B 112:13749–13755. https://doi.org/10.1021/jp8041389
Song CL, Zhao XH (2014) Structure and property modification of an oligochitosan-glycosylated and crosslinked soybean protein generated by microbial transglutaminase. Food Chem 163:114–119. https://doi.org/10.1016/j.foodchem.2014.04.089
Sorapukdee S, Tangwatcharin P (2018) Quality of steak restructured from beef trimmings containing microbial transglutaminase and impacted by freezing and grading by fat level. Asian-Australas J Anim Sci 31:129–137. https://doi.org/10.5713/ajas.17.0170
Spidel JL, Vaessen B, Albone EF, Cheng X, Verdi A, Kline JB (2017) Site-specific conjugation to native and engineered lysines in human immunoglobulins by microbial transglutaminase. Bioconjug Chem 28:2471–2484. https://doi.org/10.1021/acs.bioconjchem.7b00439
Stachel I, Schwarzenbolz U, Henle T, Meyer M (2010) Cross-linking of type I collagen with microbial transglutaminase: identification of cross-linking sites. Biomacromol 11:698–705. https://doi.org/10.1021/bm901284x
Strop P (2014) Versatility of microbial transglutaminase. Bioconjug Chem 25:855–862. https://doi.org/10.1021/bc500099v
Strop P et al (2013) Location matters: site of conjugation modulates stability and pharmacokinetics of antibody drug conjugates. Chem Biol 20:161–167. https://doi.org/10.1016/j.chembiol.2013.01.010
Strop P et al (2016) RN927C, a site-specific Trop-2 antibody–drug conjugate (ADC) with enhanced stability, is highly efficacious in preclinical solid tumor models. Mol Cancer Ther 15:2698–2708. https://doi.org/10.1158/1535-7163.MCT-16-0431
Synowiecki J, Wołosowska S (2006) Immobilization of thermostable β-glucosidase from Sulfolobus shibatae by cross-linking with transglutaminase. Enzyme Microb Technol 39:1417–1422. https://doi.org/10.1016/j.enzmictec.2006.03.028
Taghi Gharibzahedi SM, Koubaa M, Barba FJ, Greiner R, George S, Roohinejad S (2018) Recent advances in the application of microbial transglutaminase crosslinking in cheese and ice cream products: a review. Int J Biol Macromol 107:2364–2374. https://doi.org/10.1016/j.ijbiomac.2017.10.115
Tammatinna A, Benjakul S, Visessanguan W, Tanaka M (2007) Gelling properties of white shrimp (Penaeus vannamei) meat as influenced by setting condition and microbial transglutaminase. LWT: Food Sci Technol 40:1489–1497. https://doi.org/10.1016/j.lwt.2006.11.017
Tang C-H, Jiang Y, Wen Q-B, Yang X-Q (2005) Effect of transglutaminase treatment on the properties of cast films of soy protein isolates. J Biotechnol 120:296–307. https://doi.org/10.1016/j.jbiotec.2005.06.020
Tappa K, Jammalamadaka U (2018) Novel biomaterials used in medical 3D printing techniques. J Funct Biomater 9:17. https://doi.org/10.3390/jfb9010017
Taylor MM, Bumanlag L, Marmer WN, Brown E (2006) Use of enzymatically modified gelatin and casein as fillers in leather processing. J Am Leather Chem Assoc 101:169–178
Tello F, Prata AS, Rodrigues RAF, Sartoratto A, Grosso CRF (2016) Improving the performance of transglutaminase-crosslinked microparticles for enteric delivery. Food Res Int 88:153–158. https://doi.org/10.1016/j.foodres.2016.02.020
Tesfaw A, Assefa F (2014) Applications of transglutaminase in textile, wool, and leather processin. Int J Text Sci 3:64–69. https://doi.org/10.5923/j.textile.20140304.02
Ting-Jin L, Azhar-Mat E, Abdul-Alias K, Rajeev B, Min-Tze L (2011) Development of soy-based cream cheese via the addition of microbial transglutaminase, soy protein isolate and maltodextrin. Br Food J 113:1147–1172. https://doi.org/10.1108/00070701111174587
Toh WS, Loh XJ (2014) Advances in hydrogel delivery systems for tissue regeneration. Mater Sci Eng C 45:690–697. https://doi.org/10.1016/j.msec.2014.04.026
Tominaga J, Kamiya N, Doi S, Ichinose H, Goto M (2004) An enzymatic strategy for site-specific immobilization of functional proteins using microbial transglutaminase. Enzyme Microb Technol 35:613–618. https://doi.org/10.1016/j.enzmictec.2004.08.014
Treib J, Baron JF, Grauer MT, Strauss RG (1999) An international view of hydroxyethyl starches. Intensive Care Med 25:258–268. https://doi.org/10.1007/s001340050833
Uran H, Yilmaz S (2018) A research on determination of quality characteristics of chicken burgers produced with transglutaminase supplementation. J Food Sci Technol 38:19–25. https://doi.org/10.1590/1678-457X.33816
Wang Q, Jin G, Fan X, Zhao X, Cui L, Wang P (2010) Antibacterial functionalization of wool via mTGase-catalyzed grafting of ε-poly-l-lysine. Appl Biochem Biotechnol 160:2486–2497. https://doi.org/10.1007/s12010-009-8708-7
Wang F, Huang W, Kim Y, Liu R, Tilley M (2011) Effects of transglutaminase on the rheological and noodle-making characteristics of oat dough containing vital wheat gluten or egg albumin. J Cereal Sci 54:53–59. https://doi.org/10.1016/j.jcs.2011.02.010
Wang B, Adhikari B, Barrow CJ (2014) Optimisation of the microencapsulation of tuna oil in gelatin–sodium hexametaphosphate using complex coacervation. Food Chem 158:358–365. https://doi.org/10.1016/j.foodchem.2014.02.135
Wang J-H, Tang M-Z, Yu X-T, Xu C-M, Yang H-M, Tang J-B (2019) Site-specific, covalent immobilization of an engineered enterokinase onto magnetic nanoparticles through transglutaminase-catalyzed bioconjugation. Colloids Surf B 177:506–511. https://doi.org/10.1016/j.colsurfb.2019.02.018
Wen-qiong W, Lan-wei Z, Xue H, Yi L (2017) Cheese whey protein recovery by ultrafiltration through transglutaminase (TG) catalysis whey protein cross-linking. Food Chem 215:31–40. https://doi.org/10.1016/j.foodchem.2016.07.057
Woods K, Thigpen C, Wang JP, Park H, Hielscher A (2017) Mechanically tuned 3 dimensional hydrogels support human mammary fibroblast growth and viability. BMC Cell Biol 18:35. https://doi.org/10.1186/s12860-017-0151-y
Wu M, He Q, Hong Y, Wang S (2016) Preheating of kidney bean proteins enhances cross-linking and functional properties with chicken myofibrillar proteins induced by transglutaminase. LWT: Food Sci Technol 65:816–822. https://doi.org/10.1016/j.lwt.2015.09.019
Xiao H, Chatterjee A, Choi S-h, Bajjuri KM, Sinha SC, Schultz PG (2013) Genetic incorporation of multiple unnatural amino acids into proteins in mammalian cells. Angew Chem Int Ed 52:14080–14083. https://doi.org/10.1002/anie.201308137
Xing G, Rui X, Jiang M, Xiao Y, Guan Y, Wang D, Dong M (2016) In vitro gastrointestinal digestion study of a novel bio-tofu with special emphasis on the impact of microbial transglutaminase. PeerJ 4:e2754. https://doi.org/10.7717/peerj.2754
Yang G, Xiao Z, Ren X, Long H, Qian H, Ma K, Guo Y (2016) Enzymatically crosslinked gelatin hydrogel promotes the proliferation of adipose tissue-derived stromal cells. PeerJ 4:e2497. https://doi.org/10.7717/peerj.2497
Yang G, Xiao Z, Long H, Ma K, Zhang J, Ren X, Zhang J (2018) Assessment of the characteristics and biocompatibility of gelatin sponge scaffolds prepared by various crosslinking methods. Sci Rep 8:1616. https://doi.org/10.1038/s41598-018-20006-y
Yao H, Jiang F, Lu A, Zhang G (2016) Methods to design and synthesize antibody-drug conjugates (ADCs). Int J Mol Sci 17:194. https://doi.org/10.3390/ijms17020194
Yokoyama K, Nio N, Kikuchi Y (2004) Properties and applications of microbial transglutaminase. Appl Microbiol Biotechnol 64:447–454. https://doi.org/10.1007/s00253-003-1539-5
Yuan F, Lv L, Li Z, Mi N, Chen H, Lin H (2017) Effect of transglutaminase-catalyzed glycosylation on the allergenicity and conformational structure of shrimp (Metapenaeus ensis) tropomyosin. Food Chem 219:215–222. https://doi.org/10.1016/j.foodchem.2016.09.139
Zhao X, Shaw AC, Wang J, Chang CC, Deng J, Su J (2010) A novel high-throughput screening method for microbial transglutaminases with high specificity toward Gln141 of human growth hormone. J Biomol Screen 15:206–212. https://doi.org/10.1177/1087057109356206
Zhao L et al (2016) A novel smart injectable hydrogel prepared by microbial transglutaminase and human-like collagen: its characterization and biocompatibility. Mater Sci Eng C 68:317–326. https://doi.org/10.1016/j.msec.2016.05.108
Zhou JQ, He T, Wang JW (2016a) The microbial transglutaminase immobilization on carboxylated poly(N-isopropylacrylamide) for thermo-responsivity. Enzyme Microb Technol 87–88:44–51. https://doi.org/10.1016/j.enzmictec.2016.02.012
Zhou JQ, He T, Wang JW (2016b) PEGylation of cytochrome c at the level of lysine residues mediated by a microbial transglutaminase. Biotechnol Lett 38:1121–1129. https://doi.org/10.1007/s10529-016-2083-6
Zhou L et al (2017a) Abrogation of immunogenic properties of gliadin peptides through transamidation by microbial transglutaminase is acyl-acceptor dependent. J Agric Food Chem 65:7542–7552. https://doi.org/10.1021/acs.jafc.7b02557
Zhou L et al (2017) Blocking celiac antigenicity of the glutamine-rich gliadin 33-mer peptide by microbial transglutaminase. RSC Adv 7:14438–14447. https://doi.org/10.1039/C6RA27893K
Zhu Y, Tramper J (2008) Novel applications for microbial transglutaminase beyond food processing. Trends Biotechnol 26:559–565. https://doi.org/10.1016/j.tibtech.2008.06.006
Zhu C et al (2018) Novel enzymatic crosslinked hydrogels that mimic extracellular matrix for skin wound healing. J Mater Sci 53:5909–5928. https://doi.org/10.1007/s10853-017-1956-y
Zou Q, Liu X, Zhao J, Tian F, Zhang H-p, Zhang H, Chen W (2012) Microencapsulation of Bifidobacterium bifidum F-35 in whey protein-based microcapsules by transglutaminase-induced gelation. J Food Sci 77:M270–M277. https://doi.org/10.1111/j.1750-3841.2012.02673.x
Zuberbühler K, Casi G, Bernardes GJL, Neri D (2012) Fucose-specific conjugation of hydrazide derivatives to a vascular-targeting monoclonal antibody in IgG format. Chem Commun 48:7100–7102. https://doi.org/10.1039/C2CC32412A
Acknowledgements
The authors wish to thank Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Coordenação do Aperfeiçoamento de Pessoal do Ensino Superior (CAPES), Finance Code 001, and Fundação de Apoio à Pesquisa do Estado do Rio Grande do Sul (FAPERGS) for their financial support of this project and scholarships.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
All authors of this research declare to have no conflicts of interest.
Ethical approval
Research was conducted without using human or animal experimentation.
Informed consent
No informed consent was necessary to conduct this research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Duarte, L., Matte, C.R., Bizarro, C.V. et al. Review transglutaminases: part II—industrial applications in food, biotechnology, textiles and leather products. World J Microbiol Biotechnol 36, 11 (2020). https://doi.org/10.1007/s11274-019-2792-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-019-2792-9