Abstract
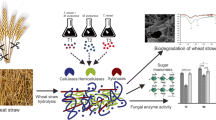
Lignocellulosic plant biomass is the world’s most abundant carbon source and has consequently attracted attention as a renewable resource for production of biofuels and commodity chemicals that could replace fossil resources. Due to its recalcitrant nature, it must be pretreated by chemical, physical or biological means prior to hydrolysis, introducing additional costs. In this paper, we tested the hypothesis that fungi which thrive on lignocellulosic material (straw, bark or soil) would be efficient in degrading untreated lignocellulose. Wheat straw was used as a model. We developed a fast and simple screening method for cellulase producers and tested one hundred Trichoderma strains isolated from wheat straw. The most potent strain—UB483FTG2/ TUCIM 4455, was isolated from substrate used for mushroom cultivation and was identified as T. guizhouense. After optimization of growth medium, high cellulase activity was already achieved after 72 h of fermentation on raw wheat straw, while the model cellulase overproducing strain T. reesei QM 9414 took 170 h and reached only 45% of the cellulase activity secreted by T. guizhouense. Maximum production levels were 1.1 U/mL (measured with CMC as cellulase substrate) and 0.7 U/mL (β-glucosidase assay). The T. guizhouense cellulase cocktail hydrolyzed raw wheat straw within 35 h. Our study shows that screening for fungi that successfully compete for special substrates in nature will lead to the isolation of strains with qualitatively and quantitatively superior enzymes needed for their digestion which could be used for industrial purposes.



Similar content being viewed by others
References
Barakat A, Chuetor S, Monlau F, Solhy A, Rouau X (2014) Eco-friendly dry chemo-mechanical pretreatments of lignocellulosic biomass: impact on energy and yield of the enzymatic hydrolysis. Appl Energy 113:97–105. https://doi.org/10.1016/j.apenergy.2013.07.015
Baroncelli R et al (2015) Draft whole-genome sequence of the biocontrol agent Trichoderma harzianum T6776. Genome Announc 3:e00647–e001615. https://doi.org/10.1128/genomeA.00647-15
Benoliel B, Torres GA, de Moraes LMP (2013) A novel promising Trichoderma harzianum strain for the production of a cellulolytic complex using sugarcane bagasse in natura. Springer Plus 2:1–7
Biely P, Markovič O, Mislovičová D (1985) Sensitive detection of endo-1,4-β-glucanases and endo-1,4-β-xylanases in gels. Anal Biochem 144:147–151. https://doi.org/10.1016/0003-2697(85)90096-X
Bischof R, Fourtis L, Limbeck A, Gamauf C, Seiboth B, Kubicek CP (2013) Comparative analysis of the Trichoderma reesei transcriptome during growth on the cellulase inducing substrates wheat straw and lactose. Biotechnol Biofuels 6:127–141. https://doi.org/10.1186/1754-6834-6-127
Chaverri P, Branco-Rocha F, Jaklitsch W, Gazis R, Degenkolb T, Samuels GJ (2015) Systematics of the Trichoderma harzianum species complex and the re-identification of commercial biocontrol strains. Mycologia 107:558–590. https://doi.org/10.3852/14-147
Cherubini F (2010) The biorefinery concept: using biomass instead of oil for producing energy and chemicals. Energy Convers Manag 51:1412–1421. https://doi.org/10.1016/j.enconman.2010.01.015
Day JP, Shattock RC (1997) Aggressiveness and other factors relating to displacements of population of Phytophthora infestans in England and Wales. Eur J Plant Pathol 103:379–391
Deacon JW (2006) Environmental conditions for growth, and tolerance of extremes. In: Deacon JW (ed) Fungal biology, 4th edn. Blackwell Publishing, Oxford, p 16
Delabona Pda S, Farinas CS, da Silva MR, Azzoni SF, Pradella JG (2012) Use of a new Trichoderma harzianum strain isolated from the Amazon rainforest with pretreated sugar cane bagasse for on-site cellulase production. Bioresour Technol 107:517–521. https://doi.org/10.1016/j.biortech.2011.12.048
Deshpande V, Eriksson KE, Pettersson B (1978) Production, purification and partial characterization of 1,4-beta-glucosidase enzymes from Sporotrichum pulverulentum. Eur J Biochem 90:191–198
Dojnov B, Grujic M, Vujcic Z (2015a) Highly efficient production of Aspergillus niger amylase cocktail by solid state fermentation using triticale grains as a well-balanced substrate. J Serb Chem Soc. https://doi.org/10.2298/jsc150317041d
Dojnov B, Grujic M, Vujcic Z (2015b) Reliable simultaneous zymographic method of characterization of cellulolytic enzymes from fungal cellulase complex. Electrophoresis 36:1724–1727. https://doi.org/10.1002/elps.201400541
Druzhinina IS, Kopchinskiy AG, Komon M, Bissett J, Szakacs G, Kubicek CP (2005) An oligonucleotide barcode for species identification in Trichoderma and Hypocrea fungal. Genet Biol 42:813–828. https://doi.org/10.1016/j.fgb.2005.06.007
Druzhinina IS, Kubicek CP, Komoń-Zelazowska M, Mulaw TB, Bissett J (2010) The Trichoderma harzianum demon: complex speciation history resulting in coexistence of hypothetical biological species, recent agamospecies and numerous relict lineages. BMC Evolut Biol 10:14. https://doi.org/10.1186/1471-2148-10-94
Druzhinina IS et al (2011) Trichoderma: the genomics of opportunistic success. Nat Rev Microbiol 9:749–759. https://doi.org/10.1038/nrmicro2637
Druzhinina IS, Chenthamara K, Zhang J, Atanasova L, Yang D, Miao Y, Rahimi MJ (2018) Massive lateral transfer of genes encoding plant cell wall-degrading enzymes to the mycoparasitic fungus Trichoderma from its plant-associated hosts. PLoS Genet 14:33. https://doi.org/10.1371/journal.pgen.1007322
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Ghose TG (1987) Measurement of cellulase activities. Pure Appl Chem 59:11
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224. https://doi.org/10.1093/molbev/msp259
Grujić M, Dojnov B, Potočnik I, Duduk B, Vujčić Z (2015) Spent mushroom compost as substrate for the production of industrially important hydrolytic enzymes by fungi Trichoderma spp. and Aspergillus niger in solid state fermentation. Int Biodeterior Biodegrad 104:290–298. https://doi.org/10.1016/j.ibiod.2015.04.029
Kahr H, Wimberger J, Schürz D, Jäger A (2013) Evaluation of the biomass potential for the production of lignocellulosic bioethanol from various agricultural residues in Austria and worldwide. Energy Procedia 40:146–155. https://doi.org/10.1016/j.egypro.2013.08.018
Kandari V, Vajpayee I, Kumar D, Gupta S (2013) Cellulase and β-glucosidase production by Trichoderma viride and Aspergillus wentiiin sub-merged fermentation utilizing pretreated lignocellulosic biomass. J Microbil Biotechnol Res 3:63–78
Khan TS, Mubeen U (2012) Wheat straw: a pragmatic. Overv Curr Res J Biol Sci 4:2
Kim S, Dale BE (2004) Global potential bioethanol production from wasted crops and crop residues. Biomass Bioenergy 26:361–375. https://doi.org/10.1016/j.biombioe.2003.08.002
Kopchinskiy A, Komon M, Kubicek CP, Druzhinina IS (2005) TrichoBLAST: a multilocus database for Trichoderma and Hypocrea identifications. Mycol Res 109:658–660
Kosanović D, Potočnik I, Duduk B, Vukojević J, Stajić M, Rekanović E, Milijašević-Marčić S (2013) Trichoderma species on Agaricus bisporus farms in Serbia and their biocontrol. Ann Appl Biol 163:218–230. https://doi.org/10.1111/aab.12048
Kubicek CP (1981) Release of carboxymethyl-cellulase and β-glucosidase from cell walls of Trichoderma reesei. Eur J Appl Microbiol Biotechnol 13:226–231
Kubicek CP, Kubicek EM (2016) Enzymatic deconstruction of plant biomass by fungal enzymes. Curr Opin Chem Biol 35:51–57. https://doi.org/10.1016/j.cbpa.2016.08.028
Kumar D, Murthy GS (2013) Stochastic molecular model of enzymatic hydrolysis of cellulose for ethanol production. Biotechnol Biofuels 6:1–20. https://doi.org/10.1186/1754-6834-6-63
Lee J-S, Saddler J, Binod P (2016) Pretreatment of biomass. Bioresour Technol 199:1. https://doi.org/10.1016/j.biortech.2015.10.092
Li XL, Spanikova S, de Vries RP, Biely P (2007) Identification of genes encoding microbial glucuronoyl esterases. Febs Lett 581:4029–4035. https://doi.org/10.1016/j.febslet.2007.07.041
Maeda RN et al (2011) Enzymatic hydrolysis of pretreated sugar cane bagasse using Penicillium funiculosum and Trichoderma harzianum cellulases. Process Biochem 46:1196–1201. https://doi.org/10.1016/j.procbio.2011.01.022
Maheswari DK, Jahan H, Paul J, Varma A (1993) Wheat straw, a potential substrate for cellulase production using Trichoderma reesei. World J Microbiol Biotechnol 9:120–122
Mandels M (1975) Microbial sources of cellulase Biotechnology and Bioengineering symposium, pp 81–105
Martinez D et al (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nature Biotechnol 26:553–560. https://doi.org/10.1038/nbt1403
Meddeb-Mouelhi F, Moisan JK, Beauregard M (2014) A comparison of plate assay methods for detecting extracellular cellulase and xylanase activity. Enzyme Microb Technol 66:16–19. https://doi.org/10.1016/j.enzmictec.2014.07.004
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Pathak P, Bhardwaj NK, Singh AK (2014) Production of crude cellulase and xylanase from Trichoderma harzianum PPDDN10 NFCCI-2925 and its application in photocopier waste paper recycling. Appl Biochem Biotechnol 172:3776–3797. https://doi.org/10.1007/s12010-014-0758-9
Rabosky DL et al (2014) BAMMtools: an R package for the analysis of evolutionary dynamics on phylogenetic trees. Methods Ecol Evol 5:701–707. https://doi.org/10.1111/2041-210x.12199
Rahnama N, Foo HL, Abdul Rahman NA, Ariff A, Md Shah UK (2014) Saccharification of rice straw by cellulase from a local Trichoderma harzianum SNRS3 for biobutanol production. BMC Biotechnol 14:1–12. https://doi.org/10.1186/s12896-014-0103-y
Rath J, Messner R, Kosma P, Altmann F, Miirz L, Kubicek CP (1995) The α-d-mannan core of a complex cell-waU heteroglycan of Trichoderma reesei is responsible for β-glucosidase activation. Arch Microbiol 164:414–419. https://doi.org/10.1007/BF02529739
Ronquist F et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Saloheimo M et al (2002) Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur J Biochem 269:4202–4211
Silva GGD, Rouau SGX (2011) Successive centrifugal grinding and sieving of wheat straw. Powder Technol 208:266–270. https://doi.org/10.1016/j.powtec.2010.08.015
Song Y, Wynn JP, Li Y, Grantham D, Ratledge C (2001) A pre-genetic study of the isoforms of malic enzyme associated with lipid accumulation in Mucor circinelloides. Microbiology 147:1507–1515. https://doi.org/10.1099/00221287-147-6-1507
Sun S, Sun S, Cao X, Sun R (2016) The role of pretreatment in improving the enzymatic hydrolysis of lignocellulosic materials. Bioresour Technol 199:49–58. https://doi.org/10.1016/j.biortech.2015.08.061
van Tilbeurgh H, Claeyssens M (1985) Detection and differentiation of cellulase components using low molecular mass fluorogenic substrates. Febs Lett 187:283–288. https://doi.org/10.1016/0014-5793(85)81260-6
Wan C, Li Y (2012) Fungal pretreatment of lignocellulosic biomass. Biotechnol Adv 30:1447–1457. https://doi.org/10.1016/j.biotechadv.2012.03.003
Zhang PYH, Himmel ME, Mielenz JR (2006) Outlook for cellulase improvement: screening and selection strategies. Biotechnol Adv 24:452–481. https://doi.org/10.1016/j.biotechadv.2006.03.003
Zitomer SW, Eveleigh D (1987) Cellulase screening by iodine staining: an artefact. Enzyme Microb Technol 9:214–216. https://doi.org/10.1016/0141-0229(87)90017-2
Acknowledgements
This study was supported by a grant from the Serbian Ministry of Education, Science and Technological Development (Project Grant Number OI172048). The work in TU Wien was supported by the Austrian Science Fund (FWF) Project Number P 25613 B20 to ISD. We thank Dr. Komal Chenthamara for help with in situ data analysis. We thank Jasna Bojčevski for English correction.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11274_2019_2774_MOESM1_ESM.tif
Fig. S1. Screening for efficient cellulase producers among Trichoderma isolates. Total liquefaction of the medium (level 4 from Table 1) occurred within 25 h for isolate T. guizhouense TG2 and first minor changes in viscosity (level 2 from Table 1) occurred within 25 h for T. reesei QM9414. (TIF 1509 kb)
11274_2019_2774_MOESM2_ESM.tif
Fig. S2. Position of Trichoderma strain TG2 in the tef1 Bayesian phylogenetic tree. Branching points supported by posterior probabilities higher than 0.94 are indicated by grey dots. Arrowindicates branching that leads to the Trichoderma harzianum species clade. Subclade names within the Trichoderma harzianum species complex, including strain TG2 are shown in bold. Tailing digits correspond to GenBank accession numbers. (TIF 12006 kb)
11274_2019_2774_MOESM3_ESM.tif
Fig. S3. Optimization of cellulase complex production by T. guizhouense TG2. Influence of a) substrate amount; b) agitation (CC- combined conditions); c) inoculation (SS-spore suspension; MD-mycelial disk; I1-inoculum1; I2-inoculum 2; I3-inoculum 3); and d) nitrogen sources, on production levels. Enzyme activities are presented as U/mL, and are displayed as the mean of three independent assays ± SEM (all below 5%). (TIF 6329 kb)
11274_2019_2774_MOESM4_ESM.pdf
Table S1. Optimization of the production of cellulase enzyme complex by TG2 with untreated wheat straw as substrate. (PDF 60 kb)
11274_2019_2774_MOESM5_ESM.pdf
Table S2. Comparison of maximal cellulase production (U/mL) of T. guizhouense TG2 and T. reesei QM9414 on raw wheat straw substrate. Enzyme activities are presented as U/mL, and are displayed as the mean of three independent assays ± SEM (all below 5%). (PDF 116 kb)
Rights and permissions
About this article
Cite this article
Grujić, M., Dojnov, B., Potočnik, I. et al. Superior cellulolytic activity of Trichoderma guizhouense on raw wheat straw. World J Microbiol Biotechnol 35, 194 (2019). https://doi.org/10.1007/s11274-019-2774-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-019-2774-y