Abstract
Aims
As a consequence of the increasing impact of climate change on crop production and food security worldwide, the need to explore agricultural systems in a sustainable manner is also intensified. The improvement of long-read metagenomics approaches might give valuable information not only on soil microbial communities, but also on their potential effects on plant phenotypes. Soil properties, climate conditions, and agricultural techniques are the main factors shaping microbial communities found in the soil and on the surface of the potatoes, influencing plant health and performance. The objective of this study was to decipher the bacterial communities in contrasting potato terroirs using long-read sequencing of the 16S rRNA gene.
Methods
To do so, 18 soil samples were taken from different potato fields in the island of Naxos (Island Terroir) and Northern Greece (Continental Terroir). Differences in soil properties and climatic conditions were also regarded to explore the possible motif of microbial structure and diversity in each region.
Results
Our results highlighted that contrasting potato terroirs strongly affected microbial community composition and diversity. A richer microbial composition in the island terroir was evident. A wide range of soil bacteria were identified, including Vicinamibacter, Neobacillus, Povalibacter, Microvirga, Thermoanaerobaculum, Arenimonas, and Rubrobacter, with different distribution patterns that resulted in characteristic microbial footprints.
Conclusions
In combination with soil analysis, microbial mapping might be a valuable tool, not only for gaining a deeper knowledge of their impact on potato production, but also for developing biomarkers that would uniquely define and characterize each potato habitat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple factors can hamper the process of plant growth and development, such as the imbalances in the microbial population and abundance in the rhizosphere, the soil acidification, as well as the rise of soilborne diseases owing to the presence of pathogenic fungi and bacteria, accompanied by the decrease in beneficial bacteria in the rhizosphere (Akram et al. 2019; Mahmud et al. 2021; Bhojiya et al. 2022; Upadhyay and Chauhan 2022). The rhizosphere is a zone of high microbial diversity and activity, acquiring nutrients from root exudates and detached plant cells (Faist et al. 2021). Climatic conditions (Mendes et al. 2013), soil properties (Breidenbach et al. 2016), agricultural management (Santos-Medellín et al. 2017), and cropping patterns (Shi et al. 2015) affect the rhizosphere and the soil microbial populations. The presence of archaea, bacteria, fungi, and other eukaryotic communities in the rhizosphere could not only improve soil, water, air quality, stimulation of organic matter decomposition, and plant nutrient absorption (Qin et al. 2017; Edwards et al. 2018; Hamonts et al. 2018), but also minimize soil-borne diseases (Zimudzi et al. 2018).
Potatoes are one of the world's most important non-cereal food crops, with its high nutritive value for millions of people around the globe (Liu et al. 2021). The quality and flavor of potatoes are influenced by a variety of factors, including soil characteristics, weather conditions, and farming practices (Bulgarelli et al. 2015; Zhalnina et al. 2018; Boutsika et al. 2022). These factors can shape the microbial communities present in the soil and on the surface of the potato tubers, which, in turn, might affect the plant's health and productivity (Faist et al. 2021). Beneficial bacteria in potato soils can enhance nutrient cycle, suppress diseases, and produce plant growth-promoting compounds (Compant et al. 2009; Mendes et al. 2013; Wagg et al. 2014). For example, soil bacteria break down complex organic substances into simpler ones that the potato plant can easily absorb (Jacoby et al. 2017). Additionally, bacteria in such soils may produce antimicrobial compounds or compete for soil resources to prevent the growth of harmful pathogens (Singh et al. 2017).
Following the unprecedented development of molecular techniques, microbiome analysis has also advanced significantly, producing vast quantities of assembled metagenomes of high quality from complex environmental samples (Fierer 2017; Boutsika et al. 2022; Albertsen 2023). Using sequencing platforms capable of producing longer reads, long-read metagenomics has the potential to provide more accurate taxonomic classification of microbial communities and to identify previously unknown microorganisms (Zhang et al. 2023). In a previous study on potato rhizosphere microbiota at different vegetation stages and regions, a plethora of opportunistic microbial strains were identified at specific growth stages, whereas few ubiquitous strains were also detected regardless of geographic location or development stage, probably being associated with potato growth (Pfeiffer et al. 2017). The bacterial diversity present in potato fields and expressed as a bacterial species equilibrium index, has been previously employed as a powerful biological indicator to determine soil quality indices (Jeanne et al. 2019). Using high-throughput sequencing of the 16S rRNA, the potential of a commercial microbial consortium (Bacillus subtilis and Trichoderma harzianum strains) to improve health and enhance productivity was demonstrated.
(Wang et al. 2019). Meanwhile, tare soils seem to influence rhizosphere microbial composition more strongly for fungi than bacteria, as revealed by ITS2, or 16S metabarcoding, respectively (Delventhal et al. 2022). The origin of bulk soil where potato tuber is grown, seem to be the main factor influencing its bacterial community diversity and composition, while the residual soil adhering to the tuber surface may serve as a new avenue for shaping beneficial microbiomes in the underground tissues (Bender et al. 2016).
In Greece, ‘Patata Naxou’ is produced in the island of ‘Naxos’, Aegean Sea, Greece, and certified as a Protected Geographical Indication (PGI) product. The distinctive properties of ‘Naxos’ potato, such as the high carotenoid content (Boutsika et al. 2023), is presumed to be due to the island’s particular soil and climatic characteristics, as well as due to the various conventional development strategies that the local producers and stakeholders are still using nowadays (European Commission 2020). In our previous studies, we reported the first integrated study combining genome-wide DNA methylation, RNA sequencing, and quantitative proteomics analysis to acquire the molecular portrait of the famous PGI potatoes during harvest and at post-harvest (Boutsika et al. 2023). The results demonstrated that the transcript expression and protein abundance of potatoes cultivated in distinct environments exhibited distinct differences, confirming previous findings that the transcriptome and the proteome represent valuable diagnostic tools for exploring plant performance in different terroirs (Wei et al. 2022). Herein, by using long-read metataxonomic approaches of the 16S rRNA gene, we aimed to obtain a thorough understanding of the microbial communities associated with potatoes in two contrasting terroirs (island vs continental), including their taxonomic composition and diversity. This study will shed light on the effect of soil on the potato holobiont and provide information on the microbial ecology of potato agriculture between two geographical regions with remarkably different environmental conditions.
Materials and methods
Sampling sites and sample collection
Potatoes (S. tuberosum L., cv. Spunta, Oldenburger, Assem, Holland) were cultivated in two contrasting regions of Greece in Naxos Island, Aegean Sea, (Island Terroir, IT), and in Chalkidiki, North Greece (Continental Terroir, CT) (Fig. 1a). Soil samples from each location (two geographic regions x three individual fields x three biological replicates = 18 soil samples) were collected in the period between the end of June 2021 (island region) till the beginning of July 2021 (continental region), along with potato tuber harvest. The potato cultivation in each geographical region followed the same crop management techniques (conventional). Composite soil samples were kept cool in a fridge with ice and returned to the lab in Ziploc bags within 24 h. After arriving at the lab, soil samples were homogenized and sieved to study soil physicochemical properties, whereas an aliquot of each sample was used immediately used for DNA isolation.
a: Map of Greece indicating the two sampling regions: 1. Continental Terroir (CT), and 2. Island Terroir (IT). For each sample location, the exact coordinates are provided in Supplementary Table 1. b: Heatmap representing soil properties (z-score normalized) of the two sampling sites. EC, electrical conductivity of the saturation extract; OM, Organic Matter; Si, Silicon
Soil properties
The soil samples were collected from the upper 30-cm layer (where the main part of the potato plants root system lays and nutrients are mainly taken up) of the two geographical regions (Fig. 1a; Supplementary Table 1). Each soil sample was consisted of five soil sub-samples randomly collected from each field and mixed to fix the homogenized soil sample. The soil samples were mechanically crumbled, air-dried, sieved with a 2-mm sieve, and prepared for assessing soil texture through particle size analysis with the hydrometer (or Bouyoucos) method (Gee and Bauder 1986). Then the following determinations were performed: % organic matter (OM) with the wet digestion method (Walkley and Black 1934), % CaCO3 with the back titration method (Loeppert and Suarez 1996), pH measured in saturated paste, and electrical conductivity (EC) measured in a soil saturation extract. Nitrate-nitrogen was extracted using 2 M KCl (Clesceri and Greenberg 1998), available phosphorus (P) according to the Olsen method (Watanabe and Olsen 1965), exchangeable calcium (Ca), potassium (K), and magnesium (Mg) with ammonium acetate at pH = 7.0 (Thomas 1982), copper (Cu), manganese (Mn), iron (Fe), and zinc (Zn) using DTPA (Lindsay and Norvell 1978), and extractable boron with the azomethine-H method (Bingham 1982). The concentrations of the exchangeable cations and extractable micronutrients (Ca, Mg, K, Fe, Mn, Zn, and Cu) were measured by ICP (OPTIMA 2100 DV, optical emission spectrometer, Perkin Elmer, Waltham, MA, USA).
Sample processing and sequencing
A total of 200 g (wet weight) of soil samples were used for DNA isolation. Genomic DNA was isolated with the DNeasy PowerSoil Pro Kit (QIAGEN, Carlsbad, USA), following the manufacturer’s protocol. The absorbance ratio of A260/280 nm and A260/230 nm were measured in a NanoDrop spectrophotometer (NanoDrop One/ NanoDrop One C, Thermo Scientific, United States) to determine the total DNA quantity and purity. The extracted DNA were then stored at -80 °C for all the downstream applications.
The full-length 16S rRNA ribosomal gene was amplified with PCR using a LongAmp Hot Start Taq 2 × Master Mix (M0533S, New England Biolabs), 16S barcoded primers, DNA template (10 ng), and nuclease-free water on an Applied Biosystems® QuantStudio® 5 Real-Time PCR System (Thermo Fischer Scientific, Waltham, MA, USA), essentially as previously described (Boutsika et al. 2023). Initial denaturation at 95 °C for 1 min was followed by 30 three-step amplification cycles: denaturation at 95 °C for 20 s, primer hybridization at 55 °C for 30 s, primer elongation at 65 °C for 2 min, and final elongation at 65 °C for 1 min. In order to sequence the 16S ribosomal gene and generate libraries, the 16S Barcoding Kit 1–24 (SQK-16S024, Oxford Nanopore Technologies, UK) was used. Purification of PCR products was done with Agecount AMPure XP beads (Beckman Coulter, USA), and quantification was done with the Qubit 4 Fluorometer and the dsDNA HS Assay Kit (Beckman Coulter, USA) (Thermo Fisher Scientific, USA). Following that, the PCR products were mixed in equimolar ratios, and the library was prepared according to the manufacturer's instructions. The library was then loaded into the MinION Mk1C (Oxford Nanopore Technologies, UK) using a MinION R9.4.1 flow cell (FLO-MIN106) and MINKNOW software (Version 1.11.5) (Oxford Nanopore Technologies) for data acquisition.
Data analysis and read processing
The Guppy program (Version 5.0.17) was used to convert MinIONTM sequence reads (i.e., FAST5 data) into FASTQ files (Oxford Nanopore Technologies). The EPI2ME software, which is based on Nextflow (DI Tommaso et al. 2017), was used to construct bacterial communities. DNA sequences from microbial samples were classified utilizing the Centrifuge program (Kim et al. 2016). This program, which is based on the Burrows-Wheeler Transformation and the Ferragina-Manzini index, enables precise and quick metataxonomic analysis. Python scripts were applied to the CSV files to match NCBI taxonomic IDs to lineages and count the number of readings per NCBI taxonomic ID.
R programming language packages were applied in RStudio (Version 4.2.2), combined with several software to further analyze the data (McMurdie and Holmes 2013). Using the “vegan” and “betapart” packages, (Baselga and Orme 2012) the alpha-diversity and beta-diversity were calculated. To calculate alpha diversity indices, we have employed the relative abundance data. Likewise, Principal coordinates analysis (PCoA) and Non-metric Multi-dimensional Scaling (NMDS) were analyzed and plotted. The PCoA analysis was based on the presence/absence of data from the total data set, while the NMDS analysis in our study involved transformations such as Wisconsin double standardization and square root transformation. These transformations were applied to adjust the scale and distribution of the data. Subsequently, analysis of similarities (ANOSIM) was also performed, using the “vegan” and “betapart” package, while heatmap plots based on z-score normalization of relative abundances of reads were generated using the “gplots” package. Lastly, the linear discriminant analysis (LDA) effect size (LEfSe) study (Segata et al. 2011) was conducted in MicrobiomeAnalyst web-based tool (Lu et al. 2023), using log-transformed data, to define the microbiological variation between the distinct groups and to discover suitable biomarkers for each geographical region (IT and CT).In addition, in an effort to compare bacteria composition and diversity between soil samples and potato tubers, we performed a meta-analysis of the dataset from BioProject accession number PRJNA854325 (Boutsika et al. 2023). In particular, we performed a Non-metric Multidimensional Scaling (NMDS) analysis between soils and tubers originating from the same field and collected or harvested at the same time points.
Statistical analysis
The results of soil analysis were firstly tested for normality using the Anderson and Darling normality test, revealing that soil data were normally distributed. Student’s T-test was then applied to determine differences in soil properties between IT (n = 9) and CT (n = 9), whereas the Average Linage clustering method with Euclidean distances was used to cluster the sampling sites in RStudio (Version 4.2.2). To evaluate the normality of our sequencing dataset, we also utilized the Anderson–Darling test that resulted in p-value < 2.2e-16, clearly evidencing against the null hypothesis of normality. The differences in the relative abundance between the two groups (IT vs CT) were evaluated using Wilcoxon rank sum test. When p < 0.05, we considered the difference significant. The Wilcoxon rank-sum test was also utilized to determine alpha diversity indices (Simpson, Shannon, and Chao) at both genera and species level, between the two regions. All analyses were based on 18 samples (two geographic regions x three individual fields x three biological replicates).
Data availability
Raw data were deposited in the National Centre for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under the BioProject accession number PRJNA970975.
Results
Soil properties in the sampling sites
Overall, soil samples obtained from IT and CT showed discernible difference in their properties (Supplementary Table 2). In particular, the soils collected from IT (Naxos Island, Aegean Sea) were classified as loamy sand or sandy loam (clay content, 11.3%; sand content, 66.0%) with significantly higher OM content (2.2%), lower pH (7.6), and higher EC than CT soils (Fig. 1b; Supplementary Table 2). By contrast, the soils of CT (Chalkidiki, Central Macedonia) were classified as clay loam (clay content, 26.0%; sand content, 47.9%) with a significantly lower OM content (1.5%), higher pH (7.9), and lower EC than IT soils. With regard to macro- and micro-nutrients, both terroirs represent nutrient-rich ecosystems, but with some differences in soil fertility. More specifically, IT had higher mean concentrations of P and Zn, while CT had higher mean concentration of Mg, Mn, Cu and B. Heatmap analysis revealed that the amount of S, Zn, P, Fe, as well as OM and EC, was higher IT soil, which was diversified from CT soils creating a separate cluster. On the contrary, it is noticeable that CT samples perform a different cluster, due to the increased amounts of K, Mn, Cu, Mg, pH, and Si. Overall, the results of the analysis of soils from IT and CT demonstrated discernible differences in terms of their physical and chemical properties, based on Euclidean distances, except for IT1, which formed a separate cluster but closer to CT than the other IT samples. These findings emphasize the need to understand how soils in different regions affect plant growth and ecosystems.
Bacterial communities’ diversity and composition
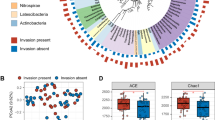
The high variability in soil properties between contrasting terroirs can lead to a wide range in microbial structure and diversity of soil microorganisms. Thus, to detect distinct differences in the soil bacterial profiles found in the two different agroecosystems, where potato is cultivated, bacterial 16S rRNA gene long-read sequencing was performed using a Nanopore MinION-based metataxonomic sequencing pipeline. Taxa abundances at various taxonomic levels – phylum, genus, and species – were calculated between the two terroirs, highlighting distinct bacterial communities between regions, which tended to differ in both composition and abundance (Fig. 2; Supplementary Tables 3, 4). At the phylum level, the top abundant phyla shared by IT and CT were Pseudomonadota, Bacillota, Actinomycetota and Bacteroidota, followed by Acidobacteriota and Thermodesulfobacteriota (Supplementary Table 4). Notably, IT had higher relative abundances of Acidobacteriota, Thermodesulfobacteriota and Verrucomicrobiota (Fig. 2a), whereas CT of Bacillota (Fig. 2b).
At the genus level, the top genera for IT were Vicinamibacter, Neobacillus, Haliangium, Gemmatimonas, and Paenibacillus (Fig. 2c), while for CT, there were Vicinamibacter, Microvirga, Haliangium, Gemmatimonas and Thermoanaerobaculum (Fig. 2d). Other abundant genera were Thiobacter, Bacillus and Fimbriiglobus for IT, as well as Brevitalea, Gaiella, Rubrobacter and Arenomicrobium for CT. Soils from IT possessed significantly more Neobacillus (Bacillota), Thiobacter and Povalibacter (Pseudomonadota) (Supplementary Table 5). By contrast, CT had higher relative abundances of Vicinamibacter, Microvirga, Thermoanaerobaculum, Brevitalea and Arenimicrobium (Acidobacteriota), as well as of Gaiella and Rubrobacter (Actinomycetota). Meanwhile, at the species level, the two terroirs shared high abundancies of Vicinamibacter silvestris, Haliangium tepidum and Fimbriiglobus ruber (Supplementary Table 3). The most discriminated abundant bacteria for IT were Neobacillus niacini, Povalibacter uvarum, and Thiobacter subterraneus, whereas for CT, there were Thermoanaerobaculum aquaticum, Arenimicrobium luteum, and Gaiella occulta.
Island vs continental terroir and spatial heterogeneity
To examine the distribution of reads assigned to taxa across various samples, venn diagrams and heatmap plots were generated for the genus and the species level (Fig. 3). For genera, the results showed that 587 taxa were shared by both terroirs, whereas the number of specific taxa were 12 and one (1), for IT and CT, respectively (Fig. 3a). At the species level, the shared taxa were 3922, while the unique taxa were 2142 and 927, for IT and CT, respectively (Fig. 3b). These data indicated that IT had a greater number of unique genera and species than CT, confirming the differentiation between the two regions (Fig. 3c, 3d). A heatmap visualization with hierarchical clustering of genera and species in all soil samples also depicted the distinct differences in bacterial community structure between the two geographical regions (Fig. 3c, 3d).
Analysis of taxa similarity among different terroirs. Venn diagrams represent shared and unique taxa at the genus (a) and the species (b) level. Heatmap plots represent the top 80 most abundant taxa across Island Terroir (IT) and Continental Terroir (CT), based on z-normalization of relative abundance data, ordered by distance-based clustering (Euclidian), at the genus (c) and the species (d) level
Alpha- and beta-diversity analysis
Alpha diversity measures (Chao1, Shannon, and Simpson indices) are presented in Fig. 4, highlighting increased genera/species richness and diversity in soils of IT compared to CT. At both genera and species level, there were significant differences between IT and CT for all alpha-diversity indices, as determined by Wilcoxon rank-sum test (p < 0.05) (Supplementary Table 5). In particular, chao1, which is an indicator of species richness (total number of species in a sample) that is sensitive to rare taxa (Chao 1987), showed higher values in IT than CT, both at the genus and the species level, indicating greater richness. Shannon’s index, which serves as an indicator of species evenness (proportional distribution of the number of each species in a sample), further confirmed that IT’s genera and species had greater diversity than CT’s. Lastly, Simpson, which is an indicator of species evenness (proportional distribution of the number of each species in a sample), also revealed the greater community diversity in IT samples. Soil samples of CT were characterized by variable intra-diversity at the genus level for richness (Fig. 4a) and at the species level for evenness (Fig. 4f), which made the box-plot range larger than in IT, suggesting that some CT replicates were highly diverse and rich, while others were characterized by their lower richness and proportional distribution in bacterial communities. Collectively, these data may indicate that IT soils, where the PGI potato originates from, had high species diversity, probably with less dominant species, whereas the CT soils represented more homogeneous bacterial communities, potentially with a dominance of a few species.
Violin plots representing the alpha-diversity of the bacteria from Island Terroir (IT) and Continental Terroir (CT). The plots compare the two regions at the genus level (a, b, c) and the species level (d, e, f) using Chao1, Shannon, and Simpson indices. Each violin plot displays the distribution of the alpha-diversity values, while the overlaid boxplot provides additional summary statistics such as the median, quartiles, and potential outliers for a comprehensive understanding of the dataset. Wilcoxon rank-sum was used to test for differences in alpha diversity indices between groups. *p < 0.05, **p < 0.01 and ***p < 0.001
Regarding beta-diversity, the principal coordinates analysis (PCoA) revealed vibrant differences between IT and CT soils (Fig. 5). More specifically, IT soil samples were separated from CT samples at both the genus and at the species level, since the plots create distinct clusters (Fig. 5a, c). The analysis of similarities (ANOSIM), based on Jaccard dissimilarity, further confirmed significant differences between the two regions (R = 0.834, P < 0.001 and R = 0.847, P < 0.001 for genera and species, respectively) (Fig. 5b, d).
Beta-diversity and community similarity analysis of the bacterial composition present in Island Terroir (IT) and Continental Terroir (CT), using the relative abundances of the genera (a, b) and species (c, d). a, c: Principal Coordinates Analysis (PCoA) plots based on Jaccard dissimilarity. b, d: Unweighted Unifrac ANOSIM analysis between the two terroirs
In an effort to compare bacterial communities that were present in the soil compared to the rhizosphere of potato tubers grown in the same IT and CT fields, an NMDS analysis using Bray–Curtis dissimilarity (-diversity) was performed using the dataset generated in this study, compared with a previously published (PRJNA854325; Boutsika et al. 2023). Results revealed significant regional differentiation for both genera and species between the potato rhizosphere and the surrounding soil (Fig. 6a), as they were clustered independently, corroborating the hypothesis that the sampling locations had distinguished microbial community composition.
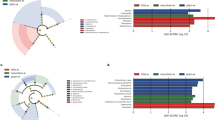
a, b: Non-metric multidimensional scaling (NMDS), using taxa relative abundances, showing ordination of bacteria found in the rhizosphere of potato tubers and the surrounding soils of IT and CT, harvested at the same time, at the genus (a) and at the species (b) level. The ordinate analysis is based on the Bray–Curtis distance matrix. Percentage following axis labels indicated percentage of total inertia explained by the respective axis. Number of samples: Island Potatoes (IP) = 23, Continental Potatoes (CP) = 18, Island Terroir (IT) = 9, and Continental Terroir (CT) = 9. c: Histogram of LDA Effect Size (LEfSe) distribution genera with significant differences in abundance between groups. Box plots (log-transformed counts) of the most differential genera in IT and CT are also shown.
Linear discriminant analysis effect size
The LEfSe analysis detected 12 and 15 bacterial clades in the soil of IT and CT, respectively, discriminating the terroir-specific microbial communities in the two geographic regions (Fig. 6b). At the genus level, candidate biomarkers for IT were Neobacillus and Povalibacter, while for CT, there were Vicinamibacter, Arenimicrobium and Rubrobacter These potential biomarkers were associated with each terroir, revealing geographic-origin dissimilarities. Notably, one genus identified as bacterial biomarker of IT, Neobacillus, was also detected both at harvest and at post-harvest potato tubers, obtained simultaneously from the same fields, as soil samples (Boutsika et al. 2023). This genus was abundant in the PGI potatoes, but it also remained abundant after storage, dominating the microbial community of potato tubers. These metataxonomic results obtained from soil and tubers showed a consistency for this genus, that was abundant in different potato fields across the island, thus it may represent an excellent putative ‘terroir’ biomarker for the traceability of the certified potatoes cultivated there.
Correlation plot analysis between genera and soil properties
A correlation plot analysis integrating soil properties and bacterial abundances for CT and IT was employed in an effort to associate specific taxa (the top 20 genera) with soil characteristics (Fig. 7). The correlations were visually represented using circles, where the size and color of the circles indicated the degree of the connection. Among the several genera that were analyzed, Vicinamibacter, Thermoanaerobium, Gaiella, Rubrobacter, Tepidisphaera, Gemmata, Arenimicrobium, Fimbriiglobus, Flavisolibacter, Brevitalea, Microvirga, and Neobacillus exhibited the strongest positive and negative associations. In particular, Vicinamibacter, Thermoanaerobium, Gaiella, Rubrobacter, and Tepidisphaera, exhibited significant interactions with seven (7) to nine (9) soil properties. Similarly, Gemmata, Arenimicrobium, Fimbriiglobus, Flavisolibacter, and Brevitalea had correlations with five (5) to six (6) soil properties, while Microvirga and Neobacillus demonstrated fewer correlations. Notably, the bacteria that exhibited the strongest associations were exclusively associated with CT. In this regard, the abundance of Vicinamibacter, Thermoanaerobium, Gaiella, and Rubrobacter was positively correlated with C, pH, Mg, Mn, Cu, and B, while also displaying strong negative correlations with Ec, P, and Zn. Brevitalea and Arenimicrobium exhibited comparable trends, with strong positive associations with Mg, Mn, Cu, and B, and negative associations with Ec. Furthermore, Gemmata and Fimbriiglobus had similar patterns characterized by strong positive correlations with C, pH, and Mg, while showing negative correlations with P and Zn. Finally, it is noteworthy that Flavisolibacter and Microvirga exhibited exclusively strong positive relationships. Flavisolibacter showed positive correlations with C, Si, pH, Mg, and Cu, while Microvirga displayed significant positive correlations with Mg, Mn, and Cu. Among the genera that were more abundant in IT, Neobacillus was positively correlated with P and negatively with pH and Mg. Overall, these findings provide some useful insights into the unique microbial-ecological dynamics that are specific to these regions, and merits further investigation.
Pearson correlation analysis of all pairwise comparisons between 16 soil characteristics and the top 20 most abundant genera. The magnitude of the correlation is depicted by the size and color of the circles. Light brown to dark brown coloring indicates a lower to higher negative correlations, while light green to dark green coloring indicates lower to higher positive correlations
Discussion
Potatoes are one of the world's most important crops (Visser et al. 2009), with their quality being highly dependent on the terroir they grow (Costantini and Bucelli 2013), and specifically on soil, geography, and climate (Lucini et al. 2020). Along with the genotypic effect, the sprouting, the quality, and the postharvest potential of potatoes, they can be also influenced by the soil properties and the environment during growth (Visser et al. 2009) and development (Buchholz et al. 2019). Furthermore, as the soil represents the main reservoir of microorganisms able to colonize potato tubers (Fierer and Jackson 2006; Genitsaris et al. 2020), it is conceivable that contrasting locations differing in the composition of the soil microbiota can also impact both potato quality and storability, even if the genotype remains the same. In the past, the majority of studies have focused on unravelling the effect of diverse bacterial communities that are present in potatoes with regard to productivity or resistance to soil-borne pathogens (Sessitsch et al. 2004; Guyer et al. 2015; Pfeiffer et al. 2017). Since the effect of potato genotype on the bacterial diversity seem to be minor or not consistent (Weinert et al. 2010), in this study, using nanopore long reads, we explored soil metataxonomics in contrasting potato fields from island and continental terroirs. Our findings demonstrated that the two areas (CT, IT) had distinguishable bacterial communities, suggesting the presence of terroir-specific bacterial species in both regions. Interestingly, the soils of IT, where the popular PGI potato is produced, had a bigger number of distinct species and genera than CT, indicating that the microbial ecosystems of the two regions varied substantially.
The genera of Neobacillus (Bacillota), Povalibacter, and Thiobacter (Pseudomonadota) have been found to be highly abundant in IT. Several species of the genus Neobacillus has been previously reported to promote plant growth through a variety of direct and indirect mechanisms, including nitrogen fixation, the use of biological controls to combat soil-borne pathogens, the production of plant growth hormones that enhance plant development, and the production of metabolites (Alawiye and Babalola 2019). The other highly abundant genus in IT, Povalibacter, is well known to assimilate N by reducing NO3− and NO2− into NH4+ (Nogi et al. 2014), usually associated with neutral pH ranges and with soils with higher amounts of soil organic matter and soil organic carbon (Kim et al. 2023). This is in line with our findings, demonstrating that IT had nearly twofold higher OM content, as well as lower pH than CT. Additionally, the abundance of this genus in long-term continuous cropped soils has been found to decrease due to the lack of organic material (Li et al. 2018), which might explain its lower abundances in CT. Another key genus in IT, Thiobacter, is able to store massive amounts of sulfur and to form conspicuous gelatinous mats, and it has been previously suggested to belong to the sulfide-oxidizing bacteria, using hydrogen sulfide as energy source (Grünke et al. 2010). This group of bacteria can therefore play a key role in transformation of sulfur in soil (Joshi et al. 2021). Although sulfur content was not determined in this study, the presence of such bacteria in the soils of IT, containing significantly higher organic matter content, may be associated with its higher availability for plants.
On the other hand, key genera that were significantly more abundant in CT included the Acidobacteriota Vicinamibacter, Microvirga, Thermoanaerobaculum, Brevitalea, Arenimicrobium, as well as the Actinomycetota Gaiella and Rubrobacter. The genus Vicinamibacter contain bacteria that can survive in a broad variety of pH levels, able to use different sugars, organic acids, and complex proteinaceous substrates to thrive (Huber et al. 2016). Significantly, these genera have been linked to the process of P solubilization, as shown by the presence of quinoprotein glucose dehydrogenase genes in their genomes (Liang et al. 2020; Wu et al. 2022). The genes catalyze the oxidation of glucose into gluconic acid, having a pivotal role in microbial P cycling in the soil (Zhao et al. 2022). Although the association between P-transforming microorganisms and the long-term nutrient inputs remain patchy at best, high P inputs are generally associated with the inhibition of growth of Acidobacteria, being oligotrophic bacteria (Dai et al. 2020; Liu et al. 2022). This is in agreement with the high P availability in IT, accompanied by lower abundances of Acidobacteria, such as Vicinamibacter, Microvirga, Thermoanaerobaculum, Brevitalea and Arenimicrobium. On the other hand, Microvirga species can be found in a wide range of ecological niches and have crucial functions in enhancing plant growth and suppressing pathogenic microorganisms (Liu et al. 2022), being commonly found in healthy soils (Wang et al. 2017). Several members of this genus are important in symbiotic nitrogen fixation, having the capacity to reduce nitrate to nitrite (Dahal and Kim 2017). Herein, no high correlation between the abundance of this genus and nitrate was evident, but instead, a high positive correlation was observed for Mg and some micronutrients i.e., Mn, Cu and B. Another genus that was significantly more abundant in CT was Thermoanaerobaculum which is known for its remarkable capacity to reduce Fe(III) and Mn(IV) (Losey et al. 2013). The concentration of Fe was not substantially different between the two terroirs, but Mn levels were higher in CT, being positively associated with the abundance of Thermoanaerobaculum. Other abundant genera found in CT, Brevitalea and Arenimicrobium, are aerobic chemoheterotrophic mesophiles that may thrive in a wide variety of pH levels, using a limited spectrum of carbon and energy sources for growth, with a preference for complex proteinaceous substrates (Wüst et al. 2016). The Brevitalea and Arenimicrobium clades are commonly found in dry soil settings, indicating their ability to adapt to arid circumstances and their dependence on certain carbon sources that are abundant in these environments (Dedysh and Yilmaz 2018).
Within the phylum Actinomycetota (or Actinobacteria), the abundances of Gaiella and Rubrobacter, were significantly higher in CT. The genus Gaiella is characterized by Gram-negative, non-motile rod-shaped bacteria, known as important organic matter decomposers, involved in carbon cycling (Albuquerque and da Costa 2014). Interestingly, the genus Gaiella was found to be significantly enriched in vermicompost soil, inhibiting the abundance of Fusarium oxysporum f. sp. lycopersici (Zhou et al. 2021), although the underlying mechanism of pathogen inhibition remains unclear. Previously, a correlation between the abundance of Gaiella and moderately high amounts of rainfall was demonstrated (Amon et al. 2023), being consistent with our report of remarkably higher rainfall levels in CT compared to IT during the last two months prior soil sampling (Supplementary Table 6). Earlier studies have also reported that the abundance of Gaiella genus can be significantly reduced in soils fertilized with pig manure (Liu et al. 2017) or poultry litter (Parente et al. 2021), which may justify its lower abundance in IT, where soils had higher OM content (Supplementary Table 2). Within the same phylum clade, Rubrobacter is another prominent and widespread genus, commonly found in cultivated fields and marine ecosystems, with variable abundance and diversity (Holmes et al. 2000; Chen et al. 2021). This genus is known for its exceptional ability to withstand radiation (Yoshinaka et al. 1973) and is commonly found in terrestrials with high-temperature, or with elevated salt concentrations (Singleton et al. 2003). However, none of these observations were confirmed in this study, as CT had lower average temperature, as well as lower EC compared to IT. Furthermore, prior studies have emphasized on the existence of extremely distinct proteins in Rubrobacter that are essential for the development of bacteriochlorophyll-based photosynthesis, providing insight into its evolutionary importance (Gupta and Khadka 2016).
An interesting finding highlighted in the current study is the identification of possible bacterial biomarkers, with statistically significant higher relative representation, specific for each terroir. In particular, the genera Neobacillus and Povalibacter served as good candidates for IT, whereas Arenimicrobium and Thermoanaerobaculum for CT. Although these potential biomarkers need to be validated over different growing seasons, our results highlight the possibility to employ such microbial biomarkers to distinguish various geographical locations, especially those related to PGI products. Another interesting note is that Neobacillus was discovered in both the soil (this study) and potato tubers harvested from the same fields simultaneously (Boutsika et al. 2023), thus representing an ideal "terroir" indicator for traceability. It is noteworthy to mention that Neobacillus, the putative a biomarker in IT region, was positively correlated with EC, suggesting its putative association with coastal regions. Therefore, this method, along with other analytical assays related to product quality, may be potentially applied towards the prevention of misleading indications of PGI origin, as well as for the protection of authentic local products and production methods from exploitation, imitation, and deception. Similar studies provide interesting linkages between grape microbial communities and the wine terroir (Kamilari et al. 2021).
On the other hand, the genera Bacillus, Paenibacillus, Haliangium, Fibriiglobus and Gemmatimonas were highly abundant in both terroirs, with no particular differences in abundance, suggesting that they can be regarded as “core” soil microbiome for potato sampling sites in Greece. Overall, these genera can have significant impacts on plant growth promotion and on the suppression of pathogen attack (Leontidou et al. 2020; Liu et al. 2022), possessing multiple ecological functions in soil ecosystems such as nutrient cycling, synthesis of phytohormones and plant stress tolerance (Timmusk et al. 2005; Grady et al. 2016; Jeong et al. 2019a, b; Saxena et al. 2020; Varliero et al. 2021). Interestingly, these genera were also relatively abundant in potato tubers harvested from these sampling sites (Boutsika et al. 2023), further strengthening their putative contribution to the “core” microbiome, despite huge differences in soil properties, cropping history and climate. By contrast, in a recent review, the genera Bradyrhizobium, Sphingobium and Microvirga have been suggested as the main representatives of the potato core microbiome (Petrushin et al. 2024). Therefore, the presence of generalist genera over different vegetation stages and years, as well as in other potato varieties and environmental conditions, needs to be further investigated to point toward an even closer association between potato cultivation and the core microbiome composition.
Although linking alterations in soil bacterial communities with agricultural productivity or quality may seem premature (Peiffer et al. 2013; Hartmann and Six 2023), the contribution of soil microbiome as a potential driver of the “terroir” signature cannot be ignored (Boutsika et al. 2023; Johnston-Monje et al. 2023). Among all the components constituting terroir, climate (including solar radiation, precipitation, humidity, temperature and wind) is a dominant factor governing not only agricultural product characteristics but also bacteria community composition and diversity. Previously, the diversity and abundance of soil bacteria have been shown to be inversely connected with temperature and positively with soil humidity (Rousk et al. 2010). Based on these considerations, the observed differences in the composition of the bacterial communities of IT and CT may be justified by differences in the average temperature (slightly higher in IT) and the average rainfall (significantly higher in CT) (Supplementary Table 6). Furthermore, bacterial metabolism may accelerate the decomposition of OM and soil nutrient release at high temperatures (Yu et al. 2022), whereas higher soil moisture may cause anaerobic conditions and induce alterations in soil diversity (Fierer et al. 2003). The Aegean Islands are famous for their low-productivity Lithic Xerorthents or Leptosol volcanic soils (Moustakas and Georgoulias 2005), with the soil being also xeric due to minimal precipitation and high evaporation rates. Therefore, we could presume, that the diverse microbiome of IT and CT may vary due to their diversified microclimate and particularly due to significantly different average temperature and precipitation levels, along with soil properties, that can jointly shape the so-called “terroir footprint”.
As soil microbiomes drive key functions in agroecosystems, regulating soil fertility, crop productivity and stress resilience (Hartmann and Six 2023), several analytical approaches have been employed to tap microbial diversity, mainly by sequencing one or two hypervariable regions of the 16S rRNA gene (Yang et al. 2016). In our study, we have sequenced the full-length 16S rRNA gene for the metataxonomic analysis by employing the Nanopore EPI2ME pipeline. Despite the limited possibility to customize workflow parameters, as well as the higher error rate characterizing nanopore sequencing, the sequencing of the full-length 16S rRNA gene allows species-level classification, enhancing taxa resolution over previous short-read technologies (Benítez-Páez et al. 2016; Ciuffreda et al. 2021). Future advances on nanopore long-read chemistry, with the implementation of different bacterial reference databases, could surpass current problems maximizing reliability and confidence on data analysis, and will be pivotal in the research towards harnessing soil microbiomes as a sustainable solution for modern agriculture.
Conclusions
This study offers useful insights into the microbial diversity and composition of potato fields in two contrasting terroirs. Significant variations in bacterial communities across the two locations depicts the necessity for region-specific soil management practices to maximize crop yields and preserve soil health. Using long-read sequencing of the 16S rRNA gene, a breakthrough method that has made closed microbial genomes routine creating a complete microbial tree of life, we uncovered that potato soils in the semi-arid region of the Aegean Island, which represents a unique Mediterranean agroecosystem, recruited more beneficial microorganisms, possibly to cope with the unfavorable environmental conditions. These results are in accordance with the notion that soil microbial biogeography is predominantly governed by regional soil properties, unique for each terroir. Despite the fact that our research was limited to three different fields per regions and one year of sampling, our results pave the way towards understanding the factors that determine microbial community structure and diversity of dissimilar terroirs, that is crucial for preserving soil quality, avoiding soil-borne plant diseases, and enhancing plant growth. Further experimentation over different years and climatic conditions are crucial to unravel how different agricultural terroirs shape potato microbiome and to understand if the distinguishable microbial footprints can contribute to the regional identity of the produced potatoes.
Data availability
Raw data of were deposited in the National Centre for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under BioProject accession number PRJNA970975.
References
Akram W, Aslam H, Ahmad SR et al (2019) Bacillus megaterium strain A12 ameliorates salinity stress in tomato plants through multiple mechanisms. J Plant Interact 14:506–518. https://doi.org/10.1080/17429145.2019.1662497
Alawiye TT, Babalola OO (2019) Bacterial Diversity and Community Structure in Typical Plant Rhizosphere. Diversity (basel) 11:179. https://doi.org/10.3390/d11100179
Albertsen M (2023) Long-read metagenomics paves the way toward a complete microbial tree of life. Nat Methods 20:30–31. https://doi.org/10.1038/s41592-022-01726-6
Amon CER, Fossou RK, Ebou AET, Koua DK, Kouadjo CG, Brou YC, Voko Bi DRR, Cowan DA, Zézé A (2023) The core bacteriobiome of Côte d’Ivoire soils across three vegetation zones. Front Microbiol 14:1220655. https://doi.org/10.3389/fmicb.2023.1220655
Baselga A, Orme CDL (2012) betapart: an R package for the study of beta diversity. MEE 3:808–812. https://doi.org/10.1111/j.2041-210X.2012.00224.x
Bender SF, Wagg C, van der Heijden MGA (2016) An Underground Revolution: Biodiversity and Soil Ecological Engineering for Agricultural Sustainability. Trends Ecol Evol 31:440–452. https://doi.org/10.1016/j.tree.2016.02.016
Benítez-Páez A, Portune KJ, Sanz Y (2016) Species-level resolution of 16S rRNA gene amplicons sequenced through the MinION™ portable nanopore sequencer. GigaScience 5(1): s13742–016–0111-z. https://doi.org/10.1186/s13742-016-0111-z
Bhojiya AA, Joshi H, Upadhyay SK et al (2022) Screening and Optimization of Zinc Removal Potential in Pseudomonas aeruginosa-HMR1 and its Plant Growth-Promoting Attributes. Bull Environ Contam Toxicol 108:468–477. https://doi.org/10.1007/s00128-021-03232-5
Bingham FT (1982) Methods of Soil Analysis. Part 2. In: Page AL (ed) Boron. American Soceity of Agronomy, Madison, WI, USA, pp 431–448
Boutsika A, Michailidis M, Ganopoulou M, et al (2023) A wide foodomics approach coupled with metagenomics elucidates the environmental signature of potatoes. iScience 26:1. https://doi.org/10.1016/j.isci.2022.105917
Boutsika A, Tanou G, Xanthopoulou A et al (2022) Insights and advances in integrating multi-omic approaches for potato crop improvement. Sci Hortic 305:111387. https://doi.org/10.1016/j.scienta.2022.111387
Breidenbach B, Pump J, Dumont MG (2016) Microbial community structure in the rhizosphere of rice plants. Front Microbiol 6:1537. https://doi.org/10.3389/fmicb.2015.01537
Buchholz F, Antonielli L, Kostić T et al (2019) The bacterial community in potato is recruited from soil and partly inherited across generations. PLoS ONE 14:e0223691. https://doi.org/10.1371/journal.pone.0223691
Bulgarelli D, Garrido-Oter R, Münch PC et al (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403. https://doi.org/10.1016/j.chom.2015.01.011
Chao A (1987) Estimating the Population Size for Capture-Recapture Data with Unequal Catchability. Biometrics 43:783. https://doi.org/10.2307/2531532
Chen RW, He YQ, Cui LQ, Li C, Shi SB, Long LJ et al (2021) Diversity and Distribution of Uncultured and Cultured Gaiellales and Rubrobacterales in South China Sea Sediments. Front Microbiol 12:657072. https://doi.org/10.3389/fmicb.2021.657072
Chin LM, Wong Vui Ling CM (2020) Effects of Elevated Temperature on the Tropical Soil Bacterial Diversity (Kesan Peningkatan Suhu terhadap Kepelbagaian Bakteria Tanah Tropika). Sains Malaysiana 49(10): 2335–2344. https://doi.org/10.17576/jsm-2020-4910-01
Ciuffreda L, Rodríguez-Pérez H, Flores C (2021) Nanopore sequencing and its application to the study of microbial communities. CSBJ 19:277–284. https://doi.org/10.1016/j.csbj.2021.02.020
Clesceri LS, Greenberg AE (1998) Standard Methods for the Examination of Water and Waste Water. American Public Health Association Washington DC 19
Compant S, Clément C, Sessitsch A (2009) Plant growth-promoting bacteria in the rhizo- and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem 42:669–678. https://doi.org/10.1016/j.soilbio.2009.11.024
Costantini CEA, Bucelli P (2013) Soil and Terroir. In: SpringerBriefs in Environment, Security, Development and Peace , Springer, Cham. Springer, Cham, pp 97–133
Dahal RH, Kim J (2017) Microvirga soli sp. nov., an alphaproteobacterium isolated from soil. IJSEM 67:127–132. https://doi.org/10.1099/ijsem.0.001582
Dai Z, Liu G, Chen H et al (2020) Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems. ISME J 14:757–770. https://doi.org/10.1038/s41396-019-0567-9
Dedysh SN, Yilmaz P (2018) Refining the taxonomic structure of the phylum Acidobacteria. IJSEM 68(12):3796–3806. https://doi.org/10.1099/ijsem.0.003062
Delventhal MsK, Skillman MsV, Li DrX et al (2022) Characterizing variation in the bacterial and fungal tare soil microbiome of the seed potato. APS. https://doi.org/10.1094/PBIOMES-11-22-0092-R
Di Tommaso P, Chatzou M, Floden EW et al (2017) Nextflow enables reproducible computational workflows. Nat Biotechnol 35:316–319. https://doi.org/10.1038/nbt.3820
Edwards J, Santos-Medellín C, Sundaresan V (2018) Extraction and 16S rRNA Sequence Analysis of Microbiomes Associated with Rice Roots. Bio Protoc 8:2884. https://doi.org/10.21769/BioProtoc.2884
Commission E (2020) Publication of an application for approval of an amendment, which is not minor, to a product specification pursuant to Article 50(2)(a) of Regulation (EU) No 1151/2012 of the European Parliament and of the Council on quality schemes for agricultural products and foodstuffs 2020/C 274/08. Off J Eur Union 274:8–29
Faist AM, Antoninka AJ, Barger NN et al (2021) Broader Impacts for Ecologists: Biological Soil Crust as a Model System for Education. Front Microbiol 11:3284. https://doi.org/10.3389/fmicb.2020.577922
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590. https://doi.org/10.1038/nrmicro.2017.87
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631. https://doi.org/10.1073/pnas.050753510
Fierer N, Schimel JP, Holden PA (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35:167–176. https://doi.org/10.1016/S0038-0717(02)00251-1
Gee GN, Bauder JW (1986) Particle Size Distribution. In: Klute A (ed) Physical and Mineralogical Methods, 2nd edn. Agronomy Society of America/Soil Science Society of America, Madison, Wisconsin, pp 383–411
Genitsaris S, Stefanidou N, Leontidou K et al (2020) Bacterial Communities in the Rhizosphere and Phyllosphere of Halophytes and Drought-Tolerant Plants in Mediterranean Ecosystems. Microorganisms 8:1708. https://doi.org/10.3390/microorganisms8111708
Grady EN, MacDonald J, Liu L et al (2016) Current knowledge and perspectives of Paenibacillus: A review. Microb Cell Factories 15:203. https://doi.org/10.1186/s12934-016-0603-7
Grünke S, Lichtschlag A, Beer D et al (2010) Novel observations of Thiobacterium, a sulfur-storing Gammaproteobacterium producing gelatinous mats. ISME J 4:1031–1043. https://doi.org/10.1038/ismej.2010.23
Gupta RS, Khadka B (2016) Evidence for the presence of key chlorophyll-biosynthesis-related proteins in the genus Rubrobacter (Phylum Actinobacteria) and its implications for the evolution and origin of photosynthesis. Photosynth Res 127:201–218. https://doi.org/10.1007/s11120-015-0177-y
Guyer A, De Vrieze M, Bönisch D et al (2015) The anti-phytophthora effect of selected potato-associated Pseudomonas strains: From the laboratory to the field. Front Microbiol 6:1309. https://doi.org/10.3389/fmicb.2015.01309
Hamonts K, Trivedi P, Garg A et al (2018) Field study reveals core plant microbiota and relative importance of their drivers. Environ Microbiol 20:124–140. https://doi.org/10.1111/1462-2920.14031
Hartmann M, Six J (2023) Soil structure and microbiome functions in agroecosystems. Nat Rev Earth Environ 4:4–18. https://doi.org/10.1038/s43017-022-00366-w
Holmes AJ, Bowyer J, Holley MP, O’Donoghue M, Montgomery M, Gillings MR (2000) Diverse, yet-to-be-cultured members of the Rubrobacter subdivision of the Actinobacteria are widespread in Australian arid soils. FEMS Microbiol Ecol 33(2):111–120. https://doi.org/10.1111/j.1574-6941.2000.tb00733.x
Huber KJ, Geppert AM, Wanner G, Fösel BU, Wüst PK, Overmann J (2016) The first representative of the globally widespread subdivision 6 Acidobacteria, Vicinamibacter silvestris gen. nov., sp. nov., isolated from subtropical savannah soil. IJSEM 66:2971–2979. https://doi.org/10.1099/ijsem.0.001131
Jacoby R, Peukert M, Succurro A, Koprivova A, Kopriva S (2017) The role of soil microorganisms in plant mineral nutrition—current knowledge and future directions. Front Plant Sci 8:1617. https://doi.org/10.3389/fpls.2017.01617
Jeanne T, Parent SÉ, Hogue R (2019) Using a soil bacterial species balance index to estimate potato crop productivity. PLoS ONE 14:e0214089. https://doi.org/10.1371/journal.pone.0214089
Jeong H, Choi SK, Ryu CM, Park SH (2019a) Chronicle of a soil bacterium: Paenibacillus polymyxa E681 as a tiny guardian of plant and human health. Front Microbiol 10:467. https://doi.org/10.3389/fmicb.2019.00467
Jeong S, Hong JK, Jho EH, Nam K (2019b) Interaction among soil physicochemical properties, bacterial community structure, and arsenic contamination: Clay-induced change in long-term arsenic contaminated soils. J Hazard Mater 378:120729. https://doi.org/10.1016/j.jhazmat.2019.06.006
Johnston-Monje D, Vergara LI, Lopez-Mejia J, White JF (2023) Plant microbiomes as contributors to agricultural terroir. Front Agron 5:1216520. https://doi.org/10.3389/fagro.2023.1216520
Joshi N, Gothalwal R, Singh M et al (2021) Novel sulphur-oxidizing bacteria consummate sulphur deficiency in oil seed crop. Arch Microbiol 203:1–6. https://doi.org/10.1007/s00203-020-02009-4
Kamilari E, Mina M, Karallis C, Tsaltas D (2021) Metataxonomic Analysis of Grape Microbiota During Wine Fermentation Reveals the Distinction of Cyprus Regional terroirs. Front Microbiol 12:726483. https://doi.org/10.3389/fmicb.2021.726483
Kim D, Song L, Breitwieser FP, Salzberg SL (2016) Centrifuge: rapid and sensitive classification of metagenomic sequences. Genome Res 26:1721–1729. https://doi.org/10.1101/gr.210641.116
Kim DH, Moon JY, Hong SY, Ahn H, Yoon YW, Kim H, et al (2023) Comparative Analysis of Microbial Community Characteristic of Acidic and Neutral Soils in Korean Orchards. Korean J Soil Sci 449–462. https://doi.org/10.7745/KJSSF.2023.56.4.449
Leontidou K, Genitsaris S, Papadopoulou A et al (2020) plant growth promoting rhizobacteria isolated from halophytes and drought-tolerant plants: genomic characterisation and exploration of phyto-beneficial traits. Sci Rep 10:14857. https://doi.org/10.1038/s41598-020-71652-0
Li WH, Liu QZ, Chen P (2018) Effect of long-term continuous cropping of strawberry on soil bacterial community structure and diversity. J Integr Agric 17(11):2570–2582. https://doi.org/10.1016/S2095-3119(18)61944-6
Liang JL, Liu J, Jia P et al (2020) Novel phosphate-solubilizing bacteria enhance soil phosphorus cycling following ecological restoration of land degraded by mining. ISME 14:1600–1613. https://doi.org/10.1038/s41396-020-0632-4
Lindsay WL, Norvell WA (1978) Development of a DTPA Soil Test for Zinc, Iron, Manganese, and Copper. Soil Sci Soc Am J 42:421–428. https://doi.org/10.2136/sssaj1978.03615995004200030009x
Liu A, Wang W, Chen X, Zheng X, Fu W, Wang G, Ji J, Guan C (2022) Phytoremediation of DEHP and heavy metals co-contaminated soil by rice assisted with a PGPR consortium: Insights into the regulation of ion homeostasis, improvement of photosynthesis and enrichment of beneficial bacteria in rhizosphere soil. Environ Pollut 314:120303. https://doi.org/10.1016/j.envpol.2022.120303
Liu B, Gu W, Yang Y et al (2021) Promoting potato as staple food can reduce the carbon–land–water impacts of crops in China. Nature Food 2:570–577. https://doi.org/10.1038/s43016-021-00337-2
Liu H, Hao Z, Yuan Y, Li C, Zhang J (2023) Application of mineral phosphorus fertilizer influences rhizosphere chemical and biological characteristics. Arch Agron Soil Sci 69(5):771–784. https://doi.org/10.1080/03650340.2022.2034792
Liu P, Jia S, He X, Zhang X, Ye L (2017) Different impacts of manure and chemical fertilizers on bacterial community structure and antibiotic resistance genes in arable soils. Chemosphere 188:455–464. https://doi.org/10.1016/j.chemosphere.2017.08.162
Loeppert RH, Suarez DL (1996) Carbonate and gypsum. In: Sparks DL, et al (ed) SSSA Book Series No. 5. SSSA and ASA, Madison, WI, pp 437–474
Losey NA, Stevenson BS, Busse HJ, Sinninghe Damsté JS, Rijpstra WIC, Rudd S, Lawson PA (2013) Thermoanaerobaculum aquaticum gen. nov., sp. nov., the first cultivated member of Acidobacteria subdivision 23, isolated from a hot spring. IJSEM 63(Pt_11). https://doi.org/10.1099/ijs.0.051425-0
Lu Y, Zhou G, Ewald J, Pang Z, Shiri T, Xia J (2023) MicrobiomeAnalyst 2.0: comprehensive statistical, functional and integrative analysis of microbiome data. Nucleic Acids Res 51:310–318. https://doi.org/10.1093/nar/gkad407
Lucini L, Rocchetti G, Trevisan M (2020) Extending the concept of terroir from grapes to other agricultural commodities: an overview. Curr Opin Food Sci 31:88–95. https://doi.org/10.1016/j.cofs.2020.03.007
Mahmud AA, Upadhyay SK, Srivastava AK, Bhojiya AA (2021) Biofertilizers: A Nexus between soil fertility and crop productivity under abiotic stress. Curr Res Environ Sustain 3:100063. https://doi.org/10.1016/j.crsust.2021.100063
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8. https://doi.org/10.1371/journal.pone.0061217
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 37:634–663. https://doi.org/10.1111/1574-6976.12028
Moustakas NK, Georgoulias F (2005) Soils developed on volcanic materials in the island of Thera, Greece. Geoderma 129:125–138. https://doi.org/10.1016/j.geoderma.2004.12.039
Nogi Y, Yoshizumi M, Hamana K, Miyazaki M, Horikoshi K (2014) Povalibacter uvarum gen. nov., sp. nov., a polyvinyl-alcohol-degrading bacterium isolated from grapes. IJSEM 64:2712–2717. https://doi.org/10.1099/ijs.0.062620-0
Parente CET, Brito EMS, Caretta CA et al (2021) Bacterial diversity changes in agricultural soils influenced by poultry litter fertilization. Braz J Microbiol 52:675–686. https://doi.org/10.1007/s42770-021-00437-y
Peiffer JA, Spor A, Koren O et al (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci USA 110:6548–6553. https://doi.org/10.1073/pnas.1302837110
Petrushin IS, Filinova NV, Gutnik DI (2024) Potato microbiome: Relationship with environmental factors and approaches for microbiome modulation. Int J Mol Sci 25:750. https://doi.org/10.3390/ijms25020750
Pfeiffer S, Mitter B, Oswald A, et al (2017) Rhizosphere microbiomes of potato cultivated in the High Andes show stable and dynamic core microbiomes with different responses to plant development. FEMS Microbiol Ecol 93. https://doi.org/10.1093/femsec/fiw242
Qin S, Yeboah S, Cao L et al (2017) Breaking continuous potato cropping with legumes improves soil microbial communities, enzyme activities and tuber yield. PLoS ONE 12:e0175934. https://doi.org/10.1371/journal.pone.0175934
Rousk J, Bååth E, Brookes PC et al (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351. https://doi.org/10.1038/ismej.2010.58
Santos-Medellín C, Edwards J, Liechty Z, et al (2017) Drought stress results in a compartment-specific restructuring of the rice root-associated microbiomes. mBio 8. https://doi.org/10.1128/mbio.00764-17
Saxena AK, Kumar M, Chakdar H et al (2020) Bacillus species in soil as a natural resource for plant health and nutrition. J Appl Microbiol 128:1583–1594. https://doi.org/10.1111/jam.14506
Segata N, Izard J, Waldron L et al (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:1–18. https://doi.org/10.1186/gb-2011-12-6-r60
Sessitsch A, Reiter B, Berg G (2004) Endophytic bacterial communities of field-grown potato plants and their plant-growth-promoting and antagonistic abilities. Can J Microbiol 50:239–249. https://doi.org/10.1139/w03-118
Shi S, Nuccio E, Herman DJ, et al (2015) Successional trajectories of rhizosphere bacterial communities over consecutive seasons. mBio 6. https://doi.org/10.1128/mbio.00746-15
Singh S, Singh SK, Chowdhury I, Singh R (2017) Understanding the Mechanism of Bacterial Biofilms Resistance to Antimicrobial Agents. Open Microbiol J 11:53. https://doi.org/10.2174/1874285801711010053
Singleton DR, Furlong,MA, Peacock AD, White DC, Coleman DC, Whitman WB (2003) Solirubrobacter pauli gen. nov., sp. nov., a mesophilic bacterium within the Rubrobacteridae related to common soil clones. IJSEM 53(Pt_11): 485–490. https://doi.org/10.1099/ijs.0.02438-0
Thomas GW (1982) Exchangeable Cations. Methods of Soil Analysis, Part 2, Chemical and Microbiological Properties. In: Page A, Miller R, Keeney D (eds) Agronomy, 2nd edn. American Society of Agronomy Inc., Madison, Wisconsin, USA, pp 159–165
Timmusk S, Grantcharova N, Wagner EG (2005) Paenibacillus polymyxa invades plant roots and forms biofilms. Appl Environ Microbiol 71:7292–7300. https://doi.org/10.1128/AEM.71.11.7292-7300.2005
Upadhyay SK, Chauhan PK (2022) Optimization of eco-friendly amendments as sustainable asset for salt-tolerant plant growth-promoting bacteria mediated maize (Zea Mays L.) plant growth, Na uptake reduction and saline soil restoration. Environ Res 211:113081. https://doi.org/10.1016/j.envres.2022.113081
Varliero G, Anesio AM, Barker GLA (2021) A Taxon-Wise Insight Into Rock Weathering and Nitrogen Fixation Functional Profiles of Proglacial Systems. Front Microbiol 12:627437. https://doi.org/10.3389/fmicb.2021.627437
Visser RGF, Bachem CWB, de Boer JM et al (2009) Sequencing the Potato genome: Outline and first results to come from the Elucidation of the sequence of the world’s third most important food crop. Am J Potato Res 86:417–429. https://doi.org/10.1007/s12230-009-9097-8
Wagg C, Bender SF, Widmer F, Van Der Heijden MGA (2014) Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci U S A 111:5266–5270. https://doi.org/10.1073/pnas.1320054111
Wang J, Song Y, Ma T, Raza W, Li J, Howland JG et al (2017) Impacts of inorganic and organic fertilization treatments on bacterial and fungal communities in a paddy soil. Appl Soil Ecol 112:42–50. https://doi.org/10.1016/j.apsoil.2017.01.005
Walkley AJ, Black IA (1934) Estimation of soil organic carbon by the chromic acid titration method. Soil Sci 37:29–38
Wang Y, Rashid MAR, Li X et al (2019) Collection and evaluation of genetic diversity and population structure of potato landraces and varieties in china. Front Plant Sci 10:139. https://doi.org/10.3389/fpls.2019.00139
Watanabe FS, Olsen SR (1965) Test of an Ascorbic Acid Method for Determining Phosphorus in Water and NaHCO3 Extracts from Soil. Soil Sci Soc Am J 29:677–678. https://doi.org/10.2136/sssaj1965.03615995002900060025x
Wei R, Ding Y, Chen N, et al (2022) Diversity and dynamics of microbial communities during spontaneous fermentation of Cabernet Sauvignon (Vitis vinifera L.) from different regions of China and their relationship with the volatile components in the wine. Food Res Intl 156:111372. https://doi.org/10.1016/j.foodres.2022.111372
Weinert N, Meincke R, Gottwald C, Heuer H, Schloter M, Berg G, Smalla K (2010) Bacterial diversity on the surface of potato tubers in soil and the influence of the plant genotype. FEMS Microbiol Ecol 74(1):114–123. https://doi.org/10.1111/j.1574-6941.2010.00936.x
Wu X, Cui Z, Peng J, Zhang F, Liesack W (2022) Genome-resolved metagenomics identifies the particular genetic traits of phosphate-solubilizing bacteria in agricultural soil. ISME Commun 16;2(1):17. https://doi.org/10.1038/s43705-022-00100-z
Wüst PK, Foesel BU, Geppert A, Huber KJ, Luckner M, Wanner G, Overmann J (2016) Brevitalea aridisoli, B. deliciosa and Arenimicrobium luteum, three novel species of Acidobacteria subdivision 4 (class Blastocatellia) isolated from savanna soil and description of the novel family Pyrinomonadaceae. IJSEM 66:3355–3366. https://doi.org/10.1099/ijsem.0.001199
Yang B, Wang Y, Qian PY (2016) Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis. BMC Bioinform 17:135. https://doi.org/10.1186/s12859-016-0992-y
Yoshinaka T, Yano K, Yamaguchi H (1973) Isolation of Highly Radioresistant Bacterium, Arthrobacter radiotolerans nov. sp. Agric Biol Chem 37(10): 2269–2275
Yu JL, Bing HJ, Chang RY, Cui YX, Shen GT, Wang XX, Zhang SP, Fang LC (2022) Microbial metabolic limitation response to experimental warming along an altitudinal gradient in alpine grasslands, eastern Tibetan Plateau. CATENA 214:1–10. https://doi.org/10.1016/j.catena.2022.106243
Zhalnina K, Louie KB, Hao Z, et al (2018) Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nature Microbiology 2018 3:4 3:470–480. https://doi.org/10.1038/s41564-018-0129-3
Zhang ZF, Liu LR, Pan YP et al (2023) Long-read assembled metagenomic approaches improve our understanding on metabolic potentials of microbial community in mangrove sediments. Microbiome 11:188. https://doi.org/10.1186/s40168-023-01630-x
Zhao F, Zhang Y, Dong W, Zhang Y, Zhang G, Sun Z, Yang L (2019) Vermicompost can suppress Fusarium oxysporum f. sp. lycopersici via generation of beneficial bacteria in a long-term tomato monoculture soil. Plant Soil 440:491–505. https://doi.org/10.1007/s11104-019-04104-y
Zhao X, Zhang Y, Cui Z, Peng L, Cao C (2022) Dynamics of phoD- and gcd- harboring microbial communities across an age sequence of biological soil crusts under sand-fixation plantation. Front Microbiol 13:831888. https://doi.org/10.3389/fmicb.2022.831888
Zhou X, Wang JT, Wang WH et al (2021) Changes in Bacterial and Fungal Microbiomes Associated with Tomatoes of Healthy and Infected by Fusarium oxysporum f. sp. lycopersici. Microb Ecol 81:1004–1017. https://doi.org/10.1007/s00248-020-01535-4
Zimudzi J, van der Waals JE, Coutinho TA et al (2018) Temporal shifts of fungal communities in the rhizosphere and on tubers in potato fields. Fungal Biol 122:928–934. https://doi.org/10.1016/j.funbio.2018.05.008
Acknowledgements
We wish to thank farmers of the Union of Agricultural Cooperatives of Naxos, Greece, for their support in soil collection.
Funding
Open access funding provided by HEAL-Link Greece. This research was funded by European Regional Development Fund (ERDF), through the Operational Program “Southern Aegean” 2014–2020, entitled “Enhancement of quality and nutritional traits of Naxos potatoes using omics-technologies. Acronym: GrEaTest-Potatoes” (0040991).
Author information
Authors and Affiliations
Contributions
Conceptualization: IM, IG; Methodology: AB, AX, GT, M-E Z, MV, I N–O; Investigation: IM, IG. Analysis: AB, AX, GT; Writing – Original Draft Preparation: AB, IM; Writing – Review & Editing: all.
Corresponding authors
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Martin Weih.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boutsika, A., Xanthopoulou, A., Tanou, G. et al. A microbiome survey of contrasting potato terroirs using 16S rRNA long-read sequencing. Plant Soil (2024). https://doi.org/10.1007/s11104-024-06686-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11104-024-06686-8