Abstract
Key message
The moss Pseudocrossidium replicatum is a desiccation-tolerant species that uses an inducible system to withstand severe abiotic stress in both protonemal and gametophore tissues.
Abstract
Desiccation tolerance (DT) is the ability of cells to recover from an air-dried state. Here, the moss Pseudocrossidium replicatum was identified as a fully desiccation-tolerant (FDT) species. Its gametophores rapidly lost more than 90% of their water content when exposed to a low-humidity atmosphere [23% relative humidity (RH)], but abscisic acid (ABA) pretreatment diminished the final water loss after equilibrium was reached. P. replicatum gametophores maintained good maximum photosystem II (PSII) efficiency (Fv/Fm) for up to two hours during slow dehydration; however, ABA pretreatment induced a faster decrease in the Fv/Fm. ABA also induced a faster recovery of the Fv/Fm after rehydration. Protein synthesis inhibitor treatment before dehydration hampered the recovery of the Fv/Fm when the gametophores were rehydrated after desiccation, suggesting the presence of an inducible protective mechanism that is activated in response to abiotic stress. This observation was also supported by accumulation of soluble sugars in gametophores exposed to ABA or NaCl. Exogenous ABA treatment delayed the germination of P. replicatum spores and induced morphological changes in protonemal cells that resembled brachycytes. Transcriptome analyses revealed the presence of an inducible molecular mechanism in P. replicatum protonemata that was activated in response to dehydration. This study is the first RNA-Seq study of the protonemal tissues of an FDT moss. Our results suggest that P. replicatum is an FDT moss equipped with an inducible molecular response that prepares this species for severe abiotic stress and that ABA plays an important role in this response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Due to climate change, rainfall frequency has decreased, and global temperatures have increased in recent years, resulting in an imbalanced ecosystem and hampering crop productivity (Fedoroff et al. 2010). It is estimated that the world population will reach more than 9 billion by 2050 (Godfray et al. 2010). Studies have suggested that a 70–100 % increase in food production is needed to feed the growing population (Godfray et al. 2010). Among all abiotic factors, drought is the environmental stressor with the highest global agronomic impact (Boyer 1982; Araus et al. 2002; Cramer et al. 2011). In only the past decade, global losses in crop production due to drought have totalled approximately $30 billion (Gupta et al. 2020). Therefore, there is a strong need to understand the mechanisms by which plants adapt to adverse environmental conditions in order to improve stress tolerance.
Drought causes water deficit (Bray 1997) and may even lead to desiccation, a condition in which bound water is the only water that remains in plant cells (Ramanjulu and Bartels, 2002). Although seeds of higher plants can withstand desiccation (Bewley 1979), the vegetative tissues of most plants do not tolerate a water content below 30–60 % (Challabathula and Bartels 2013; Zhang and Bartels 2018). Interestingly, in the course of evolution, some plants have developed desiccation tolerance (DT), which enables them to survive when free water is completely lost from the plant cells. Plants with DT can withstand complete desiccation for long periods and recover normal physiological status rapidly upon rehydration. A small number of vascular plant species have evolved to exhibit vegetative DT; these plants are known as resurrection plants Leprince and Buitink 2015; Giarola and Bartels 2015; Farrant et al. 2015; Zhang and Bartels 2018). Resurrection plants have been successfully explored as sources of genes with biotechnological potential to improve drought tolerance in transgenic plants (Villalobos et al. 2004; Garwe et al. 2006).
Bryophytes were the first plants to colonize terrestrial habitats, an event that required acquisition of unique physiological and molecular mechanisms to withstand fluctuating water availability. The ability to tolerate dehydration is widely distributed among bryophytes, but only a select group shows full DT. Several methods have been proposed to classify DT in mosses (reviewed in Wood 2007). The Austin protocol was proposed to generate a consensus methodology for DT classification in mosses (Wood 2007). This protocol classifies species with DT into two categories: (a) fully desiccation-tolerant (FDT) species, which include moss species that can survive both rapid and slow dehydration and can recover maximum efficiency of photosystem II (PSII) (Fv/Fm) values upon rehydration from a completely desiccated state, and (b) modified desiccation-tolerant (MDT) plants, which include species that survive only slow and moderate water loss (Wood 2007). The moss Tortula ruralis has been reported to be an FDT plant that has a constitutive mechanism for survival of desiccation (Oliver et al. 2000). Recently, other mosses, such as Pterygoneurum lamellatum (Stark et al. 2013), Syntrichia caninervis (Gao et al. 2014), and Bryum argenteum (Gao et al. 2015, 2017), have been reported to exhibit DT. Physcomitrium patens (formerly named Physcomitrella patens) has emerged as a model moss due to several characteristics, such as its ability to grow on simple liquid or solid media, its simple developmental pattern, its responsiveness to common phytohormones, its rapid cell cycle and growth rate, its ability to be transformed at high frequency and its ability to perform double recombination at rates similar to those shown by yeast (Cove 2005; Cove et al. 2006; Rensing et al. 2020). However, P. patens has been reported to be a mesic moss (Wood 2007) that cannot survive vegetative desiccation (Frank et al. 2005; Khandelwal et al. 2010; Koster et al. 2010; Komatsu et al. 2013). Additionally, this moss shows an inducible molecular response to dehydration that is mediated, at least in part, by the abiotic stress-related hormone abscisic acid (ABA) (Hiss et al. 2014; Ortiz-Ramírez et al. 2016; Perroud et al. 2018).
Increases in the levels of soluble sugars have been reported to occur under abiotic stress in vascular and nonvascular plants (Bhyan et al. 2012; Dong and Beckles 2019; Mayaba et al. 2001; Minami et al. 2005; Nagao et al. 2006; Cruz de Carvalho et al. 2015). During abiotic stress, soluble sugars play important roles in protecting membranes and macromolecules and in preventing enzymes from denaturing, and soluble sugars accumulate in vacuoles to enable osmotic adjustment (Nagao et al. 2006; Dinakar and Bartels 2013).
Furthermore, a molecular response occurs in response to abiotic stress, and the resulting gene expression tends to allow the plant to adapt to adverse conditions. ABA is a phytohormone that plays important roles in the regulation of many plant responses to different biotic and abiotic stresses and in plant developmental modulation (Finkelstein 2013; Humplík et al. 2017; Ma et al. 2018). The role of ABA in the regulation of the internal mechanisms in response to dehydration and other abiotic factors has been studied in mosses, mainly P. patens (Koster et al. 2010; Khandelwal et al. 2010; Takezawa et al. 2011, 2015; Saruhashi et al. 2015). Some core components of the angiosperm ABA-mediated transduction pathway are present in P. patens, indicating that some key genes of the pathway are conserved in bryophytes (Komatsu et al. 2020). The transcription factor ABI3, originally identified in Arabidopsis by screening for ABA-insensitive mutants (Koornneef et al. 1984), is also present in P. patens and plays a crucial role in the acquisition of the ABA-dependent DT of vegetative tissues of this moss (Khandelwal et al. 2010). In P. patens, a group A protein phosphatase type 2 C (PP2C), which is related to ABI1, acts as a negative regulator of ABA signalling (Sakata et al. 2009; Komatsu et al. 2009, 2013). Additionally, PpABI3 is involved in regulating the cold response and freezing tolerance in P. patens (Tan et al. 2017). However, bryophytes also express regulatory genes that are not present in vascular plants and are involved in the modulation of the ABA transduction pathway. The P. patens gene ANR is conserved in algae but absent in vascular plants and encodes a kinase required for DT (Stevenson et al. 2016). De novo transcriptome assembly has been performed for gametophores of DT moss species in response to dehydration; the examined species include S. caninervis (Gao et al. 2014) and B. argenteum (Gao et al. 2015, 2017). However, no studies have analysed the protonemata of a DT moss under conditions of dehydration using RNA-Seq.
Our group is interested in studying the moss P. replicatum (Taylor) R. H. Zander, which is widely distributed in the central highlands of Mexico and exhibits an outstanding ability to recover growth after fast desiccation. In the present work, we show that the moss P. replicatum is an FDT species harbouring an inducible and rapid molecular mechanism that prepares it for desiccation. This study represents the first RNA-Seq study of protonemal tissues of an FDT moss. We propose P. replicatum as a new model for the study of DT in mosses.
Materials and methods
Plant material
Samples of P. replicatum (Taylor) R. H. Zander were collected during the rainy season in a Juniperus deppeana forest in Ixtacuixtla, Tlaxcala, Mexico (19-20-03.3 °N, 98-21-59.9 °W, 2159 m.a.s.l.). For dehydration experiments, hydrated gametophores were collected in the field, maintained in a well-watered state during transportation to the laboratory and immediately used for experimental assays. For microscopy, desiccated samples from the National Herbarium of Mexico (MEXU) were used. A distribution map was generated using data from the MEXU records (http://www.ib.unam.mx/botanica/herbario/).
DT assay
The level of DT of P. replicatum was investigated according to the Austin protocol (Wood 2007). Briefly, freshly collected gametophores were carefully washed, and hydrated tissue samples (24 h at 90% relative humidity (RH), 20 °C) were equilibrated at two RH regimes (23% RH for fast dehydration and 67% RH for slow dehydration) for 1 or 7 d. As specified in the Austin protocol, we used saturated NH4NO3 and KCH3CO2 solutions to obtain 67% RH (− 54 MPa) and 23% RH (− 198 MPa), respectively. During dehydration, the maximum quantum yield (QY) of PSII was measured using a fluorometer (Fluorpen FP 100, PSI Instruments, Czech Republic). In dark-adapted samples, the QY is equivalent to the Fv/Fm. The samples were collected after the indicated durations of rehydration with distilled water, and the QY was measured in dark-adapted samples (10 min).
Effects of ABA on the dehydration kinetics and Fv/Fm values of P. replicatum gametophores
Freshly collected gametophores were carefully washed, and groups of hydrated samples (100 mg) were incubated in 50 ml conical Falcon tubes containing 5 ml of distilled water. Some samples were treated with ABA (10 or 100 µM) and incubated for 60 min with gentle shaking. Furthermore, samples were subjected to fast or slow dehydration by exposure to 23 or 67 % RH, respectively, as previously described. Before dehydration, the excess water was carefully removed with blotting paper. Water loss was calculated on a fresh-matter basis by recording the fresh weights of the samples until no further change was noted (Koster et al. 2010). The Fv/Fm values of the samples were recorded during the dehydration process.
Effects of protein synthesis or transcription inhibitors on the DT of P. replicatum gametophores
Essentially, a modified version of the Austin protocol was used. Gametophores were incubated with protein synthesis inhibitors or a transcription inhibitor. The inhibitors were applied before or after dehydration, and the recovery of the Fv/Fm was measured in rehydrated tissues. For application of inhibitors before dehydration, we used freshly collected gametophores that were maintained in a fully hydrated state at the laboratory. Individual groups of samples were pretreated for 24 h with different combinations of the translation inhibitors cycloheximide (300 µM, CHX), chloramphenicol (3 mM, CMP) (Proctor et al. 2007; Proctor and Smirnoff 2000), and α-amanitin (10 µM) (Rajjou et al. 2004) with or without ABA (10 µM) and further equilibrated to 67% RH for 24 h. Rehydrated samples were used to measure the Fv/Fm at 1 and 24 h. For application of inhibitors after dehydration, fully hydrated gametophores were equilibrated to 67% RH for 24 h. Furthermore, the dehydrated samples were rehydrated for 24 h with water to which different combinations of CHX, CMP, α-amanitin, and ABA had been added at the described concentrations. Finally, the PSII efficiency was measured at the indicated times.
Quantitative determination of total soluble sugars
For each treatment, 100 mg of freshly collected fully hydrated gametophores were incubated for 24 h in PpNH4 liquid medium supplemented with 400 mM sorbitol, 200 mM NaCl, or 10 µM ABA in an environmentally controlled room (photoperiod 16/8 h light/dark, 24 °C) with gentle shaking (100 rpm). The medium was discarded, and the excess liquid was removed carefully using absorbent paper. The samples were transferred to liquid nitrogen, pulverized, and homogenized at 0 °C in 5 ml of EtOH-H2O (4:1). The homogenate was centrifuged at 14,000×g for 5 min at 4 °C, and the supernatant was recovered. After evaporation of the ethanol, the samples were dissolved in distilled H2O (0.5 ml) and centrifuged at 14,000×g for 5 min at 4 °C to remove any debris. The soluble sugars in the supernatant were quantified with an anthrone-sulfuric acid assay using glucose as a standard (Yemm and Willis 1954; Komatsu et al. 2013).
Spore germination analysis and in vitro culture of P. replicatum
Axenic in vitro germination of P. replicatum spores was performed by superficial sterilization of P. replicatum sporophytes with sodium dichloroisocyanurate at 0.02% for 30 min followed by at least three washes with sterile water. Spores were released in 1 ml of sterile water with the aid of a needle, and serial dilutions were plated on PpNH4 solid medium. The germination of spores was scored every 24 h using a Nikon SMZ1500 stereomicroscope. Emerging protonemal cells with at least half the diameter of the spore were considered fully germinated. To obtain a monosporic pure line of P. replicatum, the resulting protonema from one individual spore was selected and cultured in PpNH4 medium under standard growth conditions in a growth room at 24 °C under a 16/8 h light/dark photoperiod with a light intensity of 55 µmol photons m− 2 s− 1. For analysis of the effect of ABA on protonema development, individual protonemal filaments were followed for 7 d using a Nikon TE300 inverted microscope with a 10× water immersion lens (Nikon).
RNA extraction for transcriptome analysis
For analysis of transcriptomes, total RNA was extracted with TRIzol reagent (Thermo Fisher Scientific, USA) from 7-d-old monosporic protonemal tissue, which was dehydrated by exposure to 23% RH (− 198 MPa) for 10 min (T1) or 30 min (T2). RNA from nondehydrated samples was extracted and used as a control (C). Equal amounts of total RNA obtained from three independent biological replicates were pooled for each treatment.
RNA-Seq library construction and sequencing
To compare the P. replicatum transcriptome under control versus dehydration conditions (T1 and T2), libraries were constructed for the different treatments using a TruSeq RNA Sample Preparation Kit following the manufacturer’s instructions (Illumina, Inc., San Diego, CA, USA), and single-end sequencing of the resulting libraries was carried out using an Illumina Genome Analyzer II. Briefly, poly(A)-tailed mRNA was enriched, fragmented, and used for first-strand cDNA synthesis. Subsequently, the second of strand cDNA was synthesized, and the final reactions were cleaned up before performing the end repair step. Furthermore, a single ‘A’ base was added to the 3´ ends of each fragment. Adapters were ligated to both ends of the short fragments, which were enriched by PCR to create the final double-stranded cDNA libraries. Finally, library quality control and quantification were performed with a Bioanalyzer Chip DNA 1000 series II (Agilent) and sequenced directly using a high-throughput Illumina HiSeq sequencing system (Illumina, Inc., San Diego, CA, USA). The raw reads for each library were deposited in the NCBI database, and the data can be downloaded from the Sequence Read Archive (SRA) under BioProject accession number PRJNA601777.
Global gene expression and differential gene expression analysis
To analyse the RNA-Seq data of P. replicatum, Trinity-v2.9.1 https://github.com/trinityrnaseq/trinityrnaseq/wiki programs were used for RNA-Seq de novo analysis, and the reference genome of P. patens (v3.3) was downloaded from the Phytozome database (Lang et al. 2018). First, quality analysis was performed with the program FastQC v0.11.8 (Andrews 2010), and the adapters were removed from the obtained reads with Trim Galore v0.6.6 with the default parameters (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Next, the reads were assembled using Trinity pipeline programs, and annotation was performed with Trinotate v3.1.1 (Bryant et al. 2017) and eggNOT v5.0 (Huerta-Cepas et al. 2019). Gene expression was quantified and evaluated using the Kallisto program (Bray et al. 2016). Genes with changes in their expression levels were selected via statistical testing with edgeR (Robinson et al. 2010), and the identified transcripts with reported counts were BLASTed with the reference genome of P. patens.
Results
P. replicatum is a moss widely distributed in Mexico
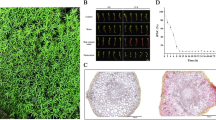
In the present work, our first aim was to obtain plant material with putative DT characteristics, specifically in the central highlands of Mexico. We identified several moss and other species with interesting characteristics; however, P. replicatum stood out and was selected for thorough characterization (Fig. 1a). P. replicatum is a moss belonging to the Pottiaceae family that grows on soil or rocks of calcareous origin and has green stems ranging from 3 to 15 mm in height with a central strand (Fig. 1d). The stem leaves are elliptic to ovate-lanceolate and 1–2 mm long with spirally revolute margins (Fig. 1c). The leaves are spirally appressed when dry (Fig. 1b). Interestingly, the photosynthetic system in P. replicatum leaves is represented by thin-walled cells in the spiral revolute leaf margins. This moss is widely distributed in Mexico (Fig. 1e) and can be found at different elevations ranging from 415 to 3600 m.a.s.l.
The species P. replicatum (Taylor) R. H. Zander is widely distributed in Mexico. The pictures show the appearance of P. replicatum gametophores when hydrated (a) or dehydrated (b). Transverse hand-cut sections of leaves (c) and stems (d) are also shown. According to the registers of MEXU, P. replicatum is widely distributed across Mexico (e) but is especially enriched in the central highland regions of the country
P. replicatum is an FDT moss
The Austin protocol (Wood 2007) was used to determine the level of DT exhibited by freshly collected gametophores. The gametophores were equilibrated for 7 d to water potentials of − 54 MPa (67% RH) and − 98 MPa (23% RH), and the Fv/Fm was undetectable under both conditions. Furthermore, the samples were watered, and the Fv/Fm values were evaluated after 1, 24 and 48 h of rehydration. Interestingly, despite such stress conditions, the Fv/Fm values were restored to values corresponding to 95% of the control levels after only 1 h of rehydration and showed Fv/Fm values higher than 0.7 only 2 h after rehydration (Fig. 2). Thus, the results clearly indicated that P. replicatum is an FDT moss that rapidly can recover its Fv/Fm to normal levels after prolonged exposure to 67% RH or 23 RH.
P. replicatum is an FDT moss. P. replicatum is an FDT moss that can recover its Fv/Fm to normal levels rapidly after prolonged exposure to 23% RH (black bars) or 67% RH (white bars). Totally hydrated gametophores prior to dehydration are indicated as controls for both groups of samples. The data shown are the means ± SDs from three independent experiments, with n = 5 at each indicated time
Effects of exogenous ABA on the dehydration and rehydration kinetics and Fv/Fm of P. replicatum gametophores
In P. patens and other mosses, ABA induces an increase in tolerance to diverse abiotic factors (Werner et al. 1991; Beckett et al. 2000; Beckett 2001; Minami et al. 2005; Nagao et al. 2005; Frank et al. 2005; Khandelwal et al. 2010; Koster et al. 2010; Komatsu et al. 2013). To study the effect of ABA on the dehydration kinetics of P. replicatum, freshly collected gametophores were subjected to fast or slow dehydration by exposure to 23% (− 198 MPa) or 67% RH (− 54 MPa), respectively, and the results were analysed using ANOVA. P. replicatum gametophores lost approximately 50% of their water content when exposed to a low-humidity atmosphere for only 30 min; however, at 67% RH, an exposure duration greater than 70 min was required to reach a similar percentage of water loss. ABA pretreatment slowed this dehydration process at both 23 RH and 67% RH (Fig. 3a). Under 23% RH conditions, the samples reached equilibrium after 80 min of dehydration; in contrast, the samples reached equilibrium after 180 min of dehydration under 67% RH. Additionally, we analysed the effect of ABA on the Fv/Fm values of P. replicatum gametophores exposed to dehydration under 23% RH and 67% RH atmospheres. ANOVA determined that a significantly different Fv/Fm response was observed between the two dehydration treatments (Fig. 3b). The Fv/Fm decayed rapidly at 23% RH, showing a drastic reduction after 30 min of treatment, but 60 min of treatment was required for the Fv/Fm values to become undetectable. Lower Fv/Fm values were recorded under the same conditions when the gametophores were pretreated with ABA after 40 min of dehydration. On the other hand, high oscillatory values of Fv/Fm were obtained under slow-dehydration conditions, even after 80 min of dehydration; ultimately, more than 180 min of dehydration was required for the Fv/Fm values to become undetectable. In contrast, when gametophores pretreated with 100 µM ABA were subjected to slow dehydration, the Fv/Fm became undetectable in 140 min. For both dehydration conditions analysed, we observed faster recovery of Fv/Fm values during the first 20 min of rehydration when ABA pretreatment was applied to the gametophores before dehydration than when ABA pretreatment was not applied (Fig. 3c). Notably, the Fv/Fm values of P. replicatum gametophores that were previously exposed to − 54 MPa or − 198 MPa were able to recover to 52 and 80 % of the control Fv/Fm values, respectively, after only 10 min of rehydration. These data suggest that ABA plays a role in the responses of PSII during the dehydration of P. replicatum.
Effects of ABA on dehydration kinetics and the Fv/Fm during the dehydration/rehydration process of gametophores of P. replicatum. a P. replicatum gametophores lost their water content when exposed to a low-humidity atmosphere (23% RH, solid symbols), but ABA pretreatment significantly diminished the final water loss after equilibrium was reached at this humidity level (ANOVA, p value = 0.05). The results obtained when gametophores were exposed to 67% RH are shown as open symbols. b P. replicatum gametophores maintained high Fv/Fm values for up to two hours during a slow dehydration process; however, ABA pretreatment induced a decrease in the Fv/Fm. c ABA pretreatment before dehydration resulted in a faster recovery of the Fv/Fm during the first 20 min of the rehydration process. Each point on the graph represents the average from three independent experiments, each with n = 5
Analysis of the effects of inhibitors of transcription and translation on the recovery of PSII efficiency after dehydration
To investigate whether the gametophores of P. replicatum require the expression of genes induced during dehydration to recover photosynthesis upon rehydration, a modified Austin protocol was conducted. α-Amanitin was used as an inhibitor of transcription, whereas CHX and CMP were used as inhibitors of protein synthesis. The inhibitors were applied independently before dehydration or during rehydration (Fig. 4), and the Fv/Fm values were measured after 1 and 24 h. The Fv/Fm values were analysed using ANOVA. Interestingly, an important delay in Fv/Fm recovery was observed in response to translational inhibitor treatment. Lower values of Fv/Fm were observed when CMP was applied before dehydration (Fig. 4a) than when inhibitors were applied during rehydration (Fig. 4b). Additionally, application of exogenous ABA together with the inhibitors was not enough to restore the Fv/Fm recovery timing, indicating that the DT exhibited by P. replicatum gametophores requires de novo protein synthesis, which could help to protect the photosynthetic apparatus from damage provoked by stress. α-Amanitin was not observed to have a clear effect under the tested conditions.
Inhibitor treatment reveals the participation of chloroplastic proteins required for the optimal function of PSII after dehydration. a Fv/Fm values obtained when inhibitors were applied before dehydration. b Fv/Fm responses obtained when inhibitors were applied during rehydration. Lower values of Fv/Fm were observed when CMP was applied before dehydration than when inhibitors were applied during rehydration. Each point on the graph represents the average ± SD from three independent experiments, each with n = 5. Samples showing significant differences according to ANOVA are indicated by asterisks (p value = 0.05)
Effect of abiotic stress on the accumulation of soluble sugars in P. replicatum gametophores
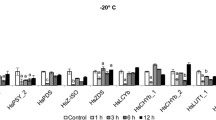
During abiotic stress, soluble sugars play important roles in protecting membranes and macromolecules and in preventing enzymes from denaturing, and soluble sugars accumulate in vacuoles to enable osmotic adjustment (Nagao et al. 2006; Dinakar and Bartels 2013). Experiments were conducted to determine whether the accumulation of soluble sugars is part of the strategy of the gametophores of P. replicatum to withstand abiotic stress. Our results indicated that application of 10 µM ABA or 200 mM NaCl induced strong accumulation of soluble sugars, resulting in levels nearly four times higher than control levels (Fig. 5). On the other hand, a twofold increase in soluble sugars was observed when gametophores were treated with 400 mM sorbitol. These observations indicate that ABA stimulation and NaCl stimulation are sufficient to induce accumulation of soluble sugars in P. replicatum gametophores and that moderate osmotic stress also induces sugar accumulation.
The gametophores of P. replicatum accumulate soluble sugars in response to ABA and abiotic stress factors. For each treatment, 100 mg of freshly collected fully hydrated gametophores were incubated for 24 h in PpNH4 liquid medium supplemented with 400 mM sorbitol, 200 mM NaCl, or 10 µM ABA. The soluble sugars were quantified with an anthrone-sulfuric acid assay. The bars represent the averages ± SDs from three independent experiments
Effects of exogenous ABA on spore germination and protonemal development
ABA is a phytohormone that also plays a role in spore germination and protonemal development (Vesty et al. 2016; Decker et al. 2006; Takezawa et al. 2011; Stevenson et al. 2016; Shinozawa et al. 2019; Arif et al. 2019). A putative effect of exogenous ABA on spore rate germination in P. replicatum was investigated. Compared with the control, all ABA concentrations tested (5–50 µM) delayed the germination kinetics (Fig. 6a). However, the highest concentration used (50 µM) not only delayed germination but also diminished the germination rate to approximately 50 %. The effect of exogenous ABA on protonemal development was also investigated by observing individual protonemal filaments for 7 d under a microscope. Application of ABA induced clear morphological changes in the protonemal cells of P. replicatum (Fig. 6b and c). While the apical cells maintained polarized growth in response to ABA, the nonapical cells of protonemal filaments acquired a small and rounded phenotype with suppressed cell expansion in response to ABA, resembling brachycytes (brood cells).
Effects of exogenous ABA on P. replicatum spore germination and protonemal development. a Each point on the graph represents the average ± SD from three independent experiments, each with n = 120–150. Additionally, an individual protonemal filament from a monosporic line was used to analyse the influence of ABA (10 µM) on protonemal development, where b is t0 and c corresponds to 7 d of exposure
RNA-Seq analysis of protonemata in response to dehydration
To investigate the potential existence of a dehydration-inducible system at the transcriptional level, a preliminary analysis of the transcriptome changes of P. replicatum in response to dehydration was conducted. The transcriptomes of protonemal samples dehydrated at 23% RH (− 198 MPa) for 10 or 30 min were compared with those of fully hydrated samples. These specific dehydration times were chosen because analysis of the dehydration kinetics of P. replicatum protonemata exposed to a 23 % RH atmosphere indicated that 10 and 30 min of treatment caused water content decreases of 30 and 70%, respectively (Supplementary file 1a). A total of 1368 genes were identified to exhibit expression changes in protonemal samples dehydrated for 10 min (T1). However, the number of genes with expression changes increased to 4220 (T2) when dehydration was extended to 30 min. A total of 1973 differentially expressed genes (DEGs) were identified using statistical analysis (Supplementary file 3); 972 corresponded to upregulated transcripts, and 821 corresponded to downregulated transcripts. When the upregulated and downregulated genes for all treatments (T1 + T2) were considered together, annotation analysis identified 175 DEGs with homology to P. patens sequences (Supplementary file 1b-A). Only 9 DEGs were detected in T1 in comparison with the hydrated control (C) (Supplementary file 1b-A). However, the number of DEGs increased to 118 when the dehydration treatment was prolonged to 30 min (Supplementary file 1b-A). Interestingly, 137 DEGs were found in T2 compared to T1, 86 of which were also DEGs in T2 compared with C (Supplementary file 1b-A). In total, 154 upregulated genes were identified in T1 + T2 compared with C, and the treatment with the most abundant upregulated genes was T2 (Supplementary file 1b-B). In contrast, a total of 27 genes were downregulated in T1 + T2 in comparison with C (Supplementary file 1b-C), and 21 of these genes were downregulated specifically in T2 (Supplementary file 1b-C). An analysis of only the nonredundant DEGs that were related to P. patens identified a total of 89 DEGs, 30 of which were shared between T1 and T2 (Supplementary file 1b-D; Supplementary file 2). A complete list of DEGs is provided in Supplementary file 3. To further analyse the obtained data, the 2-kb regions corresponding to the promoters of all DEGs were analysed using the Plant Cis-Acting Regulatory Elements (PlantCARE) database (Lescot et al. 2002), and we observed the presence of several ABRE and DRE motifs in the promoters of ABA- and drought-responsive genes (Supplementary file 1c). Furthermore, a Gene Ontology (GO) enrichment analysis was performed to compare and plot the induced and repressed genes based on a GO standardized directed acyclic graph (DAG) structured vocabulary system (Supplementary file 1d). Prediction of the subcellular localization of the products of the induced and repressed genes using CELLO2GO (Yu et al. 2014) revealed that chloroplastic protein-encoding genes were the most abundant group among the induced genes (28 %); in contrast, nuclear protein-encoding genes were the most abundant group among the repressed genes (31.8 %) (Supplementary file 1e-A and Supplementary file 1e-B, respectively).
Discussion
Anatomical and ecological characteristics of P. replicatum
Among plants collected from the central highlands of Mexico, we identified the moss P. replicatum, which stood out as an FDT plant species and was selected for further characterization. The Mexican central highlands are surrounded by three volcanoes, one of which is still active (Popocatepetl). In Mexico, this plant species has a broad distribution up to 3,200 m.a.s.l. (Fig. 1e) but is enriched in desert areas and forests in high, mountainous regions with cold weather. In the region of sample collection, the average annual minimum and maximum temperatures are 1.5 and 25 °C, respectively. The estimated annual precipitation is 700 mm, concentrated mainly from June to September, and the dry season spans several months. The leaves of P. replicatum adopt a spiral arrangement around the stem during the dehydration process, becoming tightly entwined when the desiccated state is reached. The reason for the evolution of this response is not known, but we suggest that this phenomenon may help P. replicatum maintain its internal water content by diminishing the rate of water loss during dehydration. Recently, the chloroplastic and mitochondrial genomes of P. replicatum were reported Cevallos et al. 2019, 2020, respectively). A phylogenetic analysis using 16 chloroplast protein-coding genes demonstrated that P. replicatum is a member of the Pottiaceae family and is closely related to Syntrichia ruralis (Cevallos et al. 2019).
P. replicatum is an FDT moss
We classified P. replicatum as an FDT moss. After plants were equilibrated for 7 d to water potentials of − 54 MPa (67% RH) and − 198 MPa (23% RH), the Fv/Fm values of the gametophores was undetectable. However, gametophores of P. replicatum recovered normal Fv/Fm levels after rehydration regardless of whether they were desiccated at − 54 MPa or at − 198 MPa, although a faster recovery was observed for the slower dehydration treatment. This observation might suggest that to fully recover the Fv/Fm, P. replicatum gametophores must activate an inducible molecular mechanism to quickly repair the damage generated during desiccation. T. ruralis, the most studied DT moss, has been demonstrated to employ a constitutive mechanism that helps it withstand desiccation (reviewed in Oliver and Bewley 1997; Oliver and Wood 1997; Oliver et al. 1997; Wood et al. 2000). However, T. ruralis also harbours an inducible molecular mechanism that is activated only in the rehydration stage and is dedicated to repair of membrane damage, fragmentation of vacuoles and chloroplasts, depolymerization of microtubules, and denaturation of proteins, among other alterations (Oliver and Bewley 1997; Oliver and Wood 1997; Pressel and Duckett 2010). Other mosses seem to have different strategies to withstand abiotic stress. Although P. patens is not considered to be a DT moss with a constitutive mechanism of protection from desiccation (Pressel and Duckett 2010; Koster et al. 2010), it has been shown that this moss has an inducible molecular mechanism that is activated in response to cold (Beike et al. 2015), dehydration (Koster et al. 2010), and ABA and salt treatments (Khraiwesh et al. 2015). Recently, it has been reported that 6 and 9 d of acclimation to 69 % RH are required to induce DT at 33 % RH in protonemal and gametophore tissues of P. patens, respectively (Rathnayake et al. 2019). The results obtained using the Austin protocol indicate that P. replicatum is an FDT moss that is able to recover photosynthesis after rapid and slow dehydration.
ABA plays roles in the dehydration process and in the efficiency of PSII in P. replicatum
To gain insight into the possible existence of an inducible molecular mechanism in P. replicatum that is activated in response to abiotic stress, we first investigated whether ABA plays a role during dehydration in P. replicatum. Our results indicate that ABA plays a crucial role in P. replicatum responses during dehydration of gametophores. Water loss was slightly reduced when P. replicatum gametophores were pretreated with ABA before dehydration (Fig. 3a). Similar responses have been reported for P. patens (Koster et al. 2010); ABA pretreatment also increases water retention in this moss during drying at − 52 MPa (68% RH). In contrast, the dehydration kinetics of the moss Funaria hygrometrica are not significantly altered by pretreatment with ABA (Werner et al. 1991). However, in other bryophytes such as the liverwort Marchantia polymorpha, ABA also decreases the rate of water loss in gametophytes under dehydration (Pence et al. 2005). More studies are needed to understand the different effects of ABA on the dehydration kinetics of bryophyte tissues at several developmental stages. The effect of ABA on photosynthesis was also evaluated in the current study (Fig. 3b). Application of exogenous ABA induced a strong decrease in PSII activity, which was evident when gametophores were subjected to slow dehydration (67% RH). Under controlled conditions, during both rapid and slow dehydration, the QY values became undetectable only after the water content had decreased to approximately 20%. This water content value was reached in 60 and 180 min under rapid and slow dehydration, respectively. However, when ABA pretreatment was applied, a decrease in QY was observed, especially under the slow dehydration treatment. This observation is in accordance with the responses reported for the mosses Atrichum undulatum and Atrichum androgynum, in which ABA pretreatment reduces several photosynthetic parameters (Beckett 2001; Mayaba et al. 2001). In diverse plant models, such as Arabidopsis and Craterostigma, it has been established that ABA participates in the induction of dehydration and DT (Finkelstein 2013; Sah et al. 2016; Vishwakarma et al. 2017). In bryophytes such as P. patens and F. hygrometrica, the role of ABA in acquisition of tolerance to abiotic stress has been demonstrated by evidence such as visible cellular phenotypes, plasmolysis, and growth (Werner et al. 1991; Minami et al. 2005; Khandelwal et al. 2010; Bhyan et al. 2012; Stevenson et al. 2016). Interestingly, in the present study, P. replicatum gametophores required only 1 h of pretreatment with ABA to show accelerated Fv/Fm recovery upon rehydration of desiccated gametophores, which occurred at least during the first 20 min (Fig. 3c); these findings demonstrate the role of ABA in the rapid photoprotection and/or repair of PSII in mosses during rehydration. Moreover, the results obtained using inhibitors of transcription and translation suggest that the de novo expression of chloroplastic proteins is very important for the protection and repair of PSII during the P. replicatum dehydration/rehydration response (Fig. 4). Altogether, these observations suggest that ABA induces a molecular response in P. replicatum to rapidly prepare this moss to withstand desiccation stress. Thus, once equipped with dehydration- or ABA-induced expression of a set of protective or reparative enzymes, this moss can readily and efficiently respond to dehydration stress.
ABA plays a role in the accumulation of soluble sugars in P. replicatum in response to abiotic stress
Increases in soluble sugars have been reported to occur under abiotic stress in vascular and nonvascular plants (Bhyan et al. 2012; Dong and Beckles 2019; Mayaba et al. 2001; Minami et al. 2005; Nagao et al. 2006; Cruz de Carvalho et al. 2015). During abiotic stress, soluble sugars play important roles in protecting membranes and macromolecules and in preventing enzymes from denaturing, and they accumulate in vacuoles for osmotic adjustment (Nagao et al. 2006; Dinakar and Bartels 2013). Our results indicated that the gametophores of P. replicatum accumulated soluble sugars in response to 10 µM ABA treatment and in response to moderate osmotic stress (400 mM sorbitol) and salt stress (200 mM NaCl) (Fig. 5). No alterations in the QY values were observed under such conditions (data not shown). Previous reports on vascular and nonvascular plants have indicated that application of exogenous ABA induces accumulation of soluble sugars (Orr et al. 1986; Bravo et al. 1998; Bhyan et al. 2012; Mayaba et al. 2001; Nagao et al. 2005, 2006; Akter et al. 2014; Takezawa et al. 2015). Interestingly, in P. replicatum, application of exogenous ABA induced accumulation of soluble sugars to an extent comparable to that induced by NaCl. This finding could suggest that NaCl-induced accumulation of soluble sugars is mediated by ABA in P. replicatum. Moderate osmotic stress also induced sugar accumulation, possibly via an endogenous ABA-dependent pathway. Another possible explanation is that both NaCl and osmotic stress activated similar response pathways in an ABA-independent manner. Whether P. replicatum changes its internal ABA content in response to NaCl or osmotic stress is a main question that we will address in the near future. Finally, we speculate that the observed decrease in the rate of water loss during dehydration (Fig. 3a) and the further increase in the velocity of QY recovery after rehydration (Fig. 3b) may be due, at least in part, to the ABA-mediated accumulation of soluble sugars in the gametophores of P. replicatum.
ABA affects spore germination and protonemal development in P. replicatum
ABA is a phytohormone that participates in plant development and in responses to different biotic and abiotic stresses (Humplík et al. 2017; Ma et al. 2018). To address the role of ABA during the early P. replicatum developmental stages, the effects of ABA on spore germination and protonemal growth were analysed. Spore germination of P. replicatum was less sensitive to ABA than seed germination of vascular plants (Fig. 6a). In Arabidopsis, it is well known that low ABA concentrations ranging from 3 to 5 µM are enough to completely inhibit seed germination (Finkelstein 1994; Arenas-Huertero et al. 2000). However, up to 50 µM ABA was required to inhibit 50 % of the spore germination of P. replicatum (Fig. 6a). Two important observations contradicted previous findings regarding P. patens spore germination (Vesty et al. 2016). The sensitivity of P. patens spores to ABA seems to be higher than that of P. replicatum spores because only 2 µM ABA is sufficient to significantly delay P. patens spore germination. In this study, we found that P. replicatum exhibited a better germination rate than P. patens. On the other hand, our protonema assay revealed that after 7 d of ABA treatment, the protonemata showed a phenotype characterized by cell greening, decreased cell length, and a round shape, consistent with the brood cell phenotype commonly observed to develop in P. patens in response to ABA (Fig. 6b, c) (Decker et al. 2006; Takezawa et al. 2011; Stevenson et al. 2016; Shinozawa et al. 2019; Arif et al. 2019); these results indicate that P. replicatum apical cells retain polar growth under such conditions. These findings support early ABA participation in bryophytes during land conquest and vegetative dispersion since brood cells function as dormant structures.
P. replicatum transcriptome changes in response to dehydration
Recently, de novo transcriptome assembly has been performed for gametophores of DT mosses, such as S. caninervis (Gao et al. 2014) and B. argenteum (Gao et al. 2015, 2017), under conditions of dehydration. In all three reports gametophores were tested for dehydration, and further RNA-Seq strategies were performed. However, no work has used RNA-Seq to analyse the protonemata of a DT moss in response to dehydration. To preliminarily investigate the transcriptomic response to a low-water regime in P. replicatum protonemata, we assessed the effects of fast dehydration of protonemal tissues at 23% RH over short time periods: 10 min (T1) and 30 min (T2). The RNA-Seq analyses revealed that compared with genes in the control condition, 1368 genes at T1 and 4220 genes at T2 were differentially expressed. Bioinformatics analysis of the DEGs reduced the numbers to 972 upregulated and 821 downregulated DEGs. Of these, a total of 175 DEGs that were homologous with P. patens sequences were significant when both treatments were considered together (T1 + T2), and the most abundant were upregulated genes. Consistent with the scenario in P. replicatum, the mosses S. caninervis and B. argenteum show an inducible molecular response to dehydration. However, the inducible molecular mechanism used by P. replicatum is completely different from that used by the moss T. ruralis, another FDT plant that uses a constitutive mechanism to survive desiccation (Oliver et al. 2000) but uses an inducible mechanism during rehydration (Oliver et al. 2005). This mechanism also seems to be partially present in B. argenteum (Gao et al. 2017). It could be interesting to investigate the molecular response of P. replicatum during the process of rehydration; future studies will address this topic. Nuclear and chloroplastic protein-encoding genes were the most abundant types of genes among the induced genes, accounting for more than 50% of the genes; in contrast, nuclear protein-encoding genes were the most abundant proteins among the repressed genes (31.8%). Downregulated transcripts encoding chloroplastic proteins accounted for 18.2% of the genes; among them, genes encoding important proteins required for assembly of the PSII reaction centre, such as PsbN (Torabi et al. 2014), several photosynthetic electron transfer B proteins (Goltsev et al. 2021), and PHY1 light-sensor protein kinase (Possart and Hiltbrunner 2013), were identified. Notably, inhibition of photosynthesis is a primary central response that is exhibited by both desiccation-tolerant and desiccation-sensitive plants affected by drought stress to protect the photosynthetic apparatus (Challabathula et al. 2018).
Desiccation-tolerant plants employ both conserved and novel antioxidant enzymes/metabolites to minimize oxidative damage and to protect the photosynthetic machinery. De novo-synthesized stress-induced proteins, in combination with antioxidants, are important components of the protective network. Interestingly, transcripts encoding proteins associated with photosynthesis (such as thiamine thiazole synthase; the early light inducible protein ELIP1; light-harvesting PSII subunits; the MPH1 protein, which is involved in Maintenance of PSII under High light conditions; ribulose biphosphate carboxylase small chain; chlorophyll a-/b-binding proteins; plastocyanin; and PSI-F), general/sugar metabolism (enolase, fructose-bisphosphate aldolase, D-fructose-1,6-bisphosphate-1-phosphohydrolase, phosphopyruvate hydratase, phosphoglycerate kinase, malate dehydrogenase, transketolase, and xyloglucan endotransglucosylase/hydrolase), and translation (the 60 S acidic ribosomal protein P0; the 40 S ribosomal proteins S3a, S25, and S21; peptidyl-prolyl alpha-trans isomerase; 5-methyltetrahydropteroyltriglutamate-homocysteine S-methyltransferase; the 60 S ribosomal protein L7a; and elongation factor 1-alpha), among other processes were found upregulated. Thus far, studies have revealed that most of these genes participate in responses to abiotic stress in plants either by expression analysis or forward and reverse genetics Bernacchia et al. 1995; Laxalt et al. 1996; Haldrup et al. 2000; Sharma et al. 2003; Morita-Yamamuro et al. 2004; Barkla et al. 2009; Lu et al. 2012; Omidbakhshfard et al. 2012; Hertz et al. 2013; Wang et al. 2013; Gururani et al. 2015; Liu et al. 2015; Liu and Last 2015; Piro et al. 2015; Cai et al. 2016; Joshi et al. 2016; Li et al. 2016; Yasmenn et al. 2016; Wang et al. 2016; Giarola et al. 2017; Zhou et al. 2018; Li et al. 2019; Hasanuzzaman et al. 2020; Zhao et al. 2020). During abiotic stress, proteins can be damaged, and turnover of specific proteins is required. Thus, upregulation of transcripts encoding proteins related to translation could be a strategy of P. replicatum to adapt to dehydration.
On the other hand, transcripts related to osmoprotection, oxidative stress, protein integrity/turnover (protection, repair, degradation), and signalling also accumulated during drying but were more closely related to Arabidopsis, wheat, and other plant species than P. patens (germin-like proteins, several LEA proteins, the early response to dehydration protein ERD15, aquaporins, cobalamin synthase, cytochrome c oxidase, an HR-like lesion-inducing protein-related protein, plasma membrane H+-ATPases, ubiquitin, polyubiquitin, the glycine cleavage system P protein, alpha/beta-hydrolases, Shaggy-like kinase 13, serine/threonine kinases, and basic region leucine zipper transcription factors, among others).
Finally, using the PlantCARE database, we evaluated the promoter sequences of all upregulated and downregulated genes. We found 27 promoters enriched with ABRE and DRE boxes, which are cis-elements that bind to transcription factors in response to ABA and abiotic stress, respectively (Guiltinan et al. 1990; Yamaguchi-Shinozaki and Shinozaki 1994). The ABRE sequence was the most abundant motif in both induced and repressed genes, supporting the idea of an important role for ABA during the P. replicatum dehydration response. The upregulated genes whose promoters were enriched mainly with ABRE boxes encoded proteins involved in the photosynthetic process. These observations support the idea that dehydration induces a rapid change in the expression profile of P. replicatum to readjust metabolism so that the organism can respond efficiently to stress. Further analyses of the molecular responses of P. replicatum during different developmental stages and under other conditions of dehydration will help elucidate the molecular strategy of this DT moss. Nevertheless, this study represents the first RNA-Seq study on protonemal tissues of an FDT moss. Altogether, our results indicate that P. replicatum is an FDT moss that exhibits an inducible mechanism in response to abiotic stress and ABA. This work provides a basis for ongoing studies on the functional genomics of P. replicatum as a plant model for the study of DT.
References
Akter K, Kato M, Sato Y, Kaneko Y, Takezawa D (2014) Abscisic acid-induced rearrangement of intracellular structures associated with freezing and desiccation stress tolerance in the liverwort marchantia polymorpha. J Plant Physiol 171:1334–1343. https://doi.org/10.1016/j.jplph.2014.05.004
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Araus JL, Slafer GA, Reynolds MP, Royo C (2002) Plant breeding and drought in C3 cereals: what should we breed for? Ann Bot 89:925–940. https://doi.org/10.1093/aob/mcf049
Arenas-Huertero F, Arroyo A, Zhou L, Sheen J, León P (2000) Analysis of Arabidopsis glucose insensitive mutants gin5 and gin6 reveals a central role of the plant hormone ABA in the regulation of plant vegetative development by sugar. Genes Dev 14:2085–9096. DOI:https://doi.org/10.1101/gad.14.16.2085
Arif MA, Hiss M, Tomek M, Busch H, Meyberg R, Tintelnot S, Reski R, Rensing S, Frank W (2019) ABA-induced vegetative diaspore formation in Physcomitrella patens. Front Plant Sci 10:1–18. https://doi.org/10.3389/fpls.2019.00315
Barkla BJ, Vera-Estrella R, Hernández-Coronado M, Pantoja O (2009) Quantitative proteomics of the tonoplast reveals a role for glycolytic enzymes in salt tolerance. Plant Cell 21:4044–4058. https://doi.org/10.1105/tpc.109.069211
Beckett RP, Csintalan Z, Tuba Z (2000) ABA treatment increases both the desiccation tolerance of photosynthesis, and nonphotochemical quenching in the moss Atrichum undulatum. Plant Ecol 151:65–71. https://doi.org/10.1023/A:1026546707649
Beckett RP (2001) ABA-induced tolerance to ion leakage during rehydration following desiccation in the moss Atrichum androgynum. Plant Growth Regul 35:131–135. https://doi.org/10.1023/A:1014477603983
Beike AK, Lang D, Zimmer AD, Wiist F, Trautmann D, Wiedemann G, Beyer P et al (2015) Insights from the cold transcriptome of Physcomitrella patens: global specialization pattern of conserved transcriptional regulators and identification of orphan genes involved in cold acclimation. New Phytol 205:869–881. https://doi.org/10.1111/nph.13004
Bernacchia G, Schwall G, Lottspeich F, Salamini F, Bartels D (1995) The transketolase gene family of the resurrection plant Craterostigma plantagineum: differential expression during the rehydration phase. EMBO J 14:610–618. https://doi.org/10.1002/j.1460-2075.1995.tb07037.x
Bewley JD (1979) Physiological aspects of desiccation tolerance. Annu Rev Plant Phys 30:195–238. https://doi.org/10.1146/annurev.pp.30.060179.001211
Boyer JS (1982) Plant productivity and environment. Science 2018:443–448. https://doi.org/10.1126/science.218.4571.443
Bhyan BS, Minami A, Kaneko Y, Suzuki S, Arakawa K, Sakata Y, Takezawa D (2012) Cold acclimation in the moss Physcomitrella patens involves abscisic acid-dependent signaling. J Plant Physiol 169:137–145. https://doi.org/10.1016/j.jplph.2011.08.004
Bravo LA, Zúñiga GE, Alberdi M, Corcuera LJ (1998) The role of ABA in freezing tolerance and cold acclimation in barley. Physiol Plant 103:17–23. https://doi.org/10.1034/j.1399-3054.1998.1030103.x
Bray E (1997) Plant responses to water deficit. Trends Plant Sci 2:48–54. https://doi.org/10.1016/S1360-1385(97)82562-9
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34:525–527. https://doi.org/10.1038/nbt.3519
Bryant DM, Johnson K, DiTommaso T, Tickle T, Couger MB, Payzin-Dogru D, Lee TJ, Leigh ND, Kuo T-H, Davis FG, Bateman J, Bryant S, Guzikowski AR, Tsai SL, Coyne S, Ye WW, Freeman RM, Peshkin L, Tabin CJ, Regev A, Haas BJ, Whited JL (2017) A tissue-mapped axolotl de novo transcriptome enables identification of limb regeneration factors. Cell Reports 18:762–776. https://doi.org/10.1016/j.celrep.2016.12.063
Cai B, Li Q, Xu Y, Yang L, Bi H, Ai X (2016) Genome-wide analysis of the fructose 1,6-bisphosphate aldolase (FBA) gene family and functional characterization of FBA7 in tomato. Plant Physiol Biochem 108:251–265. https://doi.org/10.1016/j.plaphy.2016.07.019
Cevallos MA, Guerrero G, Ríos S, Arroyo A, Villalobos MA, Porta H (2019) The chloroplast genome of the desiccation-tolerant moss Pseudocrossidium replicatum (Taylor) R.H. Zander. Genet Mol Biol 42:488–493. https://doi.org/10.1590/1678-4685-GMB-2018-0184
Cevallos MA, Guerrero G, Ríos S, Arroyo A, Villalobos MA, Porta H (2020) The mitogenome of Pseudocrossidium replicatum, a desiccation-tolerent moss. Mitochondrial DNA Part B 5(3):2339–2341. https://doi.org/10.1080/23802359.2020.1774436
Challabathula D, Bartels D (2013) Desiccation tolerance in resurrection plants: new insights from transcriptome, proteome and metabolome analysis. Front Plant Sci 4:482. https://doi.org/10.3389/fpls.2013.00482
Challabathula D, Zhang Q, Bartels D (2018) Protection of photosynthesis in desiccation-tolerant resurrection plants. J Plant Physiol 227:84–92. https://doi.org/10.1016/j.jplph.2018.05.002
Cove D (2005) The Moss Physcomitrella patens. Annu Rev Genet 39:39–358. https://doi.org/10.1146/annurev.genet.39.073003.110214
Cove D, Bezanilla M, Harries P, Quatrano R (2006) Mosses as model systems for the study of metabolism and development. Annu Rev Plant Biol 57:497–520. https://doi.org/10.1146/annurev.arplant.57.032905.105338
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163. https://doi.org/10.1186/1471-2229-11-163
Cruz de Carvalho R, Bernardes da Silva A, Branquinho C, Marques da Silva J (2015) Influence of dehydration rate on cell sucrose and water relations parameters in an inducible desiccation tolerant aquatic bryophyte. Environ Exp Bot 120:18–22. https://doi.org/10.1016/j.envexpbot.2015.07.002
Decker EL, Frank W, Sarnighausen E, Reski R (2006) Moss systems biology en route: phytohormones in Physcomitrella development. Plant Biol 8:397–406. https://doi.org/10.1055/s-2006-923952
Dinakar C, Bartels D (2013) Desiccation tolerance in resurrection plants: new insights from transcriptome, proteome, and metabolome analysis. Front Plant Sci 4:1–14. https://doi.org/10.3389/fpls.2013.00482
Dong S, Beckles DM (2019) Dynamic changes in the starch-sugar interconversion within plant source and sink tissues promote a better abiotic stress response. J of Plant Physiol 234–235:80–93. https://doi.org/10.1016/j.jplph.2019.01.007
Farrant JM, Cooper K, Hilgart A, Abdalla KO, Bentley J, Thomson JA, Dace HJW, Peton N, Mundree SG, Rafudeen MS (2015) A molecular physiological review of vegetative desiccation tolerance in the resurrection plant Xerophyta viscosa (Baker). Planta 242:407–426. https://doi.org/10.1007/s00425-015-2320-6
Fedoroff NV, Battisti DS, Beachy RN, Cooper PJM, Fischhoff DA, Hodges CN, Knauf VC et al (2010) Radically rethinking agriculture for the 21st century. Sci 327:833–834. https://doi.org/10.1126/science.1186834
Finkelstein RR (1994) Maternal effects govern variable dominance of two abscisic acid response mutations in Arabidopsis thaliana. Plant Physiol 105:1203–1208. https://doi.org/10.1104/pp.105.4.1203
Finkelstein R (2013) Abscisic acid synthesis and response. Arabidopsis Book 11:e0166.
Frank W, Ratnadewi D, Reski R (2005) Physcomitrella patens is highly tolerant against drought, salt and osmotic stress. Planta 220:384–394. https://doi.org/10.1007/s00425-004-1351-1
Gao B, Zhang D, Li X, Yang H, Wood A (2014) De novo assembly and characterization of the transcriptome in the desiccation-tolerant moss Syntrichia caninervis. BMC Res Notes 7:490. https://doi.org/10.1186/1756-0500-7-490
Gao B, Zhang D, Li X, Yang H, Zhang Y, Wood A (2015) De novo transcriptome characterization and gene expression profiling of the desiccation tolerant moss Bryum argenteum following rehydration. BMC Genom 16:416. https://doi.org/10.1186/s12864-015-1633-y
Gao B, Li X, Zhang D, Liang Y, Yang H, Chen M, Zhang Y et al (2017) Desiccation tolerance in bryophytes: the dehydration and rehydration transcriptomes in the desiccation-tolerant bryophyte Bryum argenteum. Sci Rep 7:7572. https://doi.org/10.1038/s41598-017-07297-3
Garwe D, Thomson JA, Mundree SG (2006) XVSAP1 from Xerophyta viscosa improves osmotic-, salinity-and high-temperature-stress tolerance in Arabidopsis. Biotechnol J 1:1137–1146. https://doi.org/10.1002/biot.200600136
Giarola V, Bartels D (2015) What can we learn from the transcriptome of the resurrection plant Craterostigma plantagineum? Planta 242:427–434. https://doi.org/10.1007/s00425-015-2327-z
Giarola V, Hou Q, Bartels D (2017) Angiosperm plant desiccation tolerance: hints from transcriptomics and genome sequencing. Trends Plant Sci 22:705–717. https://doi.org/10.1016/j.tplants.2017.05.007
Godfray HC, Beddington JR, Crute IR, Haddad L, Lawrence D, Muir JF, Pretty J, Robinson S, Thomas SM, Toulmin C (2010) Food security: the challenge of feeding 9 billion people. Science 327:812–818. https://doi.org/10.1126/science.1185383
Guiltinan MJ, Marcotte WR, Quatrano RS (1990) A plant leucine zipper protein that recognizes an abscisic acid response element. Science 250:267–271. https://doi.org/10.1126/science.2145628
Gupta A, Rico-Medina A, Caño-Delgado AI (2020) The physiology of plant responses to drought. Science 368:266–269. https://doi.org/10.1126/science.aaz7614
Gururani MA, Venkatesh J, Tran LSP (2015) Regulation of photosynthesis during abiotic stress-induced photoinhibition. Mol Plant 8:1304–1320. https://doi.org/10.1016/j.molp.2015.05.005
Haldrup A, Simpson DJ, Scheller HV (2000) Down-regulation of the psi-f subunit of photosystem i (psi) in arabidopsis thaliana: the psi-f subunit is essential for photoautotrophic growth and contributes to antenna function. J Biol Chem 275:31211–31218. https://doi.org/10.1074/jbc.m002933200
Hasanuzzaman M, Bhuyan MHMB, Parvin K, Bhuiyan TF, Anee TI, Nahar K, Hossen MS, Zulfiqar F, Alam MM, Fujita M (2020) Regulation of ROS metabolism in plants under environmental stress: a review of recent experimental evidence. Int J Mol Sci 21:8695. https://doi.org/10.3390/ijms21228695
Hertz MI, Landry DM, Willis AE, Luo G, Thompson SR (2013) Ribosomal protein S25 dependency reveals a common mechanism for diverse internal ribosome entry sites and ribosome shunting. Mol Cell Biol 33:1016–1026. https://doi.org/10.1128/MCB.00879-12
Hiss M, Laule O, Meskauskiene RM, Arif MA, Decker EL, Erxleben A, Frank W, Hanke ST, Lang D, Martin A, Neu C, Reski R, Richardt S, Schallenberg-Rüdinger M, Szövényi P, Tiko T, Wiedemann G, Wolf L, Zimmermann P, Rensing SA (2014) Large-scale gene expression profiling data for the model moss physcomitrella patens aid understanding of developmental progression, culture and stress conditions. Plant J 79:530–539. https://doi.org/10.1111/tpj.12572
Huerta-Cepas J, Szklarczyk D, Heller D, Hernández-Plaza A, Forslund SK, Cook H, Mende DR, Letunic I, Rattei T, Jensen LJ, von Mering C, Bork P (2019) eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47:D309–D314. https://doi.org/10.1093/nar/gky1085
Humplík JF, Bergougnoux V, Volkenburgh EV (2017) To stimulate or inhibit? that is the question for the function of abscisic acid. Trends Plant Sci 22:830–841. https://doi.org/10.1016/j.tplants.2017.07.009
Joshi R, Karan R, Singla-Pareek SL, Pareek A (2016) Ectopic expression of Pokkali phosphoglycerate kinase-2 (OsPGK2-P) improves yield in tobacco plants under salinity stress. Plant Cell Rep 35:27–41. https://doi.org/10.1007/s00299-015-1864-z
Khandelwal A, Cho SH, Marella H, Sakata Y, Perroud PF, Pan A, Quatrano RS (2010) Role of ABA and ABI3 in desiccation tolerance. Sci 327:546. https://doi.org/10.1126/science.1183672
Khraiwesh B, Qudeimat E, Thimma M, Chaiboonchoe A, Jijakli K, Alzahmi A, Arnoux M, Salehi-Ashtiani K M (2015) Genome-wide expression analysis offers new insights into the origin and evolution of Physcomitrella patens stress response. Sci Rep 5:17434. https://doi.org/10.1038/srep17434
Komatsu K, Nishikawa Y, Ohtsuka T et al (2009) Functional analyses of the ABI1-related protein phosphatase type 2 C reveal evolutionarily conserved regulation of abscisic acid signaling between Arabidopsis and the moss Physcomitrella patens.. Plant Mol Biol 70:327–340. https://doi.org/10.1007/s11103-009-9476-z
Komatsu K, Suzuki N, Kuwamura M, Nishikawa Y, Nakatani M, Ohtawa H, Takezawa M D (2013) Group A PP2Cs evolved in land plants as key regulators of intrinsic desiccation tolerance. Nat Commun 4:1–9. https://doi.org/10.1038/ncomms3219
Komatsu K, Takezawa D, Sakata Y (2020) Decoding ABA and osmostress signalling in plants from an evolutionary point of view. Plant Cell Environ 43:2894–2911. https://doi.org/10.1111/pce.13869
Koornneef M, Reuling G, Karssen CM (1984) The isolation and characterization of abscisic acid-insensitive mutants of Arabidopsis thaliana. Physiol Plant 61:377–383. https://doi.org/10.1111/j.1399-3054.1984.tb06343.x
Koster KL, Balsamo RA, Espinoza C, Oliver MJ (2010) Desiccation sensitivity and tolerance in the moss Physcomitrella patens: Assessing limits and damage. Plant Growth Regul 62:293–302. https://doi.org/10.1007/s10725-010-9490-9
Lang D, Ullrich KK, Murat F, Fuchs J, Jenkins J, Haas FB, Piednoel M, Gundlach H, Van Bel M, Meyberg R, Vives C, Morata J, Symeonidi A, Hiss M, Muchero W, Kamisugi Y, Saleh O, Blanc G, Decker EL, van Gessel N, Grimwood J, Hayes RD, Graham SW, Gunter LE, McDaniel SF, Hoernstein SNW, Larsson A, Li FW, Perroud PF, Phillips J, Ranjan P, Rokshar DS, Rothfels CJ, Schneider L, Shu S, Stevenson DW, Thümmler F, Tillich M, Villarreal Aguilar JC, Widiez T, Wong GK, Wymore A, Zhang Y, Zimmer AD, Quatrano RS, Mayer KFX, Goodstein D, Casacuberta JM, Vandepoele K, Reski R, Cuming AC, Tuskan GA, Maumus F, Salse J, Schmutz J, Rensing SA (2018) The Physcomitrella patens chromosome-scale assembly reveals moss genome structure and evolution. Plant J. 93(3):515–533. https://doi.org/10.1111/tpj.13801
Laxalt AM, Cassia RO, Sanllorenti PM, Madrid EA, Andreu AB, Daleo GR, Conde RD, Lamattina L (1996) Accumulation of cytosolic glyceraldehyde-3-phosphate dehydrogenase RNA under biological stress conditions and elicitor treatments in potato. Plant Mol Biol 30:961–972. https://doi.org/10.1007/BF00020807
Leprince O, Buitink J (2015) Introduction to desiccation biology: from old borders to new frontiers. Planta 242:369–78
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acid Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Li C, Wang M, Wu XM, Chen DH, Lv HJ, Shen JL, Qiao Z, Zhang W (2016) THI1, a thiamine thiazole synthase, interacts with Ca2+-dependent protein kinase CPK3 and modulates the S-type anion channels and stomatal closure in arabidopsis. Plant Physiol 170:1090–1104. https://doi.org/10.1104/pp.15.01649
Li X, Wei W, Li F, Zhang L, Deng X, Liu Y, Yang S (2019) The plastidial glyceraldehyde-3-phosphate dehydrogenase is critical for abiotic stress response in wheat. Int J Mol Sci 20:1104. https://doi.org/10.3390/ijms20051104
Liu D, Li W, Cheng J, Hou L (2015) AtPGK2, a member of PGKs gene family in Arabidopsis, has a positive role in salt stress tolerance. Plant Cell Tissue Organ Cult 120:251–262. https://doi.org/10.1007/s11240-014-0601-6
Liu L, Last RL (2015) MPH1 is a thylakoid membrane protein involved in protecting photosystem II from photodamage in land plants. Plant Signaling Behavior 10:10. https://doi.org/10.1080/15592324.2015.1076602
Lu W, Tang X, Huo Y, Xu R, Qi S, Huang J, Zheng C, Wu C (2012) Identification and characterization of fructose 1,6-bisphosphate aldolase genes in Arabidopsis reveal a gene family with diverse responses to abiotic stresses. Gene 503:65–74. https://doi.org/10.1016/j.gene.2012.04.042
Ma Y, Cao J, He J, Chen Q, Li X, Yang Y (2018) Molecular mechanism for the regulation of ABA homeostasis during plant development and stress responses. Int J Mol Sci 19:3643. https://doi.org/10.3390/ijms19113643
Mayaba N, Beckett RP, Csintalan Z, Tuba Z (2001) ABA increases the desiccation tolerance of photosynthesis in the afromontane understorey moss Atrichum androgynum. Ann Bot 88:1093–1100. https://doi.org/10.1006/anbo.2001.1552
Minami A, Nagao M, Ikegami K, Koshiba T, Arakawa K, Fujikawa S, Takezawa D (2005) Cold acclimation in bryophytes: low-temperature-induced freezing tolerance in Physcomitrella patens is associated with increases in expression levels of stress-related genes but not with increase in level of endogenous abscisic acid. Planta 220:414–423. https://doi.org/10.1007/s00425-004-1361-z
Morita-Yamamuro C, Tsutsui T, Tanaka A, Yamaguchi J (2004) Knock-out of the plastid ribosomal protein S21 causes impaired photosynthesis and sugar-response during germination and seedling development in Arabidopsis thaliana. Plant Cell Physiol 45:781–788. https://doi.org/10.1093/pcp/pch093
Nagao M, Minami A, Arakawa K, Fujikawa S, Takezawa D (2005) Rapid degradation of starch in chloroplasts and concomitant accumulation of soluble sugars associated with ABA-induced freezing tolerance in the moss Physcomitrella patens. J Plant Physiol 162:169–180. https://doi.org/10.1016/j.jplph.2004.06.012
Nagao M, Oku K, Minami A, Mizuno K, Sakurai M, Arakawa K, Fujikawa S, Takezawa D (2006) Accumulation of theanderose in association with development of freezing tolerance in the moss Physcomitrella patens. Phytochemistry 67:702–709. https://doi.org/10.1016/j.phytochem.2006.01.031
Oliver MJ, Bewley JD (1997) Desiccation-tolerance of plant tissues: a mechanistic overview. In: Janick J J (ed) Horticultural Reviews. John Wiley & Sons, New Jersey, pp 295–316
Oliver MJ, AJ Wood (1997) Desiccation tolerance in mosses. In: Koval TM (ed) Stress-inducible process in high eukaryotic cells. Springer, Boston, pp 1–26
Oliver MJ, Wood AJ, O’Mahony P (1997) How some plants recover from vegetative desiccation: a repair based strategy. Acta Physiol Plant 19:419–425. https://doi.org/10.1007/s11738-997-0038-1
Oliver MJ, Tuba Z, Mishler BD (2000) The evolution of vegetative desiccation tolerance in land plants. Plant Ecol 151:85–100. https://doi.org/10.1023/A:1026550808557
Oliver MJ, Velten J, Mishler BD (2005) Desiccation tolerance in bryophytes: a reflection of the primitive strategy for plant survival in dehydrating habitats? Integr Comp Biol 45:788–799. https://doi.org/10.1093/icb/45.5.788
Omidbakhshfard MA, Omranian N, Ahmadi FS, Nikoloski Z, Mueller-Roeber B (2012) Effect of salt stress on genes encoding translation-associated proteins in Arabidopsis thaliana. Plant Signaling Behavior 7:095–1102. https://doi.org/10.4161/psb.21218
Orr W, Keller WA, Singh J (1986) Induction of freezing tolerance in an embryogenic cell suspension culture of Brassica Napus by abscisic acid at room temperature. J Plant Physiol 126:26–32. https://doi.org/10.1016/S0176-1617(86)80212-7
Ortiz-Ramírez C, Hernandez-Coronado M, Thamm A, Catarino B, Wang M, Dolan L, Feijó JA, Becker JD (2016) A transcriptome atlas of Physcomitrella patens provides insights into the evolution and development of land plants. Mol Plant 9:205–2020. https://doi.org/10.1016/j.molp.2015.12.002
Pence VC, Dunford SS, Redella S (2005) Differential effects of abscisic acid on desiccation tolerance and carbohydrates in three species of liverworts. J Plant Physiol 162:1331–1337. https://doi.org/10.1016/j.jplph.2005.02.002
Perroud PF, Haas FB, Hiss M, Ullrich KK, Arbolesi A, Amirebrahimi M, Barry K et al (2018) The Physcomitrella patens gene atlas project: large-scale RNA-Seq based expression data. Plant J 95:168–182. https://doi.org/10.1111/tpj.13940
Piro A, Marín-Guirao L, Serra IA, Spadafora A, Sandoval-Gil JM, Bernardeau-Esteller J, Fernandez JMR, Mazzuca S (2015) The modulation of leaf metabolism plays a role in salt tolerance of Cymodocea nodosa exposed to hypersaline stress in mesocosms. Front Plant Sci 6:464. https://doi.org/10.3389/fpls.2015.00464
Possart A, Hiltbrunner A (2013) An evolutionarily conserved signaling mechanism mediates far-red light responses in land plants. Plant Cell 25:102–114. https://doi.org/10.1105/tpc.112.104331
Pressel S, Duckett JG (2010) Cytological insights into the desiccation biology of a model system: moss protonemata. New Phytol 185:944–963. https://doi.org/10.1111/j.1469-8137.2009.03148.x
Proctor MCF, Smirnoff N (2000) Rapid recovery of photosystems on rewetting desiccation-tolerant mosses: chlorophyll fluorescence and inhibitor experiments. J Exp Bot 51:1695–1704. https://doi.org/10.1093/jexbot/51.351.1695
Proctor MCF, Ligrone R, Duckett JG (2007) Desiccation tolerance in the moss Polytrichum formosum: physiological and fine-structural changes during desiccation and recovery. Ann Bot 99:75–93. https://doi.org/10.1093/aob/mcl246
Rajjou L, Gallardo K, Debeaujon I, Vandekerchhove J, Job C, Job D (2004) The effect of alpha-amanitin on the Arabidopsis seed proteome highlights the distinct roles of stored and neosynthesized mRNAs during germination. Plant Physiol 134:1598–1613. https://doi.org/10.1104/pp.103.036293
Ramanjulu S, Bartels D (2002) Drought- and desiccation-induced modulation of gene expression in plants. Plant Cell Environ 25:141–151. https://doi.org/10.1046/j.0016-8025.2001.00764.x
Rathnayake KN, Nelson S, Seeve C, Oliver MJ, Koster KL (2019) Acclimation and endogenous abscisic acid in the moss Physcomitrella patens during acquisition of desiccation tolerance. Physiol Plant 167:317–329. https://doi.org/10.1111/ppl.12892
Rensing SA, Goffinet B, Meyberg R, Wu SZ, Bezanilla M (2020) The moss Physcomitrium (Physcomitrella) patens: a model organism for non-seed plants. Plant Cell 32:1361–1376. https://doi.org/10.1105/tpc.19.00828
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Sah SK, Reddy KR, Li J (2016) Abscisic acid and abiotic stress tolerance in crop plants. Front Plant Sci 7:571. https://doi.org/10.3389/fpls.2016.00571
Sakata Y, Komatsu K, Taji T, Tanaka S (2009) Role of PP2C-mediated ABA signalling in the moss Physcomitrella patens. Plant Signal Behav 4:887–889. https://doi.org/10.4161/psb.4.9.9464
Saruhashi M, Ghosh TK, Arai K, Ishizaki Y, Hagiwara K, Komatsu K, Shiwa Y, Izumikawa K, Yoshikawa H, Umezawa T, Sakata Y, Takezawa D (2015) Plant Raf-like kinase for ABA and stress signaling. PNAS. https://doi.org/10.1073/pnas.1511238112
Stress-induced changes in Peptidyl-Prolyl cis-trans isomerase activity of Sorghum bicolor seedlings. Biol Plant 47:367–371.
Shinozawa A, Otake R, Takezawa D, Umezawa T, Komatsu K, Tanaka K, Amagai A, Ishikawa S, Hara Y, Kamisugi Y, Cuming AC, Hori K, Ohta H, Takahashi F, Shinozaki K, Hayashi T, Taji T, Sakata Y (2019) SnRK2 protein kinases represent an ancient system in plants for adaptation to a terrestrial environment. Commun Biol 2:30. https://doi.org/10.1038/s42003-019-0281-1
Stark LR, Greenwood JL, Brinda JC, Oliver MJ (2013) The desert moss Pterygoneurum Lamellatum (Pottiaceae) exhibits an inducible ecological strategy of desiccation tolerance: effects of rate of drying on shoot damage and regeneration. Am J Bot 100:1522–1531. https://doi.org/10.3732/ajb.1200648
Stevenson SR, Kamisugi Y, Trinh CH, Schmutz J, Jenkins JW, Grimwood J, Muchero W (2016) Genetic analysis of Physcomitrella patens identifies ABSCISIC ACID NON-RESPONSIVE, a regulator of ABA responses unique to basal land plants and required for desiccation tolerance. Plant Cell 28:1310–1327. https://doi.org/10.1105/tpc.16.00091
Takezawa D, Komatsu K, Sakata Y (2011) ABA in bryophytes: how a universal growth regulator in life became a plant hormone? J Plant Res 124:437–453. https://doi.org/10.1007/s10265-011-0410-5
Takezawa D, Watanabe N, Ghosh TK, Saruhashi M, Suzuki A, Ishiyama K, Somemiya S (2015) Epoxycarotenoid-mediated synthesis of abscisic acid in Physcomitrella patens implicating conserved mechanisms for acclimation to hyperosmosis in embryophytes. New Phytol 206:209–219. https://doi.org/10.1111/nph.13231
Tan T, Sun Y, Peng X, Wu G, Bao F, He Y, Zhou H, Lin H (2017) ABSCISIC ACID INSENSITIVE3 is involved in cold response and freezing tolerance regulation in Physcomitrella patens. Front Plant Sci 8:1599. https://doi.org/10.3389/fpls.2017.01599
Torabi S, Umate P, Manavski N, Plöchinger M, Kleinknecht L, Bogireddi H, Herrmann RG, Wanner G, Schröder WP, Meurer J (2014) PsbN is required for assembly of the photosystem II reaction center in Nicotiana tabacum. Plant Cell 26:1183–1199. https://doi.org/10.1105/tpc.113.120444
Vesty EF, Saidi Y, Moody LA, Holloway D, Whitbread A, Needs S, Choudhary A (2016) The decision to germinate is regulated by divergent molecular networks in spores and seeds. New Phytol 211:952–966. https://doi.org/10.1111/nph.14018
Villalobos MA, Bartels D, Iturriaga G (2004) Stress tolerance and glucose insensitive phenotypes in arabidopsis overexpressing the CpMYB10 transcription factor gene. Plant Phys 135:309–324. https://doi.org/10.1104/pp.103.034199
Vishwakarma K, Upadhyay N, Kumar N, Gaurav Y, Singh J, Mishra RK, Kumar V (2017) Abscisic acid signaling and abiotic stress tolerance in plants: a review on current knowledge and future prospects. Front Plant Sci 8:161. https://doi.org/10.3389/fpls.2017.00161
Wang J, Lan P, Gao H, Zheng L, Li W, Schmidt W (2013) Expression changes of ribosomal proteins in phosphate- and iron-deficient Arabidopsis roots predict stress-specific alterations in ribosome composition. BMC Genom 14:783. https://doi.org/10.1186/1471-2164-14-783
Wang QJ, Sun H, Dong QL, Sun TY, Jin ZX, Hao YJ, Yao YX (2016) The enhancement of tolerance to salt and cold stresses by modifying the redox state and salicylic acid content via the cytosolic malate dehydrogenase gene in transgenic apple plants. Plant Biotechnol J 14:1986–1997. https://doi.org/10.1111/pbi.12556
Werner O, Espín RM, Bopp M, Atzorn R (1991) Abscisic-acid-induced drought tolerance in Funaria hygrometrica Hedw. Planta 186:99–103. https://doi.org/10.1007/BF00201503
Wood AJ, Oliver MJ, Cove DJ (2000) Bryophytes as model system. Bryologist 103:128–133. https://doi.org/10.1639/0007-2745
Wood AJ (2007) The nature and distribution of vegetative desiccation-tolerance in hornworts, liverworts and mosses. Bryologist 110:163–177. https://doi.org/10.1639/0007-2745
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264. https://doi.org/10.1105/tpc.6.2.251
Yemm EW, Willis AJ (1954) The estimation of carbohydrates in plant extracts by anthrone. Biochem J 57:508–514. https://doi.org/10.1042/bj0570508
Yu CS, Cheng CW, Su WC, Chang KC, Huang S-W, Hwang JK, Lu CH (2014) CELLO2GO: a web server for protein subcellular localization prediction with functional gene ontology annotation. PLoS ONE 9(6):e99368. https://doi.org/10.1371/journal.pone.0099368
Zhang Q, Bartels D (2018) Molecular responses to dehydration and desiccation in desiccation-tolerant angiosperm plants. J Exp Bot 69:3211–3222. https://doi.org/10.1093/jxb/erx489
Zhao Y, Kong H, Guo Y, Zou Z (2020) Light-harvesting chlorophyll a/b-binding protein-coding genes in jatropha and the comparison with castor, cassava and arabidopsis. PeerJ 8:e8465. https://doi.org/10.7717/peerj.8465
Zhou XT, Wang F, Ma YP, Jia LJ, Liu N, Wang HY, Zhao P, Xia GX, Zhong NQ (2018) Ectopic expression of SsPETE2, a plastocyanin from Suaeda salsa, improves plant tolerance to oxidative stress. Plant Sci 268:1–10. https://doi.org/10.1016/j.plantsci.2017.12.006
Acknowledgements
We thank Biól. Enrique Ortiz Bermúdez (IB-UNAM) and Biól. Ana Susana Estrada Márquez (IB-UNAM) for their kind help in constructing the distribution map of P. replicatum. We thank Dr. Ricardo Grande of the Unidad Universitaria de Secuenciación Masiva y Bioinformática (IBT-UNAM) for his technical support with RNA-Seq, and we thank Dr. Fernanda Iruegas Bocardo for technical assistance. We thank Ing. Gabriela Guerrero and Dr. Luis Lozano from Unidad de Análisis Bioinformáticos (CCG-UNAM) for technical advice. This work was supported by the Instituto Politécnico Nacional SIP-20201783, SIP-20201977, CONACYT Ciencia Básica 62174 and CONACYT Ciencia Básica A1-S-35357 Grants. AAB and MAVL are grateful for financial support from CONACYT to obtain the participation of MAMN throughout the project Cátedras CONACYT 1452. SRM is grateful for the CONACYT PhD fellowship.
Author information
Authors and Affiliations
Contributions
MAVL and AAB conceived and designed the research. SRM conducted the experiments. CD and JLM contributed to sample collection, taxonomic identification of P. replicatum and ecological data. MB, LC, and LL contributed to soluble sugar quantification and provided microscopic and analytical tools. MAMN, MSP, and EV performed experiments for RNA-Seq and the bioinformatics analysis of the transcriptome data. IEMM enriched the discussion of the results. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article belongs to the Topical Collection: Molecular Biology of Mosses.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ríos-Meléndez, S., Valadez-Hernández, E., Delgadillo, C. et al. Pseudocrossidium replicatum (Taylor) R.H. Zander is a fully desiccation-tolerant moss that expresses an inducible molecular mechanism in response to severe abiotic stress. Plant Mol Biol 107, 387–404 (2021). https://doi.org/10.1007/s11103-021-01167-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-021-01167-3