Abstract
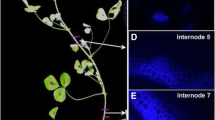
Sugarcane (Saccharum spp. hybrids) accumulates high concentrations of sucrose in its mature stalk and a considerable portion of carbohydrate metabolism is also devoted to cell wall synthesis and fibre production. We examined tissue-specific expression patterns to explore the spatial deployment of pathways responsible for sucrose accumulation and fibre synthesis within the stalk. We performed expression profiling of storage parenchyma, vascular bundles and rind dissected from a maturing stalk internode of sugarcane, identifying ten cellulose synthase subunit genes and examining significant differences in the expression of their corresponding transcripts and those of several sugar transporters. These were correlated with differential expression patterns for transcripts of genes encoding COBRA-like proteins and other cell wall metabolism-related proteins. The sugar transporters genes ShPST2a, ShPST2b and ShSUT4 were significantly up-regulated in storage parenchyma while ShSUT1 was up-regulated in vascular bundles. Two co-ordinately expressed groups of cell wall related transcripts were also identified. One group, associated with primary cell wall synthesis (ShCesA1, ShCesA7, ShCesA9 and Shbk2l3), was up-regulated in parenchyma. The other group, associated with secondary cell wall synthesis (ShCesA10, ShCesA11, ShCesA12 and Shbk-2), was up-regulated in rind. In transformed sugarcane plants, the ShCesA7 promoter conferred stable expression of green fluorescent protein preferentially in the storage parenchyma of the maturing stalk internode. Our results indicate that there is spatial separation for elevated expression of these important targets in both sucrose accumulation and cell wall synthesis, allowing for increased clarity in our understanding of sucrose transport and fibre synthesis in sugarcane.







Similar content being viewed by others
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Anisimova M, Gascuel O (2006) Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol 55:539–552. doi:10.1080/10635150600755453
Appenzeller L, Doblin M, Barreiro R et al (2004) Cellulose synthesis in maize: isolation and expression analysis of the cellulose synthase (CesA) gene family. Chem Mater Sci 11:287–299. doi:10.1023/b:cell.0000046417.84715.27
Baker SS, Wilhelm KS, Thomashow MF (1994) The 5′-region of Arabidopsis thaliana cor15a has cis-acting elements that confer cold-, drought- and ABA-regulated gene expression. Plant Mol Biol 24:701–713. doi:10.1007/BF00029852
Basnayake SWV, Morgan TC, Wu L, Birch RG (2012) Field performance of transgenic sugarcane expressing isomaltulose synthase. Plant Biotechnol J 10:217–225. doi:10.1111/j.1467-7652.2011.00655.x
Bernard V, Brunaud V, Lecharny A (2010) TC-motifs at the TATA-box expected position in plant genes: a novel class of motifs involved in the transcription regulation. BMC Genomics 11:166
Bonnett GD, Hewitt ML, Glassop D (2006) Effects of high temperature on the growth and composition of sugarcane internodes. Aust J Agric Res 57:1087–1095. doi:10.1071/AR06042
Botha FC (2009) Energy yield and cost in a sugarcane biomass system. Proc Aust Soc Sugarcane Technol 31:1–10
Bower R, Elliott AR, Potier BAM, Birch RG (1996) High-efficiency, microprojectile-mediated cotransformation of sugarcane, using visible or selectable markers. Mol Breed 2:239–249
Brady SM, Song S, Dhugga KS et al (2007) Combining expression and comparative evolutionary analysis. The COBRA gene family. Plant Physiol 143:172–187. doi:10.1104/pp.106.087262
Bull TA, Glasziou KT (1963) The evolutionary significance of sugar accumulation in Saccharum. Aust J Biol Sci 16:737–742
Burton RA, Shirley NJ, King BJ et al (2004) The CesA gene family of barley. Quantitative analysis of transcripts reveals two groups of co-expressed genes. Plant Physiol 134:224–236
Burton RA, Ma G, Baumann U et al (2010) A customized gene expression microarray reveals that the brittle stem phenotype fs2 of barley is attributable to a retroelement in the HvCesA4 cellulose synthase gene. Plant Physiol 153:1716–1728. doi:10.1104/pp.110.158329
Cardoso-Silva CB, Costa EA, Mancini MC et al (2014) De novo assembly and transcriptome analysis of contrasting sugarcane varieties. PLoS One. doi:10.1371/journal.pone.0088462
Carson DL, Huckett BI, Botha FC (2002a) Sugarcane ESTs differentially expressed in immature and maturing internodal tissue. Plant Sci 162:289–300
Carson DL, Huckett BI, Botha FC (2002b) Differential gene expression in sugarcane leaf and internodal tissues of varying maturity. S Afr J Bot 68:434–442
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552. doi:10.1093/oxfordjournals.molbev.a026334
Casu RE, Grof CPL, Rae AL et al (2003) Identification of a novel sugar transporter homologue strongly expressed in maturing stem vascular tissues of sugarcane by expressed sequence tag and microarray analysis. Plant Mol Biol 52:371–386
Casu RE, Dimmock CM, Chapman SC et al (2004) Identification of differentially expressed transcripts from maturing stem of sugarcane by in silico analysis of stem expressed sequence tags and gene expression profiling. Plant Mol Biol 54:503–517
Casu RE, Jarmey JM, Bonnett GD, Manners JM (2007) Identification of transcripts associated with cell wall metabolism and development in the stem of sugarcane by Affymetrix Genechip Sugarcane Genome Array expression profiling. Funct Integr Genomics 7:153–167
Chevenet F, Brun C, Bañuls A-L et al (2006) TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinform 7:439. doi:10.1186/1471-2105-7-439
Ching A, Dhugga KS, Appenzeller L et al (2006) Brittle stalk 2 encodes a putative glycosylphosphatidylinositol-anchored protein that affects mechanical strength of maize tissues by altering the composition and structure of secondary cell walls. Planta 224:1174–1184
Cho J-I, Burla B, Lee D-W et al (2010) Expression analysis and functional characterization of the monosaccharide transporters, OsTMTs, involving vacuolar sugar transport in rice (Oryza sativa). New Phytol 186:657–668. doi:10.1111/j.1469-8137.2010.03194.x
Conesa A, Götz S, García-Gómez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. doi:10.1093/bioinformatics/bti610
Damaj M, Kumpatla S, Emani C et al (2010) Sugarcane DIRIGENT and O-METHYLTRANSFERASE promoters confer stem-regulated gene expression in diverse monocots. Planta 231:1439–1458
de Maria Felix J, Papini-Terzi F, Rocha F et al (2009) Expression profile of signal transduction components in a sugarcane population segregating for sugar content. Trop Plant Biol 2:98–109
Dereeper A, Guignon V, Blanc G et al (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469. doi:10.1093/nar/gkn180
Dubouzet JG, Sakuma Y, Ito Y et al (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763. doi:10.1046/j.1365-313X.2003.01661.x
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Eom J-S, Cho J-I, Reinders A et al (2011) Impaired function of the tonoplast-localized sucrose transporter in rice, OsSUT2, limits the transport of vacuolar reserve sucrose and affects plant growth. Plant Physiol 157:109–119. doi:10.1104/pp.111.176982
Fong Chong B, Abeydeera WPP, Glassop D et al (2010) Co-ordinated synthesis of gentiobiitol and sorbitol, evidence of sorbitol glycosylation in transgenic sugarcane. Phytochemistry 71:736–741
Franks T, Birch RG (1991) Gene transfer into intact sugarcane cells using microprojectile bombardment. Funct Plant Biol 18:471–480. doi:10.1071/PP9910471
Goldemberg J (2008) The Brazilian biofuels industry. Biotechnol Biofuels 1:6
Grivet L, Arruda P (2001) Sugarcane genomics: depicting the complex genome of an important tropical crop. Curr Opin Plant Biol 5:122–127
Grof C, Byrt C, Patrick J (2014) Phloem transport of resources. In: Moore PH, Botha FC (eds) Sugarcane: physiology, biochemistry, and functional biology. Wiley, Chichester, pp 267–305
Guindon S, Dufayard JF, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. doi:10.1093/sysbio/syq010
Hamerli D, Birch RG (2011) Transgenic expression of trehalulose synthase results in high concentrations of the sucrose isomer trehalulose in mature stems of field-grown sugarcane. Plant Biotechnol J 9:32–37. doi:10.1111/j.1467-7652.2010.00528.x
Hawker JS (1985) Sucrose. In: Dey PM, Dixon RA (eds) Biochemistry of storage carbohydrates in green plants. Academic Press, London, pp 1–52
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Hwang Y, Karrer E, Thomas B et al (1998) Three cis-elements required for rice α-amylase Amy3D expression during sugar starvation. Plant Mol Biol 36:331–341. doi:10.1023/A:1005956104636
Iskandar HM, Casu RE, Fletcher AT et al (2011) Identification of drought-response genes and a study of their expression during sucrose accumulation and water deficit in sugarcane culms. BMC Plant Biol 11:12
Jackson P (2013) Energy cane. In: Saha MC, Bhandari HS, Bouton JH (eds) Bioenergy feedstocks, 1st edn. Wiley, Hoboken, pp 117–149
Jacobsen KR, Fisher DG, Moore PH (1992) Developmental changes in the anatomy of the sugarcane stem in relation to phloem unloading and sucrose storage. Bot Acta 105:p70–p80
Joyce P, Kuwahata M, Turner N, Lakshmanan P (2010) Selection system and co-cultivation medium are important determinants of Agrobacterium-mediated transformation of sugarcane. Plant Cell Rep 29:173–183
Jung B, Ludewig F, Schulz A et al (2015) Sucrose accumulation in sugar beet taproots. Nat Plants 1:1–6. doi:10.1038/nplants.2014.1
Kizis D, Pagès M (2002) Maize DRE-binding proteins DBF1 and DBF2 are involved in rab17 regulation through the drought-responsive element in an ABA-dependent pathway. Plant J 30:679–689. doi:10.1046/j.1365-313X.2002.01325.x
Kotake T, Aohara T, Hirano K et al (2011) Rice Brittle culm 6 encodes a dominant-negative form of CesA protein that perturbs cellulose synthesis in secondary cell walls. J Exp Bot 62:2053–2062. doi:10.1093/jxb/erq395
Li Y, Qian Q, Zhou Y et al (2003) BRITTLE CULM1, which encodes a COBRA-like protein, affects the mechanical properties of rice plants. Plant Cell 15:2020–2031. doi:10.1105/tpc.011775.chanical
Liu L, Shang-Guan K, Zhang B et al (2013) Brittle culm1, a COBRA-like protein, functions in cellulose assembly through binding cellulose microfibrils. PLoS Genet 9:e1003704. doi:10.1371/journal.pgen.1003704
McCormick AJ, Cramer MD, Watt DA (2008) Differential expression of genes in the leaves of sugarcane in response to sugar accumulation. Trop Plant Biol 1:142–158
McQualter RB, Fong Chong B, Meyer K et al (2005) Initial evaluation of sugarcane as a production platform for p-hydroxybenzoic acid. Plant Biotechnol J 3:29–41
Milne RJ, Byrt CS, Patrick JW, Grof CP (2013) Are sucrose transporter expression profiles linked with patterns of biomass partitioning in Sorghum phenotypes? Front Plant Sci. doi:10.3389/fpls.2013.00223
Moyle RL, Birch R (2013a) Diversity of sequences and expression patterns among alleles of a sugarcane loading stem gene. Theor Appl Genet 126:1775–1782. doi:10.1007/s00122-013-2091-z
Moyle RL, Birch RG (2013b) Sugarcane Loading Stem Gene promoters drive transgene expression preferentially in the stem. Plant Mol Biol 82:51–58. doi:10.1007/s11103-013-0034-3
Mudge S, Osabe K, Casu R et al (2009) Efficient silencing of reporter transgenes coupled to known functional promoters in sugarcane, a highly polyploid crop species. Planta 229:549–558
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nass LL, Pereira PAA, Ellis D (2007) Biofuels in Brazil: an overview. Crop Sci 47:2228–2237. doi:10.2135/cropsci2007.03.0166
Nordin K, Vahala T, Palva ET (1993) Differential expression of two related, low-temperature-induced genes in Arabidopsis thaliana (L.) Heynh. Plant Mol Biol 21:641–653. doi:10.1007/BF00014547
Pandey A, Soccol CR, Nigam P, Soccol VT (2000) Biotechnological potential of agro-industrial residues. I: sugarcane bagasse. Bioresour Technol 74:69–80
Parra G, Bradnam K, Rose AB, Korf I (2011) Comparative and functional analysis of intron-mediated enhancement signals reveals conserved features among plants. Nucleic Acids Res 39:5328–5337. doi:10.1093/nar/gkr043
Petrasovits LA, Purnell MP, Nielsen LK, Brumbley SM (2007) Production of polyhydroxybutyrate in sugarcane. Plant Biotechnol J 5:162–172
Petrasovits LA, Zhao L, McQualter RB et al (2012) Enhanced polyhydroxybutyrate production in transgenic sugarcane. Plant Biotechnol J 10:569–578. doi:10.1111/j.1467-7652.2012.00686.x
Preacher KJ (2001) Calculation for the Chi square test: an interactive calculation tool for Chi square tests of goodness of fit and independence [Computer software]. Available from http://quantpsy.org
Prestridge DS (1991) SIGNAL SCAN: a computer program that scans DNA sequences for eukaryotic transcriptional elements. Comput Appl Biosci (CABIOS) 7:203–206. doi:10.1093/bioinformatics/7.2.203
Rae AL, Perroux JM, Grof CPL (2005) Sucrose partitioning between vascular bundles and storage parenchyma in the sugarcane stem: a potential role for the ShSUT1 sucrose transporter. Planta 220:817–825
Rae AL, Casu RE, Perroux JM et al (2011) A soluble acid invertase is directed to the vacuole by a signal anchor mechanism. J Plant Physiol 168:983–989
Rae AL, Martinelli AP, Dornelas MC (2014) Anatomy and morphology. In: Moore PH, Botha FC (eds) Sugarcane: physiology, biochemistry, and functional biology, 1st edn. Wiley, Ames, pp 19–34
Ranocha P, Denancé N, Vanholme R et al (2010) Walls are thin 1 (WAT1), an Arabidopsis homolog of Medicago truncatula NODULIN21, is a tonoplast-localized protein required for secondary wall formation in fibers. Plant J 63:469–483. doi:10.1111/j.1365-313X.2010.04256.x
Rebetzke GJ, Van Herwaarden AF, Jenkins C et al (2008) Quantitative trait loci for water-soluble carbohydrates and associations with agronomic traits in wheat. Aust J Agric Res 59:891–905. doi:10.1071/AR08067
Reinders A, Sivitz AB, Starker CG et al (2008) Functional analysis of LjSUT4, a vacuolar sucrose transporter from Lotus japonicus. Plant Mol Biol 68:289–299
Richmond TA, Somerville CR (2000) The cellulose synthase superfamily. Plant Physiol 124:495–498
Rose AB, Elfersi T, Parra G, Korf I (2008) Promoter-proximal introns in Arabidopsis thaliana are enriched in dispersed signals that elevate gene expression. Plant Cell 20:543–551
Sangnark A, Noomhorm A (2004) Effect of dietary fiber from sugarcane bagasse and sucrose ester on dough and bread properties. LWT Food Sci Technol 37:697–704
Schenk PM, Remans T, Sági L et al (2001) Promoters for pregenomic RNA of banana streak badnavirus are active for transgene expression in monocot and dicot plants. Plant Mol Biol 47:399–412
Schnyder H (1993) The role of carbohydrate storage and redistribution in the source–sink relations of wheat and barley during grain filling—a review. New Phytol 123:233–245. doi:10.1111/j.1469-8137.1993.tb03731.x
Schobert C, Baker L, Szederkényi J et al (1998) Identification of immunologically related proteins in sieve-tube exudate collected from monocotyledonous and dicotyledonous plants. Planta 206:245–252. doi:10.1007/s004250050396
Schulz A, Beyhl D, Marten I et al (2011) Proton-driven sucrose symport and antiport are provided by the vacuolar transporters SUC4 and TMT1/2. Plant J 68:129–136. doi:10.1111/j.1365-313X.2011.04672.x
Sindhu A, Langewisch T, Olek A et al (2007) Maize brittle stalk2 encodes a COBRA-like protein expressed in early organ development but required for tissue flexibility at maturity. Plant Physiol 145:1444–1459. doi:10.1104/pp.107.102582
Tanaka K, Murata K, Yamazaki M et al (2003) Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol 133:73–83
Taylor NG (2008) Cellulose biosynthesis and deposition in higher plants. New Phytol 178:239–252. doi:10.1111/j.1469-8137.2008.02385.x
Thimm O, Bläsing O, Gibon Y et al (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Usadel B, Nagel A, Thimm O et al (2005) Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes, and comparison with known responses. Plant Physiol 138:1195–1204
Uys L, Botha FC, Hofmeyr JHS, Rohwer JM (2007) Kinetic model of sucrose accumulation in maturing sugarcane culm tissue. Phytochemistry 68:2375–2392
Walsh KB, Sky RC, Brown SM (2005) The anatomy of the pathway of sucrose unloading within the sugarcane stalk. Funct Plant Biol 32:367–374
Wang M-L, Goldstein C, Su W et al (2005) Production of biologically active GM-CSF in sugarcane: a secure biofactory. Transgenic Res 14:167–178
Watt DA, McCormick AJ, Govender C et al (2005) Increasing the utility of genomics in unravelling sucrose accumulation. Field Crops Res 92:149–158
Wormit A, Trentmann O, Feifer I et al (2006) Molecular identification and physiological characterization of a novel monosaccharide transporter from Arabidopsis involved in vacuolar sugar transport. Plant Cell 18:3476–3490
Wu L, Birch RG (2007) Doubled sugar content in sugarcane plants modified to produce a sucrose isomer. Plant Biotechnol J 5:109–117
Xue GP (2002) Characterisation of the DNA-binding profile of barley HvCBF1 using an enzymatic method for rapid, quantitative and high-throughput analysis of the DNA-binding activity. Nucleic Acids Res 30:e77
Yang T, Poovaiah BW (2002) A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J Biol Chem 277:45049–45058. doi:10.1074/jbc.M207941200
Zhang Q, Cheetamun R, Dhugga K et al (2014) Spatial gradients in cell wall composition and transcriptional profiles along elongating maize internodes. BMC Plant Biol 14:27
Acknowledgments
The authors would like to thank Michael Hewitt (CSIRO) for expert technical assistance, Larry Cooper and Dave Olsen (Queensland Department of Agriculture and Fisheries) for transgenic sugarcane care, and Dr. Clare Bolton, Prapat Punpee and Gerard Scalia (BSES Limited, now Sugar Research Australia) for assistance with sugarcane transformation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Casu, R.E., Rae, A.L., Nielsen, J.M. et al. Tissue-specific transcriptome analysis within the maturing sugarcane stalk reveals spatial regulation in the expression of cellulose synthase and sucrose transporter gene families. Plant Mol Biol 89, 607–628 (2015). https://doi.org/10.1007/s11103-015-0388-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0388-9