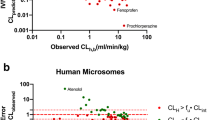
Abstract
Purpose
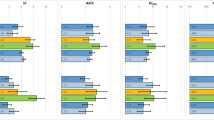
To examine the interlaboratory variability in CLint values generated with human hepatocytes and determine trends in variability and clearance prediction accuracy using physicochemical and pharmacokinetic parameters.
Methods
Data for 50 compounds from 14 papers were compiled with physicochemical and pharmacokinetic parameter values taken from various sources.
Results
Coefficients of variation were as high as 99.8% for individual compounds and variation was not dependent on the number of prediction values included in the analysis. When examining median values, it appeared that compounds with a lower number of rotatable bonds had more variability. When examining prediction uniformity, those compounds with uniform in vivo underpredictions had higher CLint, in vivo values, while those with non-uniform predictions typically had lower CLint, in vivo values. Of the compounds with uniform predictions, only a small number were uniformly predicted accurately. Based on this limited dataset, less lipophilic, lower intrinsic clearance, and lower protein binding compounds yield more accurate clearance predictions.
Conclusions
Caution should be taken when compiling in vitro CLint values from different laboratories as variations in experimental procedures (such as extent of shaking during incubation) may yield different predictions for the same compound. The majority of compounds with uniform in vitro values had predictions that were inaccurate, emphasizing the need for a better mechanistic understanding of IVIVE. The non-uniform predictions, often with low turnover compounds, reaffirmed the experimental challenges for drugs in this clearance range. Separating new chemical entities by lipophilicity, intrinsic clearance, and protein binding may help instill more confidence in IVIVE predictions.





Similar content being viewed by others
Abbreviations
- BDDCS :
-
Biopharmaceutics Drug Disposition Classification System
- CL int :
-
Intrinsic clearance
- CL H :
-
Hepatic clearance
- CV :
-
Coefficient of variation
- fu :
-
Fraction unbound
- HBA :
-
Number of hydrogen bond acceptors
- HBD :
-
Number of hydrogen bond donors
- IVIVE :
-
In vitro to in vivo extrapolation
- MRT :
-
Mean residence time
- MW :
-
Molecular weight
- PSA :
-
Polar surface area
- VD ss :
-
Steady state volume of distribution
References
Bowman CM, Benet LZ. Hepatic clearance predictions from in vitro-in vivo extrapolation and the biopharmaceuitcs drug disposition classification system. Drug Metab Dispos. 2016;44:1731–5.
Wood FL, Houston JB, Hallifax D. Clearance prediction methodology needs fundamental improvement: trends common to rat and human hepatocytes/microsomes and implications for experimental methodology. Drug Metab Dispos. 2017;45:1178–88.
Sohlenius-Sternbeck AK, Jones C, Ferguson D, Middleton BJ, Projean D, Floby E, et al. Practical use of the regression offset approach for the prediction of in vivo intrinsic clearance from hepatocytes. Xenobiotica. 2012;42:841–53.
Nagilla R, Frank KA, Jolivette LJ, Ward KW. Investigation of the utility of published in vitro intrinsic clearance data for prediction of in vivo clearance. J Pharmacol Toxicol Methods. 2006;53:106–16.
Stringer R, Nicklin PL, Houston JB. Reliability of human cryopreserved hepatocytes and liver microsomes as in vitro systems to predict metabolic clearance. Xenobiotica. 2008;38:1313–29.
Hakooz N, Ito K, Rawden H, Gill H, Lemmers L, Boobis AR, et al. Determination of a human hepatic microsomal scaling factor for predicting in vivo drug clearance. Pharm Res. 2006;23:533–9.
Barter ZE, Bayliss MK, Beaune PH, Boobis AR, Carlile DJ, Edwards RJ, et al. Scaling factors for the extrapolation of in vivo metabolic drug clearance from in vitro data: reaching a consensus on values of human microsomal protein and hepatocellularity per gram liver. Curr Drug Metab. 2007;8:33–45.
Hallifax D, Galetin A, Houston JB. Prediction of metabolic clearance using fresh human hepatocytes: comparison with cryopreserved hepatocytes and hepatic microsomes for five benzodiazepines. Xenobiotica. 2008;38:353–67.
Floby E, Johansson J, Hoogstraate J, Hewitt NJ, Hill J, Sohlenius-Sternbeck A-K. Comparison of intrinsic metabolic clearance in fresh and cryopreserved human hepatocytes. Xenobiotica. 2009;39:656–62.
Akabane T, Gerst N, Naritomi Y, Masters JN, Tamura K. A practical and direct comparison of intrinsic metabolic clearance of several non-CYP enzyme substrates in freshly isolated and cryopreserved hepatocytes. Drug Metab Pharmacokinet. 2012;27:181–91.
Blanchard N, Alexandre E, Abadie C, Lavé T, Heyd B, Mantion G, et al. Comparison of clearance predictions using primary cultures and suspensions of human hepatocytes. Xenobiotica. 2005;35:1–15.
Hallifax D, Rawden HC, Hakooz N, Houston JB. Prediction of metabolic clearance using cryopreserved human hepatocytes: kinetic characteristics for five benzodiazepines. Drug Metab Dispos. 2005;33:1852–8.
Jacobson L, Middleton B, Holmgren J, Eirefelt S, Fröjd M, Blomgren A, et al. An optimized automated assay for determination of metabolic stability using hepatocytes: assay validation, variance component analysis, and in vivo relevance. Assay Drug Dev Technol. 2007;5:403–15.
Lau YY, Sapidou E, Cui X, White RE, Cheng K-C. Development of a novel in vitro model to predict hepatic clearance using fresh, cryopreserved, and sandwich-cultured hepatocytes. Drug Metab Dispos. 2002;30:1446–54.
Lu C, Li P, Gallegos R, Uttamsingh V, Xia CQ, Miwa GT, et al. Comparison of intrinsic clearance in liver microsomes and hepatocytes from rats and humans: evaluation of free fraction and uptake in hepatocytes. Drug Metab Dispos. 2006;34:1600–5.
McGinnity DF, Soars MG, Urbanowicz RA, Riley RJ. Evaluation of fresh and cryopreserved hepatocytes as in vitro drug metabolism tools for the prediction of metabolic clearance. Drug Metab Dispos. 2004;32:1247–53.
Naritomi Y, Terashita S, Kagayama A, Sugiyama Y. Utility of hepatocytes in predicting drug metabolism: comparison of hepatic intrinsic clearance in rats and humans in vivo and in vitro. Drug Metab Dispos. 2003;31:580–8.
Riley RJ, McGinnity DF, Austin RP. A unified model for predicting human hepatic, metabolic clearance from in vitro intrinsic clearance data in hepatocytes and microsomes. Drug Metab Dispos. 2005;33:1304–11.
Soars MG, Burchell B, Riley RJ. In vitro analysis of human drug glucuronidation and prediction of in vivo metabolic clearance. J Pharmacol Exp Ther. 2002;301:382–90.
Sohlenius-Sternbeck A-K, Afzelius L, Prusis P, Neelissen J, Hoogstraate J, Johansson J, et al. Evaluation of the human prediction of clearance from hepatocyte and microsome intrinsic clearance for 52 drug compounds. Xenobiotica. 2010;40:637–49.
Gaulton A, Hersey A, Nowotka M, Bento AP, Chambers J, Mendez D, et al. The ChEMBL database in 2017. Nucleic Acids Res. 2017;45:D945–54.
Obach RS, Lombardo F, Waters NJ. Trend analysis of a database of intravenous pharmacokinetic parameters in humans for 670 drug compounds. Drug Metab Dispos. 2008;36:1385–405.
Benet LZ, Broccatelli F, Oprea TI. BDDCS applied to over 900 drugs. AAPS J. 2011;13:519–47.
Hosey CM, Chan R, Benet LZ. BDDCS predictions, self-correcting aspects of BDDCS assignments, BDDCS assignment corrections, and classification for more than 175 additional drugs. AAPS J. 2016;18:251–60.
El-Kattan AF, Varma MV, Steyn SJ, Scott DO, Maurer TS, Bergman A. Projecting ADME behavior and drug-drug interactions in early discovery and development: application of the extended clearance classification system. Pharm Res. 2016;33:3021–33.
Houston JB, Carlile DJ. Prediction of hepatic clearance from microsomes, hepatocytes, and liver slices. Drug Metab Rev. 1997;29:891–922.
Obach RS, Baxter JG, Liston TE, Silber BM, Jones BC, MacIntyre F, et al. The prediction of human pharmacokinetic parameters from preclinical and in vitro metabolism data. J Pharmacol Exp Ther. 1997;283:46–58.
Verber DF, Johnson SR, Cheng H-Y, Smith BR, Ward KW, Kopple KD. Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem. 2002;45:2615–23.
Wood FL, Houston JB, Hallifax D. Importance of the unstirred water layer and hepatocyte membrane integrity in vitro for quantification of intrinsic metabolic clearance. Drug Metab Dispos. 2018;46:268–78.
Clark DE. Rapid calculation of polar molecular surface area and its application to the prediction of transport phenomena. 1. Prediction of intestinal absorption. J Pharm Sci. 1999;88:807–14.
Hallifax D, Foster JA, Houston JB. Prediction of human metabolic clearance from in vitro systems: retrospective analysis and prospective view. Pharm Res. 2010;27:2150–61.
Foster JA, Houston JB, Hallifax D. Comparison of intrinsic clearances in human liver microsomes and suspended hepatocytes from the same donor livers: clearance-dependent relationship and implications for prediction of in vivo clearance. Xenobiotica. 2011;41:124–36.
Di L, Obach RS. Addressing the challenges of low clearance in drug research. AAPS J. 2015;17:352–7.
Baker M, Parton T. Kinetic determinants of hepatic clearance: plasma protein binding and hepatic uptake. Xenobiotica. 2007;37:1110–34.
Soars MG, McGinnity DF, Grime K, Riley RJ. The pivotal role of hepatocytes in drug discovery. Chem Biol Interact. 2007;168:2–15.
Acknowledgments and Disclosures
CMB was supported by the National Science Foundation Graduate Research Fellowship Program [Grant 1144247] and a Pharmaceutical Research and Manufacturers of America Foundation Pre-doctoral Fellowship in Pharmaceutics; LZB is a member of the UCSF Liver Center supported by NIH Grant [P30 DK026743].
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 22.9 kb)
Rights and permissions
About this article
Cite this article
Bowman, C.M., Benet, L.Z. Interlaboratory Variability in Human Hepatocyte Intrinsic Clearance Values and Trends with Physicochemical Properties. Pharm Res 36, 113 (2019). https://doi.org/10.1007/s11095-019-2645-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11095-019-2645-0