Abstract
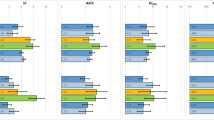
Extensive studies have been conducted to predict in vivo metabolic clearance from in vitro human liver metabolism parameters (i.e., in vitro–in vivo extrapolation (IVIVE)) with little success. Here, deriving IVIVE from first principles, we show that the product of fraction unbound in the blood and the predicted in vivo intrinsic clearance determined from hepatocyte or microsomal incubations is the lower boundary condition for in vivo hepatic clearance and the prerequisite for IVIVE predictions to be valid, regardless of extraction ratio. For 60–80% of drugs evaluated here, this product is markedly less than the in vivo measured clearance, a result that violates the lower boundary of the predictive relationship. This can only be explained by (a) suboptimal in vitro metabolic stability assay conditions, (b) significant error in the assumption that in vitro intrinsic clearance determinations will predict in vivo intrinsic clearance simply by scaling-up the amount of enzyme (in vitro incubation to in vivo liver), and/or (c) the methods of determining fraction unbound are incorrect. We further suggest that widely employed organ blood flow values underpredict the effective blood flow within the organ by approximately 2.5-fold, thus impacting IVIVE of high clearance compounds. We propose future pathways that should be investigated in terms of the relationship to experimentally measured clearance values, rather than model-dependent intrinsic clearance. IVIVE outcome can be improved by estimating the ratio of unbound drug concentration in the liver tissue to the liver plasma, examining the assumption of the free drug theory (i.e., there are no transporter effects at the blood cell membrane) and the finding that the upper limit of organ clearance may be greater than blood flow entering the organ.



Similar content being viewed by others
Abbreviations
- ADME:
-
Absorption, distribution, metabolism, excretion
- AUC:
-
Area under the concentration–time curve
- BDDCS:
-
Biopharmaceutics Drug Disposition Classification System
- \( \frac{B}{P} \) :
-
Blood to plasma partitioning ratio
- CL:
-
Clearance
- CLint :
-
Intrinsic clearance
- CYP3A:
-
Cytochrome P450 3A
- D :
-
Dose
- ECCS:
-
Extended Clearance Classification System
- F :
-
Bioavailability
- f u,B :
-
Fraction of unbound drug in the blood
- f u, inc :
-
Fraction of unbound drug in an in vitro incubation
- f u,P :
-
Fraction of unbound drug in the plasma
- IVIVE:
-
In vitro–in vivo extrapolation
- NME:
-
New molecular entity
- Q H :
-
Hepatic blood flow
References
Page KM. Validation of early human dose prediction: a key metric for compound progression in drug discovery. Mol Pharm. 2015;13:609–20.
Krauss M, Hofmann U, Schafmayer C, Igel S, Schlender J, Mueller C, et al. Translational learning from clinical studies predicts drug pharmacokinetics across patient populations. NPJ Syst Biol Appl. 2017;3:11.
Rané A, Wilkinson GR, Shand DG. Prediction of hepatic clearance ratio from in vitro measurement of intrinsic clearance. J Pharmacol Exp Ther. 1977;200:420–4.
Houston JB. Utility of in vitro drug metabolism data in predicting in vivo metabolic clearance. Biochem Pharmacol. 1994;47:1469–79.
Houston JB. Prediction of hepatic clearance from microsomes, hepatocyte and liver slices. Drug Metab Rev. 1997;29:891–922.
Iwatsubo T, Hirota N, Ooie T, Suzuki H, Shimada N, Chiba K, et al. Prediction of in vivo drug metabolism in the human liver from in vitro metabolism data. Pharmacol Ther. 1997;73:147–71.
Obach RS, Baxter JG, Liston TE, Silber BM, Jones BC, MacIntyre F, et al. The prediction of human pharmacokinetic parameters from preclinical and in vitro metabolism data. J Pharmacol Exp Ther. 1997;283:46–58.
Obach RS. Prediction of human clearance of twenty-nine drugs from hepatic microsomal intrinsic clearance data: an examination of in vitro half-life approach and nonspecific binding to microsomes. Drug Metab Dispos. 1999;27:1350–9.
Bowman CM, Benet LZ. Hepatic clearance predictions from in vitro-in vivo extrapolation and the Biopharmaceutics Drug Disposition Classification System. Drug Metab Dispos. 2016;44:1731–5.
Wood FL, Houston JB, Hallifax D. Clearance prediction methodology needs fundamental improvement: trends common to rat and human hepatocytes/microsomes and implications for experimental methodology. Drug Metab Dispos. 2017;45:1178–88.
Takano J, Maeda K, Bolger MB, Sugiyama Y. The prediction of the relative importance of CYP3A/P-gp to the nonlinear intestinal absorption of drugs by advanced compartmental absorption and transit model. Drug Metab Dispos. 2016;44:1808–18.
Chiba M, Ishii Y, Sugiyama Y. Prediction of hepatic clearance in human from in vitro data for successful drug development. AAPS J. 2009;2:262–76.
Wood FL, Houston JB, Hallifax D. Importance of the unstirred water layer and hepatocyte membrane integrity in vitro for quantification of intrinsic metabolic clearance. Drug Metab Dispos. 2018;46:268–78.
Hallifax D, Houston JB. Evaluation of hepatic clearance prediction using in vitro data: emphasis on fraction unbound in plasma and drug ionization using a database of 107 drugs. J Pharm Sci. 2012;101:2645–52.
Riccardi KA, Tess DA, Lin J, Patel R, Ryu R, Atkinson K, et al. A novel unified approach to predict human hepatic clearance from both enzyme- and transporter-mediated mechanisms using suspended human hepatocytes. Drug Metab Dispos. 2019;47:484–92.
Rowland M, Benet LZ, Graham GG. Clearance concepts in pharmacokinetics. J Pharmacokinet Biopharm. 1973;1:123–35.
Wilkinson GR, Shand DG. Commentary: a physiologic approach to hepatic drug clearance. Clin Pharmacol Ther. 1975;18:377–90.
Rowland M, Tozer TN. Clinical pharmacokinetics and pharmacodynamics: concepts and applications. 4th ed. Philadelphia: Lippincott Williams & Wilkens; 2010.
Klotz U, Avant GR, Hoyumpa A, Schenker S, Wilkinson GR. The effects of age and liver disease on the disposition and elimination of diazepam in adult man. J Clin Invest. 1975;55:347–59.
Mangoni AA, Jackson SHD. Age-related changes in pharmacokinetics and pharmacodynamics: basic principles and practical applications. Br J Clin Pharmacol. 2004;57:6–14.
Wu C-Y, Benet LZ. Predicting drug disposition via application of BCS: transport/absorption/elimination interplay and development of a Biopharmaceutics Drug Disposition Classification System. Pharm Res. 2005;22:11–23.
Hallifax D, Houston JP. Use of segregated hepatocyte scaling factors and cross-species relationships to resolve clearance dependence in the prediction of human hepatic clearance. Drug Metab Dispos. 2019;47:320–7.
Bowman CM, Benet LZ. Interlaboratory variability in human hepatocyte intrinsic clearance values and trends with physicochemical properties. Pharm Res. 2019;36:113.
Bowman CM, Benet LZ. In vitro-in vivo extrapolation and hepatic clearance-dependent underprediction. J Pharm Sci. 2019;108:2500–4.
Bowman CM, Benet LZ. In vitro-in vivo inaccuracy: the CYP3A4 anomaly. Drug Metab Dispos. 2019;47:1368–71.
Bowman CM, Benet LZ. An examination of protein binding and protein-facilitated uptake relating to in vitro-in vivo extrapolation. Eur J Pharm Sci. 2018;123:502–14.
Bowman CM, Okochi H, Benet LZ. The presence of a transporter-induced protein binding shift: a new explanation for protein-facilitated uptake and improvement for in vitro-in vivo extrapolation. Drug Metab Dispos. 2019;47:358–63.
Benet LZ, Liu S, Wolfe AR. The universally unrecognized assumption in predicting drug clearance and organ extraction ratio. Clin Pharmacol Ther. 2018;103:521–5.
Lombardo F, Berellini G, Obach RS. Trend analysis of a database of intravenous pharmacokinetic parameters in humans for 1352 drug compounds. Drug Metab Dispos. 2018;46:1466–77.
Acknowledgments
The authors thank members of our laboratory Dr. Jin Dong (present address: Simulations Plus), Alan R. Wolfe, Shufang Liu (present address: SUNY Buffalo), and Dr. Giovani Bocci (present address: University of New Mexico) for the very helpful discussions.
Funding
This work was supported in part by a Mary Anne Koda-Kimble Seed Award for Innovation. Dr. Benet is a member of the UCSF Liver Center supported by NIH Grant P30 DK026743. Ms. Sodhi was supported in part by an American Foundation for Pharmaceutical Education Pre-Doctoral Fellowship, NIGMS Grant R25 GM56847 and a Louis Zeh Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Benet, L.Z., Sodhi, J.K. Investigating the Theoretical Basis for In Vitro–In Vivo Extrapolation (IVIVE) in Predicting Drug Metabolic Clearance and Proposing Future Experimental Pathways. AAPS J 22, 120 (2020). https://doi.org/10.1208/s12248-020-00501-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-020-00501-9