Abstract
Background
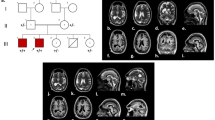
Primary autosomal recessive microcephaly (MCPH) is a rare developmental disorder characterized by cognitive impairment, delayed neurodevelopment, and reduced brain size. It is a genetically heterogeneous condition, and several genes have been identified as associated with MCPH.
Methods and results
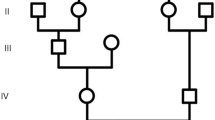
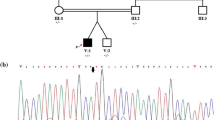
In this study, we utilized whole-exome sequencing (WES) to identify disease-causing variations in two brothers from an Iranian family affected by MCPH, who had consanguineous parents. In the patients, we detected a novel homozygous missense mutation (c.806A > G, p.Gln269Arg) in the TEDC1 gene in one of the patients. Co-segregation analysis using Sanger sequencing confirmed that this variant was inherited from parents. The identified variant was evaluated for its pathogenicity and novelty using various databases. Additionally, bioinformatics tools were employed to predict the three-dimensional structure of the mutant TEDC1 protein.
Conclusions
This study presents the second documented report of a mutation in the TEDC1 gene associated with MCPH. The identification of this novel biallelic mutation as a causative factor for MCPH in the proband further underscores the utility of genetic testing techniques, such as WES, as reliable diagnostic tools for individuals with this condition.




Similar content being viewed by others
Data availability
The raw data supporting the conclusions of this manuscript will be made available to any qualified researcher upon contact with the corresponding author.
References
Jayaraman D, Bae BI, Walsh CA (2018) The genetics of primary microcephaly. Annu Rev Genomics Hum Genet 19:177–200
Hanzlik E, Gigante J (2017) Microcephaly. Children 4(6):47
Passemard S, Kaindl AM, Verloes A (2013) Microcephaly. Handbook of clinical neurology. Elsevier, Amsterdam, pp 129–141
Adachi Y et al (2011) Congenital microcephaly with a simplified gyral pattern: associated findings and their significance. AJNR Am J Neuroradiol 32(6):1123–1129
Jean F, Stuart A, Tarailo-Graovac M (2020) Dissecting the genetic and etiological causes of primary microcephaly. Front Neurol 11:570830
Becerra-Solano LE, Mateos-Sánchez L, López-Muñoz E (2021) Microcephaly, an etiopathogenic vision. Pediatr Neonatol 62(4):354–360
Devakumar D, Bamford A, Ferreira MU, Broad J, Rosch RE, Groce N, Breuer J, Cardoso MA, Copp AJ, Alexandre P, Rodrigues LC, Abubakar I (2018) Infectious causes of microcephaly: epidemiology, pathogenesis, diagnosis and management. Lancet Infect Dis 18(1):e1–e13
Woods CG, Parker A (2013) Investigating microcephaly. Arch Dis Child 98(9):707–713
Harris SR (2015) Measuring head circumference: update on infant microcephaly. Can Fam Physician 61(8):680–684
Von der Hagen M et al (2014) Diagnostic approach to microcephaly in childhood: a two-center study and review of the literature. Dev Med Child Neurol 56(8):732–741
Zaqout S, Kaindl AM (2021) Autosomal recessive primary microcephaly: not just a small brain. Front Cell Dev Biol 9:784700
Stark AE (2016) The genetic epidemiology of the form of microcephaly ascribed to mutation at the WDR62 locus. Ann Transl Med 4(15):281
Iegiani G et al (2023) The impact of TP53 activation and apoptosis in primary hereditary microcephaly. Front Neurosci 17:10
Zaqout S, Morris-Rosendahl D, Kaindl AM (2017) Autosomal recessive primary microcephaly (MCPH): an update. Neuropediatrics 48(03):135–142
Siskos N et al (2017) Molecular genetics of microcephaly primary hereditary: an overview. Brain Sci 11(5):581
Naveed M et al (2018) Comprehensive review on the molecular genetics of autosomal recessive primary microcephaly (MCPH). Genet 100:e7
Dawidziuk M et al (2021) Exome sequencing reveals novel variants and expands the genetic landscape for congenital microcephaly. Genes (Basel). https://doi.org/10.3390/genes12122014
Boonsawat P et al (2019) Elucidation of the phenotypic spectrum and genetic landscape in primary and secondary microcephaly. Genet Med 21(9):2043–2058
Stathatos GG et al (2021) Delta and epsilon tubulin in mammalian development. Trends Cell Biol 31(9):774–787
Szklarczyk D et al (2021) The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49(D1):D605–D612
Engels H et al (2012) A phenotype map for 14q32.3 terminal deletions. Am J Med Genet A 158(4):695–706
Breslow DK et al (2018) A CRISPR-based screen for Hedgehog signaling provides insights into ciliary function and ciliopathies. Nat Genet 50(3):460–471
Gergely F (2019) Centriolar satellites are acentriolar assemblies of centrosomal proteins. EMBO J 38(14):e101082
Filipuzzi I et al (2017) Direct interaction of Chivosazole F with actin elicits cell responses similar to Latrunculin A but distinct from chondramide. ACS Chem Biol 12(9):2264–2269
Kataoka M et al (2016) Exome sequencing for bipolar disorder points to roles of de novo loss-of-function and protein-altering mutations. Mol Psychiatry 21(7):885–893. https://doi.org/10.1038/mp.2016.69
Acknowledgements
We would like to express our appreciation to the patient, their family members, and all the individuals who participated in this study for their contribution.
Funding
The current study was financially supported by Golestan University of Medical Sciences (Grant Number 113339).
Author information
Authors and Affiliations
Contributions
AS: Conceptualization, Methodology, Formal analysis, Data curation. ZMAS: Methodology, Investigation, Formal analysis, Writing—original draft. FV: Methodology, Investigation, Formal analysis, Writing—original draft. FM: Methodology, Investigation, Formal analysis, Writing—original draft. TK: Visualization, Methodology, Investigation, Data curation, Writing—original draft. NR: Methodology, Investigation, Data Collection. MO: Conceptualization, Supervision, Validation, Writing—review and editing.
Corresponding author
Ethics declarations
Conflicts of interest
There is none to declare.
Ethical approval
Our investigation was conducted in accordance with the guidelines set forth by the Ethics Committee of Golestan University of Medical Sciences (Ethics Code: IR.GOUMS.REC.1401.540).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sarli, A., Al Sudani, Z.M., Vaghefi, F. et al. Second report of TEDC1-related microcephaly caused by a novel biallelic mutation in an Iranian consanguineous family. Mol Biol Rep 51, 181 (2024). https://doi.org/10.1007/s11033-023-09136-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-09136-3