Abstract
Background
Plant microbiome acts as an interface between plants and their environment, aiding in the functioning of the ecosystem, such as protection against abiotic and biotic stress along with improving nutrient uptake. The rhizosphere is an essential interface for the interaction between plants and microbes and plays a substantial part in the removal as well as uptake of heavy metals and antibiotics from contaminated locations. Eichhornia crassipes is a promising plant that contains a rich community of microbes in its rhizosphere. Microorganism’s association with plants embodies a crucial pathway via which humans can also be exposed to antibiotic-resistant genes and bacteria.
Methods and results
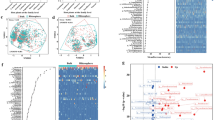
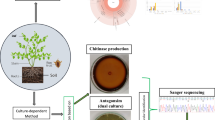
In our earlier study enhanced removal of ciprofloxacin was observed by plant growth-promoting Microbacterium sp. WHC1 in the presence of E. crassipes root exudates. Therefore, the V3-V4, hypervariable region of the 16 S rRNA gene was studied to assess the bacterial diversity and functional profiles of the microbiota associated with plant roots. Using the QIIME software program, 16 S rRNA data from the Next Generation Sequencing (NGS) platform was examined. Alpha diversity including Chao1, Observed Shannon, and Simpson index denote significantly higher bacterial diversity. Proteobacteria (79%) was the most abundant phylum which was present in the root samples followed by Firmicutes (8%) and Cyanobacteria (8%). Sulfuricurvum (36%) is the most abundant genus belonging to the family Helicobacteraceae and the species kujiense in the genus Sulfuricurvum is the most abundant species present in the root sample. Also, the bacterial communities in the rhizoplane of Eichhornia crassipes harbor the genes conferring resistance to beta-lactams, tetracycline, fluoroquinolones, and penams.
Conclusion
Metagenomic studies on the E. crassipes microbiome showed that the bacterial communities constituting the root exudates of the Eichhornia aid them to survive in a polluted environment.









Similar content being viewed by others
Data availability
The Illumina sequencing data has been submitted to NCBI with Bio-project ID: PRJNA926139.
Change history
01 February 2024
A Correction to this paper has been published: https://doi.org/10.1007/s11033-024-09227-9
References
Amalina F, Abd Razak AS, Krishnan S, Zularisam AW, Nasrullah M (2022) Water hyacinth (Eichhornia crassipes) for organic contaminants removal in water–a review. J Hazard Mater Adv 7:100092
Madikizela LM (2021) Removal of organic pollutants in water using water hyacinth (Eichhornia crassipes). J Environ Manage 295:113153
Samal K, Kar S, Trivedi S (2019) Ecological floating bed (EFB) for decontamination of polluted water bodies: design, mechanism and performance. J Environ Manage 251:109550
Sasmaz M, Topal EIA, Obek E, Sasmaz A (2015) The potential of Lemna gibba L. and Lemna minor L. to remove Cu, Pb, Zn, and as in gallery water in a mining area in Keban, Turkey. J Environ Manage 163:246–253
Mechora Š, Stibilj V, Germ M (2015) Response of duckweed to various concentrations of selenite. Environ Sci Pollut Res 22:2416–2422
Di Luca GA, Mufarrege MM, Hadad HR, Maine MA (2019) Nitrogen and phosphorus removal and Typha domingensis tolerance in a floating treatment wetland. Sci Total Environ 650:233–240
Manorama Thampatti KC, Beena VI, Meera AV, Ajayan AS (2020) Phytoremediation of metals by aquatic macrophytes. Phytoremediation. https://doi.org/10.1007/978-3-030-00099-8_6
Deng F, Zeng F, Shen Q, Abbas A, Cheng J, Jiang W, Chen ZH (2022) Molecular evolution and functional modification of plant miRNAs with CRISPR. Trends Plant Sci 27(9):890–907
Sodhi KK, Kumar M, Dhaulaniya AS, Balan B, Singh DK (2021) Enhanced ciprofloxacin removal by plant growth-promoting Microbacterium sp. WHC1 in presence of Eichhornia crassipes root exudates. Environ Sustain 4:143–153
Aydin S, Arabacı DN, Shahi A, Fakhri H, Ovez S (2022) Enhanced removal of antibiotics using Eichhornia crassipes root biomass in an aerobic hollow-fiber membrane bioreactor. Biofouling 38(3):223–234
Velpandian T, Halder N, Nath M, Das U, Moksha L, Gowtham L, Batta SP (2018) Un-segregated waste disposal: an alarming threat of antimicrobials in surface and ground water sources in Delhi. Environ Sci Pollut Res 25:29518–29528
Sodhi KK, Kumar M, Balan B, Dhaulaniya AS, Singh DK (2020) Isolation and characterization of amoxicillin-resistant bacteria and amoxicillin-induced alteration in its protein profiling and RNA yield. Arch Microbiol 202:225–232
Olivares-Rieumont S, Lima L, De la Rosa D, Graham DW, Columbie I, Santana JL, Sánchez MJ (2007) Water hyacinths (Eichhornia crassipes) as indicators of heavy metal impact of a large landfill on the Almendares river near Havana, Cuba. Bull Environ Contam Toxicol 79:583–587
Yuan S, Cohen DB, Ravel J, Abdo Z, Forney LJ (2012) Evaluation of methods for the extraction and purification of DNA from the human microbiome. PLoS ONE 7(3):e33865
Mittal P, Prasoodanan PK, Dhakan V, Kumar DB, Sharma VK (2019) Metagenome of a polluted river reveals a reservoir of metabolic and antibiotic resistance genes. Environ Microbiome 14:1–12
Sodhi KK, Kumar M, Singh DK (2021) Assessing the bacterial diversity and functional profiles of the river Yamuna using Illumina MiSeq sequencing. Arch Microbiol 203:367–375
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet C, Al-Ghalith GA, Caporaso JG (2018) QIIME 2: reproducible, interactive, scalable, and extensible microbiome data science. Nat Biotechnol 37(8):852–857
Balvočiūtė M, Huson DH (2017) SILVA, RDP, greengenes, NCBI and OTT—how do these taxonomies compare? BMC Genomics 18(2):1–8
Wickham H (2011) ggplot2. Wiley Interdiscip Rev 3(2):180–185
Ondov BD, Bergman NH, Phillippy AM (2011) Interactive metagenomic visualization in a web browser. BMC Bioinformatics 12(1):1–10
Liu D, Xu Z, Fan C, Zhou Y (2021) Development of fire risk visualization tool based on heat map. J Loss Prev Process Ind 71:104505
Thukral AK (2017) A review on measurement of alpha diversity in biology. Agric Res J 54(1):1–10
Willis AD (2019) Rarefaction, alpha diversity, and statistics. Front Microbiol 10:2407
Xiang X, Wang H, Tian W, Wang R, Gong L, Xu Y, Man B (2023) Composition and function of bacterial communities of bryophytes and their underlying sediments in the dajiuhu peatland, central China. J Earth Sci 34(1):133–144
Regueira-Iglesias A, Balsa-Castro C, Blanco-Pintos T, Tomás I (2023) Critical review of 16S rRNA gene sequencing workflow in microbiome studies: from primer selection to advanced data analysis. Mol Oral Microbiol. https://doi.org/10.1111/omi.12434
Ortiz-Estrada ÁM, Gollas-Galván T, Martínez-Córdova LR, Martínez-Porchas M (2019) Predictive functional profiles using metagenomic 16S rRNA data: a novel approach to understanding the microbial ecology of aquaculture systems. Rev Aquac 11(1):234–245
Douglas GM, Beiko RG, Langille MG (2018) Predicting the functional potential of the microbiome from marker genes using PICRUSt. Microbiome Anal. https://doi.org/10.1007/978-1-4939-8728-3_11
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, Von Mering C, Bork P (2017) Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol Biol Evol 34(8):2115–2122
Yin Y, Wang J (2021) Predictive functional profiling of microbial communities in fermentative hydrogen production system using PICRUSt. Int J Hydrog Energy 46(5):3716–3725
Alcock BP, Raphenya AR, Lau TT, Tsang KK, Bouchard M, Edalatmand A et al (2020) CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res 48(D1):D517–D525
Sodhi KK, Mishra LC, Singh CK, Kumar M (2022) Perspective on the heavy metal pollution and recent remediation strategies. Curr Res Microb Sci. https://doi.org/10.1016/j.crmicr.2022.100166
Sodhi KK, Singh CK (2022) Recent development in the sustainable remediation of antibiotics: a review. Total Environ Res Themes. https://doi.org/10.1016/j.totert.2022.100008
Sodhi KK, Singh CK, Kumar M, Singh DK (2023) Whole-genome sequencing of Alcaligenes sp. strain MMA: insight into the antibiotic and heavy metal resistant genes. Front Pharmacol 14:1144561. https://doi.org/10.3389/fphar.2023.1144561
Bhat SH, Darzi AB, Dar MS, Ganaie MM, Bakhshi SH (2011) Correlation of soil physico-chemical factors with VAM fungi distribution under different agroecological conditions. Int J Pharma Bio Sci 2(2):107
Das A, David AA, Swaroop N, Thomas T, Rao S, Hasan A (2018) Assessment of physico-chemical properties of river bank soil of Yamuna in Allahabad city, Uttar Pradesh. Int J Chem Stud 6(3):2412–2417
Liao K, Wu S, Zhu Q (2016) Can soil pH be used to help explain soil organic carbon stocks? CLEAN–Soil Air Water 44(12):1685–1689
Neina D (2019) The role of soil pH in plant nutrition and soil remediation. Appl Environ Soil Sci 2019:1–9
Fu M, Zheng L (2016) Effects of different forms of nitrogen on rhizosphere microbial community structure of Eichhorniacrassipes (Pontederiaceae). Rev Biol Trop 64(1):213–220
Ávila MP, Oliveira-Junior ES, Reis MP, Hester ER, Diamantino C, Veraart AJ et al (2019) The water hyacinth microbiome: link between carbon turnover and nutrient cycling. Microbial Ecol 78:575–588
Fulekar MH, Fulekar J (2020) Bioremediation technology: hazardous waste management. CRC Press, Florida
Kumar A, Devi S, Agrawal H, Singh S, Singh J (2020) Rhizoremediation: a unique plant microbiome association of biodegradation. Plant Microbe Symbiosis. https://doi.org/10.1007/978-3-030-36248-5_11
Ma Y, Rajkumar M, Zhang C, Freitas H (2016) Beneficial role of bacterial endophytes in heavy metal phytoremediation. J Environ Manage 174:14–25
Ojuederie OB, Babalola OO (2017) Microbial and plant-assisted bioremediation of heavy metal polluted environments: a review. Int J Environ Res Public Health 14(12):1504
Irawati W, Parhusip AJN, Sopiah N, Tnunay JA (2017) The role of heavy metals-resistant bacteria acinetobacter sp. in copper phytoremediation using Eichhornia crasippes [(Mart.) solms]. KnE Life Sci. https://doi.org/10.18502/kls.v3i5.995
Kaur P, Singh S, Kumar V, Singh N, Singh J (2018) Effect of rhizobacteria on arsenic uptake by macrophyte Eichhorniacrassipes (Mart.) solms. Int J Phytoremediation 20(2):114–120
Sharma R, Kumar A, Singh N, Sharma K (2021) 16S rRNA gene profiling of rhizospheric microbial community of Eichhorniacrassipes. Mol Biol Rep 48(5):4055–4064
Al-Dabbagh B, Elhaty IA, Elhaw M, Murali C, Al Mansoori A, Awad B, Amin A (2019) Antioxidant and anticancer activities of chamomile (Matricaria recutita L). BMC Res Notes 12(1):1–8
Kong N, Wang Z (2022) Response of plant diversity of urban remnant mountains to surrounding urban spatial morphology: a case study. Urban Ecosyst. https://doi.org/10.1007/s11252-021-01154-y
Huang D, Qin X, Peng Z, Liu Y, Gong X, Zeng G, Hu Z (2018) Nanoscale zero-valent iron assisted phytoremediation of Pb in sediment: impacts on metal accumulation and antioxidative system of Lolium perenne. Ecotoxicol Environ Safety 153:229–237
Montes-Osuna N, Cernava T, Gómez-Lama Cabanás C, Berg G, Mercado-Blanco J (2022) Identification of volatile organic compounds emitted by two beneficial endophytic pseudomonas strains from olive roots. Plants 11(3):318
Shree P, Singh CK, Sodhi KK, Surya JN, Singh DK (2023) Biofilms: understanding the structure and contribution towards bacterial resistance in antibiotics. Med Microecol. https://doi.org/10.1016/j.medmic.2023.100084
War Nongkhlaw FM, Joshi SR (2014) Epiphytic and endophytic bacteria that promote growth of ethnomedicinal plants in the subtropical forests of Meghalaya, India. Revista De Biología Tropical 62(4):1295–1308
Shahid A, Muzammil S, Aslam B, Ashfaq UA, Hayat S, Bilal M, Khurshid M (2023) Antibiotics and antibiotic-resistant bacteria in the environment: sources and impacts. Degradation of antibiotics and antibiotic-resistant bacteria from various sources. Academic Press, Massachusetts, pp 39–65
Acknowledgements
The authors would like to acknowledge the University of Delhi.
Funding
Not Applicable.
Author information
Authors and Affiliations
Contributions
CKS & KKS: conceptualization, experimentation, writing, and editing. DKS: conceptualization and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable.
Consent for publication
All the authors give consent for publication and declare no conflict of interest.
Research involving human and animal participants
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The second affiliation is corrected as 'Department of Zoology, Sri Guru Tegh Bahadur Khalsa College, University of Delhi, Delhi-110007, India.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, C.K., Sodhi, K.K. & Singh, D.K. Understanding the bacterial community structure associated with the Eichhornia crassipes rootzone. Mol Biol Rep 51, 35 (2024). https://doi.org/10.1007/s11033-023-08979-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-08979-0