Abstract
Background
Aberrant DNA methylation has been implicated in the development of gastric cancer (GC). In our previous study, we demonstrated that fructose-1,6-bisphosphatase-2 (FBP2), an enzyme that suppresses cell glycolysis and growth, is downregulated in GC due to promoter methylation. However, the precise mechanism underlying this process remains unknown. Thus, this study aimed to elucidate the mechanisms involved in FBP2 promoter hypermethylation.
Methods and results
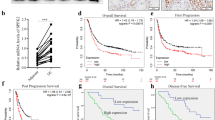
The methylation levels in GC and normal adjacent tissues were quantified using methylation-specific polymerase chain reaction. FBP2 promoter was frequently hypermethylated in primary GC tissues compared to adjacent normal tissues. To explore the functional consequences of this hypermethylation, we employed small interfering RNA-mediated knockdown of DNA methyltransferase 3a (DNMT3a) in GC cells. FBP2 expression increased following DNMT3a knockdown, suggesting that reduced methylation of the FBP2 promoter contributed to this upregulation. To further investigate this interaction, chromatin immunoprecipitation assays were conducted. The results confirmed an interaction between DNMT3a and the FBP2 promoter region, providing evidence that DNMT3a-mediated hypermethylation of the FBP2 promoter promotes GC progression.
Conclusions
This study provides evidence that DNMT3a is involved in the hypermethylation of the FBP2 promoter and regulation of GC cell metabolism. Hypermethylation of the FBP2 promoter may be a promising prognostic biomarker in GC.



Similar content being viewed by others
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
Abbreviations
- DMEM:
-
Dulbecco’s Modified Eagle
- DNMTs:
-
DNA methyltransferases
- FBP:
-
Fructose-1,6-bisphosphatase
- GC:
-
Gastric cancer
- HRP:
-
Horseradish peroxidase
- MSP:
-
Methylation-specific polymerase chain reaction products
- PVDF:
-
Polyvinylidene fluoride
- RIPA:
-
Radioimmunoprecipitation assay
- SD:
-
Standard deviation
- siRNA:
-
Small interfering ribonucleic acid
References
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Smyth EC, Nilsson M, Grabsch HI et al (2020) Gastric cancer. Lancet 396:635–648. https://doi.org/10.1016/s0140-6736(20)31288-5
Nishiyama A, Nakanishi M (2021) Navigating the DNA methylation landscape of cancer. Trends Genet 37:1012–1027. https://doi.org/10.1016/j.tig.2021.05.002
Subhankar Biswas CM (2017) Epigenetics in cancer: fundamentals and beyond. Pharmacol Ther 173:118–134
Dor Y, Cedar H (2018) Principles of DNA methylation and their implications for biology and medicine. Lancet 392:777–786. https://doi.org/10.1016/S0140-6736(18)31268-6
Usui G, Matsusaka K, Mano Y et al (2021) DNA methylation and genetic aberrations in gastric cancer. Digestion 102:25–32. https://doi.org/10.1159/000511243
Sun L, Zhang H, Gao P (2022) Metabolic reprogramming and epigenetic modifications on the path to cancer. Protein Cell 13:877–919. https://doi.org/10.1007/s13238-021-00846-7
Jin H, Liu X, Wang X et al (2009) M1976 Warburg effect revisited: An epigenetic link between aerobic glycolysis and gastric carcinogenesis. Gastroenterology 136:459. https://doi.org/10.1016/s0016-5085(09)62111-9
Li H, Wang J, Xu H et al (2013) Decreased fructose-1,6-bisphosphatase-2 expression promotes glycolysis and growth in gastric cancer cells. Mol Cancer 12:110. https://doi.org/10.1186/1476-4598-12-110
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108. https://doi.org/10.1038/nprot.2008.73
Chandrashekar DS, Bashel B, Balasubramanya SAH et al (2017) UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19:649–658. https://doi.org/10.1016/j.neo.2017.05.002
Chandrashekar DS, Karthikeyan SK, Korla PK et al (2022) UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25:18–27. https://doi.org/10.1016/j.neo.2022.01.001
Győrffy B (2023) Discovery and ranking of the most robust prognostic biomarkers in serous ovarian cancer. GeroScience 45(3):1889–1898. https://doi.org/10.1007/s11357-023-00742-4
Ehrlich M (2019) DNA hypermethylation in disease: mechanisms and clinical relevance. Epigenetics 14(1141):1163
Canale M, Casadei-Gardini A, Ulivi P et al (2020) Epigenetic mechanisms in gastric cancer: Potential new therapeutic opportunities. Int J Mol Sci 21:5500. https://doi.org/10.3390/ijms21155500
Ebrahimi V, Soleimanian A, Ebrahimi T et al (2020) Epigenetic modifications in gastric cancer: Focus on DNA methylation. Gene 742:144577. https://doi.org/10.1016/j.gene.2020.144577
Ge T, Gu X, Jia R et al (2022) Crosstalk between metabolic reprogramming and epigenetics in cancer: updates on mechanisms and therapeutic opportunities. Cancer Commun (Lond) 42:1049–1082. https://doi.org/10.1002/cac2.12374
Thakur C, Chen F (2019) Connections between metabolism and epigenetics in cancers. Semin Cancer Biol 57:52–58. https://doi.org/10.1016/j.semcancer.2019.06.0
Zhang W, Xu J (2017) DNA methyltransferases and their roles in tumorigenesis. Biomark Res 5:1
Zhang J, Yang C, Wu C et al (2020) DNA methyltransferases in cancer: Biology, paradox, aberrations, and targeted therapy. Cancers (Basel) 12:2123. https://doi.org/10.3390/cancers12082123
Zhu A, Hopkins KM, Friedman RA, Bernstock JD, Broustas CG, Lieberman HB (2021) DNMT1 and DNMT3B regulate tumorigenicity of human prostate cancer cells by controlling RAD9 expression through targeted methylation. Carcinogenesis 42:220–231
Chen B-F, Chan W-Y (2014) The de novo DNA methyltransferase DNMT3A in development and cancer. Epigenetics 9:669–677. https://doi.org/10.4161/epi.28324
Lu J, Zhen S, Tuo X et al (2022) Downregulation of DNMT3A attenuates the Warburg effect, proliferation, and invasion via promoting the inhibition of miR-603 on HK2 in ovarian cancer. Technol Cancer Res Treat 21:15330338221110668. https://doi.org/10.1177/15330338221110668
Zhou A, Yin Y, Yu M et al (2022) GTPBP4 promotes hepatocellular carcinoma progression and metastasis via the PKM2 dependent glucose metabolism. Redox Biol 56:102458
Funding
This project received support from the National Natural Science Foundation of China (Grant Number 81802807).
Author information
Authors and Affiliations
Contributions
YH collected data and drafted the manuscript. YH and HL contributed to cell experiment, statistical analysis and interpretation of all the data. HL and HL designed the study and reviewed the manuscript. All authors have read and approved the final manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
None.
Ethical approval
This study was conducted with the approval of the Institutional Ethical Standards Committee of Renji Hospital (No. 2017-114-CR-02).
Consent to participate
All participants provided written informed consent before their inclusion in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, Y., Lu, H. & Li, H. DNA methyltransferase 3a-induced hypermethylation of the fructose-1,6-bisphosphatase-2 promoter contributes to gastric carcinogenesis. Mol Biol Rep 51, 78 (2024). https://doi.org/10.1007/s11033-023-08966-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-08966-5