Abstract
The Southern rice black-streaked dwarf virus (SRBSDV), a novel Fijivirus, poses a major threat to rice production in East Asia. Analysis of the gene expression of SRBSDV-infected rice plants may reveal the molecular basis of interactions between the virus, its vector and rice plants. Reliable reference genes are required for accurate qRT-PCR analysis. However, no reliable, valid reference genes for examining gene expression in SRBSDV-infected rice plants have so far been identified. We examined the expression of eight candidate reference genes in the leaves of SRBSDV-infected, and healthy, rice plants at different points in time after virus inoculation. We used four dedicated algorithms, geNorm, BestKeeper, NormFinder and RefFinder, to evaluate the performance of these candidate genes. The RefFinder results indicate that 18S, EF1α and UBQ10 are suitable reference genes. In addition, we used these three reference genes to analyze the expression of key genes involved in the isoprenoid metabolic pathway in rice leaves after infection by SRBSDV. The results of this analysis reveal that SRBSDV may suppress the production of the rice plant volatiles that attract natural enemies of its vector Sogatella furcifera, thereby increasing the likelihood of pathogen transmission.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The quantitative reverse transcription polymerase chain reaction (qRT-PCR) is the most sensitive, precise and reproducible method for quantifying the mRNA transcript levels of individual genes. However, normalization with reference genes is essential to minimize inconsistencies in protocols and variation in the quality and stability of RNA and enzyme efficiency [1,2,3,4]. For example, numerous studies have demonstrated that the expression levels of some reference genes can vary in different developmental stages and experimental conditions [5,6,7,8]. There is, therefore, an urgent need to identify reference genes suitable for specific developmental stages and experimental conditions. Indeed, more than one reference gene is usually required to normalize qRT-PCR data [9, 10].
In 2008, a devastating disease, the Southern rice black-streaked dwarf virus (SRBSDV), was reported in rice (Oryza sativa) [11]. The rice plants infected with SRBSDV showing dwarfism, dark leaves, twisting of leaf tips, excessive tillering, small waxy swellings on the stalks and leaf-backs, up-growing rootlets on stem nodes, and seriously diseased plants may die of withering [12]. Diseased plants often showing small spikes and barren grains or none at all, causing serious loss in rice production [12, 13]. The white-backed planthopper (WBPH, Sogatella furcifera) is the only known vector of SRBSDV. There has been ongoing research on the effects of SRBSDV on rice and the response of rice to SRBSDV and its vectors [14,15,16,17,18,19]. No one has yet, however, identified suitable reference genes for investigating gene expression in SRBSDV-infected rice plants.
Infection of host cells by a virus causes certain metabolic pathways to be over- or under-expressed resulting in both physiological and phenotypic changes in the host [20,21,22]. Viruses can mediate the host selection behavior of vectors by altering host plant volatiles thereby attracting potential vectors to infected plants [23]. For example, non-viruliferous S. furcifera are more attracted by SRBSDV-infected rice plants than healthy rice plants [24]. Moreover, researchers have revealed that SRBSDV suppresses plant defenses against S. furcifera [15, 16, 19, 25]. Isoprenoids are important components of plant volatiles. However, there has been no research on the mRNA transcript levels of the key genes involved in the isoprenoid metabolic pathway in SRBSDV-infected rice plants.
In this paper, we present the results of experiments designed to identify suitable reference genes in the leaves of SRBSDV-infected, and healthy, rice plants. In addition, we compare the mRNA transcript levels of key genes involved in the isoprenoid metabolic pathway in SRBSDV-infected rice plants with those in healthy rice plants.
Materials and methods
Rice plants, S. furcifera, and SRBSDV materials
The seeds of the rice cultivar ‘Fengyuanyou 299’ used in this study were purchased from Hunan Longping Seed Industry Co., Ltd. A number of 4-leaf stage rice seedlings were individually transplanted into the pots (9 cm in diameter and 15 cm in height). Seedlings were cultured in insect-proof cages in a greenhouse at 24–32 °C. S. furcifera were collected from rice plants in Changsha, Hunan Province, China and reared on healthy rice plants for more than three generation in an illuminated incubator (26 ± 1 °C, 85% relative humidity (RH), 16 h of light and 8 h of darkness). SRBSDV-infected rice plants were kindly provided by Prof. Guohui Zhou (South China Agricultural University, Guangzhou, China).
Collection of healthy, and SRBSDV-infected, rice plants leaves
Third instar S. furcifera nymphs were identified and collected under a stereoscopic microscope. These nymphs were then randomly assigned to two groups, one fed on SRBSDV-infected rice plants to obtain viruliferous S. furcifera, and the other on healthy rice plants to obtain uninfected S. furcifera. Fifteen groups of 10 viruliferous S. furcifera were transferred to individual early-tillering rice plants. Five groups of 10 healthy S. furcifera were also transferred to individual early-tillering rice plants. All insects were subsequently removed after feeding on rice plants for 3 days. 0.2 g of the leaves of each rice plant were collected 0, 15, 25, 35 and 45 days (different disease stages) after insects had been removed and stored at − 80 °C. The presence of SRBSDV in the rice plants inoculated by viruliferous S. furcifera was detected by RT-PCR (reverse-transcription polymerase chain reaction) 45 days after insects had been removed as described previously [24]. Five samples of SRBSDV-infected rice plants were then used in subsequent experiments.
RNA extraction and cDNA synthesis
Total RNA was isolated using the MiniBEST Universal RNA Extraction Kit (TaKaRa, Tokyo, Japan) according to the manufacturer’s instruction. A NanoDrop spectrophotometer (Thermo Scientific NanoDrop 1000, Wilmington, USA) was used to measure the concentration and purity of total RNA. 125 ng of total RNA was used to synthesize first-strand cDNA using the PrimeScript™ RT reagent Kit with gDNA Eraser (TaKaRa), according to the manufacturer’s instructions.
Candidate genes
Eight candidate reference genes (UBQ10, ACTIN, GAPDH, UBQ5, EF1α, 25S, 18S, β-TUB) were selected and their sequences downloaded from the NCBI database (http://www.ncbi.nlm.nih.gov/) [26,27,28]. The sequences of 10 key genes (CMS, DMD, FPPS, GGPPS, SPS3, HDS, IPI, MTC, PHS, SQS) involved in the isoprenoid metabolic pathway were downloaded from the National Rice Data Centre of China (http://www.ricedata.cn/gene/index.htm). The names of these 18 genes are listed in Table 1.
Quantitative real-time PCR
qRT-PCR experiments were carried out using the TB Green Premix Ex Taq™ II (TaKaRa) and a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, Hercules, USA). The qRT-PCR primers of the 18 genes were designed using information on the National Center for Biotechnology Information profile server (http://www.ncbi.nlm.nih.gov/tools/primer-blast). The efficiency of qRT-PCR primers was determined using a fivefold dilution series of healthy rice cDNA. The primers and their amplification efficiencies are shown in Table 1. The qRT-PCR protocol was as follows: preheating for 30 s at 95 °C; 40 cycles of 5 s at 95 °C (denaturation), 30 s at 60 °C (annealing); followed by melting curve analysis (HRM rate = 0.5 °C) after the amplification step.
Data analysis
The stability of expression of each candidate reference gene was ranked using three different types of Microsoft Excel-based software: geNorm [29], Normfinder [30] and BestKeeper [31]. In addition, RefFinder [32] was used to rank the overall stability of all candidate reference genes. The relative expression levels of the key genes involved in the isoprenoid metabolic pathway in rice leaves infected by SRBSDV were quantified using the 2−ΔΔCt method [33]. The statistical significance of differences in mean expression levels between healthy and SRBSDV-infected rice plants was assessed with Student’s t-test in the SPSS program for windows (SPSS Inc., Chicago, USA).
Results
Performance of qRT-PCR primers
A standard curve was generated for each primer pair using the fivefold dilution series of healthy rice cDNA. The amplification efficiency (E) and correlation coefficient (R2) of each primer pair were obtained from the standard curve and are shown in Table 1. The correlation coefficients for all primer pairs ranged from 0.984 to 0.994. All primers had acceptable amplification efficiencies of between 90.4 and 108.8.
Expression levels of candidate reference genes
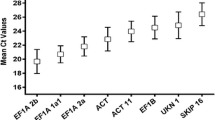
The expression levels of candidate reference genes were quantified using Cq (quantification cycle) values in the leaves of SRBSDV-infected and healthy rice plants. These assays are summarized as Box-Whisker plots in Fig. 1, the box represents the 25th and 75th percentile range and the median is indicated by a horizontal lines across the box. Average Cq values were within 8.93 and 34.72 cycles. 25S had the lowest Cq values and β-TUB the highest.
Expression profiles of the 8 candidate reference genes in leaves of A SRBSDV-infected rice plant and B healthy rice plant. The expression level of candidate reference genes was given as quantification cycle number (Cq-value) and analyzed by using box which represents the 25th and 75th percentiles, the median is indicated by a horizontal lines across the box
Stability of candidate reference genes
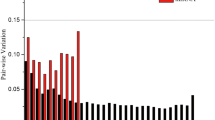
For the SRBSDV-infected rice plants. geNorm ranks the candidate reference genes based on the stability measure (M value) which is defined as the mean variation of a given gene in relation to all the others tested. The results of geNorm showed that 18S and EF1α had the most stable expression with an M value of 0.683 and ACTIN was the least stable with an M value of 1.503 (Table 2). The geNorm manual also determines the minimal number of reference genes required for normalization using a pairwise variation, it suggests that when the pairwise variation (Vn/n + 1) > 0.15, more than n + 1 genes should be used as internal reference genes. All Vn/n + 1 values were > 0.15 (Fig. 2A), suggesting that at least 3 genes should be used as control genes. The NormFinder results suggest that 18S is the best reference gene (Table 2), whereas the BestKeeper results suggest that 25S is the most suitable (Table 2). The overall RefFinder stability ranking of the candidate reference genes, from most to least stable, was: 18S > EF1α > UBQ10 > 25S > β-TUB > GAPDH > UBQ5 > ACTIN (Fig. 3A).
The stability of the 8 candidate reference genes was also analyzed in the leaves of healthy rice plants. EF1α and 18S were the most stable reference genes with a geNorm M value of 0.699 and UBQ5 was the least stable (Table 2). All Vn/n + 1 values were > 0.15, suggesting that more than 2 reference genes are required (Fig. 2B). The NormFinder results suggest that 18S is the most stable gene (Table 2) whereas BestKeeper found 25S to be the most suitable (Table 2). The RefFinder ranking, from most to least stable, was: 18S > UBQ10 > EF1α > 25S > β-TUB > ACTIN > GAPDH > UBQ5 (Fig. 3B).
Transcript levels of key enzymes genes involved in isoprenoid metabolic pathway
Based on the RefFinder ranking, 18S, EF1α and UBQ10 were selected to normalize different expression levels of the key genes involved in the isoprenoid metabolic pathway in SRBSDV-infected, and healthy, rice plants.
Compared with healthy rice plants, SQS, SPS and FPPS were significantly (t = 3.241, df = 3, p = 0.048; t = 3.207, df = 4, p = 0.049 and t = 27.059, df = 4, p < 0.01, respectively) up-regulated in the leaves of SRBSDV-infected rice plants at 0 day post inoculation, whereas IPI was significantly down-regulated (t = 6.017, df = 3, p < 0.01) (Fig. 4). At 15 days post inoculation, SPS, PHS and MTC were significantly up-regulated (t = 4.929, df = 4, p < 0.01; t = 2.918, df = 4, p = 0.043 and t = 3.191, df = 4, p = 0.033, respectively) and HDS was significantly down-regulated (t = 3.621, df = 4, p = 0.022) (Fig. 4). At 25 days post inoculation, SQS and MTC were significantly up-regulated (t = 4.042, df = 4, p = 0.016 and t = 6.614, df = 4, p < 0.01, respectively) and HDS and DMD were significantly down-regulated (t = 2.925, df = 4, p = 0.043 and t = 6.800, df = 4, p < 0.01, respectively) (Fig. 4). There were no significantly up-regulated genes 35 days after inoculation but SQS, FPPS, HDS, GGPPS, DMD and CMS were significantly down-regulated (t = 4.024, df = 4, p = 0.016; t = 3.355, df = 4, p = 0.028; t = 3.563, df = 4, p = 0.024; t = 3.818, df = 4, p = 0.019; t = 5.054, df = 4, p < 0.01 and t = 2.784, df = 4, p = 0.050, respectively) (Fig. 4). There were also no significantly up-regulated genes 45 days after inoculation but SQS, FPPS, IPI, HDS, DMD and CMS were significantly down-regulated (t = 13.110, df = 4, p <0.01; t = 6.711, df = 4, p < 0.01; t = 7.815, df = 4, p < 0.01; t = 6.727, df = 4, p < 0.01; t = 14.373, df = 4, p < 0.01 and t = 3.211, df = 4, p = 0.033, respectively) (Fig. 4).
Discussion
Reliable reference genes play a vital role in gene expression profiling. The fundamental assumption is that reference gene expression is consistent regardless of tissue type and experimental conditions. There is, however, no universal reference gene suitable for all species or experimental conditions [34]. Although many studies have assessed reference genes in different tissues and development stages of rice plants, and after subjecting plants to different kinds of stress [28, 35,36,37,38,39,40,41], to the best of our knowledge, no one has attempted to identify stable reference genes in the leaves of SRBSDV-infected rice plants.
We assessed the stability of 8 reference genes using four statistical models. According to the RefFinder results, 18S, UBQ10 and EF1α were the best reference genes for the leaves of SRBSDV-infected rice plant at different times after inoculation. The infection of SRBSDV can cause profound metabolic changes in rice plants and affecting the expression of different genes [16, 19, 42]. Therefore, our results represent the critical first step in investigating changes in gene expression in the leaves of SRBSDV-infected rice plants.
18S was ranked the most stable gene by geNorm, and the third most stable by NormFinder and BestKeeper in SRBSDV-infected rice plants. 18S has also been identified as a reliable reference gene during the growth stages of rice [43]. The genes encoding elongation factor 1-alpha and ubiquitin have also been considered reliable reference genes under different experimental conditions [44]. We also found UBQ10 and EF1α to be reliable reference genes in the leaves of SRBSDV-infected rice plants. EF1α has been identified as a stable reference gene in rice subject to low temperatures [36], and UBQ10 has been recommended as a reliable reference gene in rice plant leaves subject to salt stress [28]. GAPDH is commonly regarded as the “classical” reference gene [45], and has been suggested as a reliable reference gene in rice seedlings suffering from iron toxicity [35]. However, we found the expression of GAPDH to be quite variable. This variation in results demonstrates the importance of identifying suitable reference genes for specific experimental conditions.
The optimal number of reference genes required to normalize gene expression levels was analyzed using pairwise variation (Vn/n + 1) values in geNorm. All pairwise variation values were > 0.15 which suggests that a minimum of 3 reference genes are required [46]. Since rice seedlings grew from the tillering stage to the booting stage over the 45 day experimental period, the observed large changes in gene expression are not unexpected.
Most of the genes involved in the isoprenoid metabolic pathway were up-regulated 0 day after viruliferous S. furcifera nymphs were removed from rice plants (Fig. 4). However, nymphs had already and fed on plants for 3 days before they were removed and viruliferous S. furcifera spend more time salivating and ingesting phloem sap than virus-free S. furcifera [47]. Plant growth, nutrient composition, and secondary compounds can all be affected by viral infection [15, 48, 49]. Secondary compounds are important signals linking plants, pests and natural enemies [50]. Compared to healthy rice plants, the expression of SQS, PHS, FPPS, IPI, HDS, GGPPS, DMD and CMS was suppressed in SRBSDV-infected rice plants 35 and 45 days after virus inoculation (Fig. 4). Moreover, the mRNA transcript levels of MTC, FPPS and SQS, which catalyze the synthesis of monoterpenes and sesquiterpenes, had rapidly declined 35 and 45 days after virus inoculation (Fig. 4). SRBSDV titres are high in rice 35–50 days after inoculation [16]. Therefore, we suspect that SRBSDV suppressed the chemical defenses of rice against S. furcifera, thereby enhancing pathogen transmission. Predators of pest insects are also attracted by some terpenoids, such as monoterpenes and sesquiterpenes [51].
In addition, the expression of PHS in SRBSDV-infected rice plants was lower than in healthy rice plants. Since the down-regulation of PHS has been found to result in a dwarf phenotype of Oncidium hybrid [52] and the viral titre was high 25, 35 and 45 days after virus inoculation, we think that the dwarf phenotype of SRBSDV-infected rice could be related to the down-regulation of PHS.
References
Bustin SA, Nolan T (2004) Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J Biomol Tech 15(3):155–166
Yeung AT, Holloway BP, Adams PS, Shipley GL (2004) Evaluation of dual-labeled fluorescent DNA probe purity versus performance in real-time PCR. BioTechniques 36(2):266–270, 272, 274–265. https://doi.org/10.2144/04362RR01
Derveaux S, Vandesompele J, Hellemans J (2010) How to do successful gene expression analysis using real-time PCR. Methods 50(4):227–230. https://doi.org/10.1016/j.ymeth.2009.11.001
Mahoney DJ, Carey K, Fu MH, Snow R, Cameron-Smith D, Parise G, Tarnopolsky MA (2004) Real-time RT-PCR analysis of housekeeping genes in human skeletal muscle following acute exercise. Physiol Genom 18(2):226–231. https://doi.org/10.1152/physiolgenomics.00067.2004
Yamakawa H, Hirose T, Kuroda M, Yamaguchi T (2007) Comprehensive expression profiling of rice grain filling-related genes under high temperature using DNA microarray. Plant Physiol 144(1):258–277. https://doi.org/10.2307/40065337
Nicot N, Hausman JF, Hoffmann L, Evers D (2005) Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot 56(421):2907–2914. https://doi.org/10.1093/jxb/eri285
Goidin D, Mamessier A, Staquet MJ, Schmitt D, Berthier-Vergnes O (2001) Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and β-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Anal Biochem 295(1):17–21. https://doi.org/10.1006/abio.2001.5171
Fischer M, Skowron M, Berthold F (2005) Reliable transcript quantification by real-time reverse transcriptase-polymerase chain reaction in primary neuroblastoma using normalization to averaged expression levels of the control genes HPRT1 and SDHA. J Mol Diagn 7(1):89–96. https://doi.org/10.1677/jme.0.0290023
Daijun L, Salvaterra PM (2011) Robust RT-qPCR data normalization: validation and selection of internal reference genes during post-experimental data analysis. PLoS ONE 6(3):e17762. https://doi.org/10.1371/journal.pone.0017762
Xiao X, Ma J, Wang J, Wu X, Li P, Yao Y (2014) Validation of suitable reference genes for gene expression analysis in the halophyte Salicornia europaea by real-time quantitative PCR. Front Plant Sci 5:788. https://doi.org/10.3389/fpls.2014.00788
Zhou G, Wen J, Cai D, Li P, Xu D, Zhang S (2008) Southern rice black-streaked dwarf virus: a new proposed Fijivirus species in the family Reoviridae. Chin Sci Bull 53(23):3677–3685. https://doi.org/10.1007/s11434-008-0467-2
Zhou G, Xu D, Xu D, Zhang M (2013) Southern rice black-streaked dwarf virus: a white-backed planthopper-transmitted Fijivirus threatening rice production in Asia. Front Microbiol 4:270. https://doi.org/10.3389/fmicb.2013.00270
Lv M, Xie L, Yang J, Chen J, Zhang H-M (2016) Complete genomic sequence of maize rough dwarf virus, a Fijivirus transmitted by the small brown planthopper. Genome Announc 4(1):e01529-15. https://doi.org/10.1128/genomea.01529-15
Zhang HM, Yang J, Chen JP, Adams MJ (2008) A black-streaked dwarf disease on rice in China is caused by a novel Fijivirus. Arch Virol 153(10):1893–1898. https://doi.org/10.1007/s00705-008-0209-4
He X, Xu H, Gao G, Zhou X, Zheng X, Sun Y, Yang Y, Tian J, Lu Z (2014) Virus-mediated chemical changes in rice plants impact the relationship between non-vector planthopper Nilaparvata lugens Stål and its egg parasitoid Anagrus nilaparvatae Pang et Wang. PLoS ONE 9(8):e105373. https://doi.org/10.1371/journal.pone.0105373
Lu G, Zhang T, He Y, Zhou G (2016) Virus altered rice attractiveness to planthoppers is mediated by volatiles and related to virus titre and expression of defence and volatile-biosynthesis genes. Sci Rep 6:38581. https://doi.org/10.1038/srep38581
Xu HX, He XC, Zheng XS, Yang YJ, Zhang JF, Lu ZX (2016) Effects of SRBSDV-infected rice plants on the fitness of vector and non-vector rice planthoppers. J Asia-Pacif Entomol 19(3):707–710. https://doi.org/10.1016/j.aspen.2016.06.016
Wang Z, Yu L, Jin L, Wang W, Zhao Q, Ran L, Li X, Chen Z, Guo R, Wei Y, Yang Z, Liu E, Hu D, Song B (2017) Evaluation of rice resistance to southern rice black-streaked dwarf virus and rice ragged stunt virus through combined field tests, quantitative real-time PCR, and proteome analysis. Viruses 9(2):E37. https://doi.org/10.3390/v9020037
Li P, Liu H, Li F, Liao X, Ali S, Hou M (2018) A virus plays a role in partially suppressing plant defenses induced by the viruliferous vectors. Sci Rep 8(1):9027. https://doi.org/10.1038/s41598-018-27354-9
Whitham SA, Quan S, Chang HS, Cooper B, Zhu T, Wang X, Hou YM (2010) Diverse RNA viruses elicit the expression of common sets of genes in susceptible Arabidopsis thaliana plants. Plant J 33(2):271–283. https://doi.org/10.1046/j.1365-313X.2003.01625.x
Babu M, Griffiths JS, Huang TS, Wang A (2008) Altered gene expression changes in Arabidopsis leaf tissues and protoplasts in response to Plum pox virus infection. BMC Genom 9:325. https://doi.org/10.1186/1471-2164-9-325
Ascencioibáñez JT, Sozzani R, Lee TJ, Chu TM, Wolfinger RD, Cella R (2008) Global analysis of arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection. Plant Physiol 148(1):436–454. https://doi.org/10.2307/40066189
Martini X, Willett DS, Kuhns EH, Stelinski LL (2016) Disruption of vector host preference with plant volatiles may reduce spread of insect-transmitted plant pathogens. J Chem Ecol 42(5):357–367. https://doi.org/10.1007/s10886-016-0695-x
Wang H, Xu D, Pu L, Zhou G (2014) Southern rice black-streaked dwarf virus alters insect vectors’ host orientation preferences to enhance spread and increase rice ragged stunt virus co-infection. Phytopathology 104(2):196–201. https://doi.org/10.1094/PHYTO-08-13-0227-R
Wang L, Hu K, He H, Ding W, Li Y (2017) Southern rice black-streaked dwarf virus-induced volatiles from rice plants and behavioral responses of adult Sogatella furcifera (Hemiptera: Delphacidae) to the components of these volatiles. Acta Entomol Sin 60(4):412–420
Li QF, Sun SSM, Yuan DY, Yu HX, Gu MH, Liu QQ (2010) Validation of candidate reference genes for the accurate normalization of real-time quantitative RT-PCR data in rice during seed development. Plant Mol Biol Rep 28(1):49–57. https://doi.org/10.1007/s11105-009-0124-1
Bevitori R, Oliveira MB, Grossi-de-Sá MF, Lanna AC, da Silveira RD, Petrofeza S (2014) Selection of optimized candidate reference genes for qRT-PCR normalization in rice (Oryza sativa L.) during Magnaporthe oryzae infection and drought. Genet Mol Res 13(4):9795–9805. https://doi.org/10.4238/2014.November.27.7
Moraes GP, Benitez LC, do Amaral MN, Vighi IL, Auler PA, da Maia LC, Bianchi VJ, Braga EJ (2015) Evaluation of reference genes for RT-qPCR studies in the leaves of rice seedlings under salt stress. Genet Mol Res 14(1):2384–2398. https://doi.org/10.4238/2015.March.27.24
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):RESEARCH0034
Andersen CL, Jensen JL, Ørntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64(15):5245–5250. https://doi.org/10.1158/0008-5472.CAN-04-0496
Mallona I, Lischewski S, Weiss J, Hause B, Egea-Cortines M (2010) Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol 10:4. https://doi.org/10.1186/1471-2229-10-4
Wang X, Xiong M, Wang J, Lei C, Zhu F (2015) Reference gene stability of a synanthropic fly, Chrysomya megacephala. Parasites Vectors 8:565. https://doi.org/10.1186/s13071-015-1175-9
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45. https://doi.org/10.1093/nar/29.9.e45
Bustin S (2002) Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J Mol Endocrinol 29(1):23–39. https://doi.org/10.1677/jme.0.0290023
Santos FICD, Marini N, Santos RSD, Hoffman BSF, Alves-Ferreira M, de Oliveira AC (2018) Selection and testing of reference genes for accurate RT-qPCR in rice seedlings under iron toxicity. PLoS ONE 13(3):e0193418. https://doi.org/10.1371/journal.pone.0193418
Auler PA, Benitez LC, do Amaral MN, Vighi IL, Rodrigues GS, da Maia LC, Braga EJB (2017) Selection of candidate reference genes and validation for real-time PCR studies in rice plants exposed to low temperatures. Genet Mol Res 16(2):gmr16029695. https://doi.org/10.4238/gmr16029695
Auler PA, Benitez LC, do Amaral MN, Vighi IL, Dos Santos Rodrigues G, da Maia LC, Braga EJ (2017) Evaluation of stability and validation of reference genes for RT-qPCR expression studies in rice plants under water deficit. J Appl Genet 58(2):163–177. https://doi.org/10.1007/s13353-016-0374-1
Wang Z, Wang Y, Yang J, Hu K, An B, Deng X, Li Y (2016) Reliable selection and holistic stability evaluation of reference genes for rice under 22 different experimental conditions. Appl Biochem Biotechnol 179(5):753–775. https://doi.org/10.1007/s12010-016-2029-4
Xu H, Bao JD, Dai JS, Li Y, Zhu Y (2015) Genome-wide identification of new reference genes for qRT-PCR normalization under high temperature stress in rice endosperm. PLoS ONE 10(11):e0142015. https://doi.org/10.1371/journal.pone.0142015
Tan QQ, Zhu L, Li Y, Liu W, Ma WH, Lei CL, Wang XP (2015) A de novo transcriptome and valid reference genes for quantitative real-time PCR in Colaphellus bowringi. PLoS ONE 10(2):e0118693. https://doi.org/10.1371/journal.pone.0118693
Narsai R, Ivanova A, Ng S, Whelan J (2010) Defining reference genes in Oryza sativa using organ, development, biotic and abiotic transcriptome datasets. BMC Plant Biol 10:56. https://doi.org/10.1186/1471-2229-10-56
Xu D, Mou G, Wang K, Zhou G (2014) MicroRNAs responding to southern rice black-streaked dwarf virus infection and their target genes associated with symptom development in rice. Virus Res 190:60–68. https://doi.org/10.1016/j.virusres.2014.07.007
Kim BR, Nam HY, Kim SU, Kim SI, Chang YJ (2003) Normalization of reverse transcription quantitative-PCR with housekeeping genes in rice. Biotechnol Lett 25(21):1869–1872. https://doi.org/10.1023/a:1026298032009
Han X, Lu M, Chen Y, Zhan Z, Cui Q, Wang Y (2012) Selection of reliable reference genes for gene expression studies using real-time PCR in tung tree during seed development. PLoS ONE 7(8):e43084. https://doi.org/10.1371/journal.pone.0043084
de Jonge HJ, Fehrmann RS, de Bont ES, Hofstra RM, Gerbens F, Kamps WA, de Vries EG, van der Zee AG, te Meerman GJ, ter Elst A (2007) Evidence based selection of housekeeping genes. PLoS ONE 2(9):e898. https://doi.org/10.1371/journal.pone.0000898
Koramutla MK, Aminedi R, Bhattacharya R (2016) Comprehensive evaluation of candidate reference genes for qRT-PCR studies of gene expression in mustard aphid, Lipaphis erysimi (Kalt). Sci Rep 6:25883. https://doi.org/10.1038/srep25883
Lei W, Li P, Han Y, Gong S, Yang L, Hou M (2016) EPG recordings reveal differential feeding behaviors in Sogatella furcifera in response to plant virus infection and transmission success. Sci Rep 6:30240. https://doi.org/10.1038/srep30240
Xu HX, He XC, Zheng XS, Yang YJ, Lu ZX (2014) Influence of rice black streaked dwarf virus on the ecological fitness of non-vector planthopper Nilaparvata lugens (Hemiptera: Delphacidae). Insect Sci 21(4):507–514. https://doi.org/10.1111/1744-7917.12045
Fiebig M, Poehling HM, Borgemeister C (2004) Barley yellow dwarf virus, wheat, and Sitobion avenae: a case of trilateral interactions. Entomol Exp Appl 110(1):11–21. https://doi.org/10.1111/j.0013-8703.2004.00115.x
Obara N, Hasegawa MO (2002) Induced volatiles in elicitor-treated and rice blast fungus-inoculated rice leaves. Biosci Biotechnol Biochem 66(12):2549–2559. https://doi.org/10.1271/bbb.66.2549
Khan ZR, James DG, Midega CAO, Pickett JA (2008) Chemical ecology and conservation biological control. Biol Control 45(2):210–224. https://doi.org/10.1016/j.biocontrol.2007.11.009
Liu JX, Chiou CY, Shen CH, Chen PJ, Liu YC, Jian CD, Shen XL, Shen FQ, Yeh KW (2014) RNA interference-based gene silencing of phytoene synthase impairs growth, carotenoids, and plastid phenotype in Oncidium hybrid orchid. Springerplus 3:478. https://doi.org/10.1186/2193-1801-3-478
Funding
This work was funded by the National Natural Science Foundation of China (31572005) and Research Foundation of Education Bureau of Hunan Province, China (15A090).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hu, K., Qiu, L., Ding, W. et al. Evaluation of reference genes and expression of key genes involved in the isoprenoid metabolic pathway of rice leaves after infection by the Southern rice black-streaked dwarf virus. Mol Biol Rep 46, 3945–3953 (2019). https://doi.org/10.1007/s11033-019-04841-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04841-4