Abstract
Novel pyrrolo[2,3-d]pyrimidine-based analogues were designed, synthesized, and evaluated for their ability to inhibit the α-amylase enzyme in order to treat diabetes. In vitro antidiabetic analysis demonstrated excellent antidiabetic action for compounds 5b, 6c, 7a, and 7b, with IC50 values in the 0.252–0.281 mM range. At a 200 μg/mL concentration, the exceptional percent inhibition values for compounds 5a, 5b, 5d, and 6a varied from 97.79 ± 2.86% to 85.56 ± 4.13% overperforming the standard (acarbose). Molecular docking of all compounds performed with Bacillus paralicheniformis α-amylase enzyme. The most active compounds via in vitro and non-toxic via in silico ADMET and molecular docking analysis, hybrids 6c, 7a, and 7b displayed binding affinity from − 8.2 and − 8.5 kcal/mol. Molecular dynamic simulations of most active compound 5b and 7a investigated into the active sites of the Bacillus paralicheniformis α-amylase enzyme for a 100-ns indicating the stability of hybrid-protein complex. Consistent RGyr values for the two complexes under study further suggest that the system's proteins are closely packed in the dynamic state. Synthesized analogs’ in vitro biological assessments, ADMET, molecular docking, and MD modelling reveal that 5b, 6c, 7a, and 7b hybrid analogs may be employed in the development of future antidiabetic drugs.
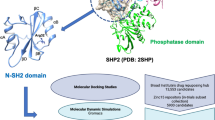
Graphical abstract








Similar content being viewed by others
References
Deepa M, Bhansali A, Anjana R et al (2014) Knowledge and awareness of diabetes in urban and rural India: The Indian Council of Medical Research India Diabetes Study (Phase I): Indian Council of Medical Research India Diabetes 4. Indian J Endocrinol Metab 18:379. https://doi.org/10.4103/2230-8210.131191
Zala AR, Naik HN, Ahmad I et al (2023) Design and synthesis of novel 1,2,3-triazole linked hybrids: molecular docking, MD simulation, and their antidiabetic efficacy as α-amylase inhibitors. J Mol Struct 1285:135493. https://doi.org/10.1016/j.molstruc.2023.135493
Ligthelm RJ, Kaiser M, Vora J, Yale JF (2012) Insulin use in elderly adults: risk of hypoglycemia and strategies for care. J Am Geriatr Soc 60:1564–1570. https://doi.org/10.1111/J.1532-5415.2012.04055.X
Lazar C, Kluczyk A, Kiyota T, Konishi Y (2004) Drug evolution concept in drug design: 1. Hybridization method. J Med Chem 47(27):6973–6982. https://doi.org/10.1021/jm049637
Viegas-Junior C, Danuello A, da Bolzani V et al (2007) Molecular hybridization: a useful tool in the design of new drug prototypes. Curr Med Chem 14:1829–1852. https://doi.org/10.2174/092986707781058805
Zala AR, Rajani DP, Kumari P (2023) Design, synthesis, molecular docking and in silico ADMET investigations of novel piperidine-bearing cinnamic acid hybrids as potent antimicrobial agents. J Iran Chem Soc. https://doi.org/10.1007/s13738-023-02801-1
Szumilak M, Wiktorowska-Owczarek A, Stanczak A (2021) Hybrid drugs—a strategy for overcoming anticancer drug resistance? Molecules 26:2601. https://doi.org/10.3390/molecules26092601
Kerru N, Singh P, Koorbanally N et al (2017) Recent advances (2015–2016) in anticancer hybrids. Eur J Med Chem 142:179–212. https://doi.org/10.1016/j.ejmech.2017.07.033
Abbot V, Sharma P, Dhiman S et al (2017) Small hybrid heteroaromatics: resourceful biological tools in cancer research. RSC Adv 7:28313–28349. https://doi.org/10.1039/C6RA24662A
Nepali K, Sharma S, Sharma M et al (2014) Rational approaches, design strategies, structure activity relationship and mechanistic insights for anticancer hybrids. Eur J Med Chem 77:422–487. https://doi.org/10.1016/j.ejmech.2014.03.018
Chen LZ, Shu HY, Wu J et al (2021) Discovery and development of novel pyrimidine and pyrazolo/thieno-fused pyrimidine derivatives as potent and orally active inducible nitric oxide synthase dimerization inhibitor with efficacy for arthritis. Eur J Med Chem. https://doi.org/10.1016/J.EJMECH.2021.113174
Gomha SM, Hassaneen HME (2011) Synthesis and antimicrobial activity of some new pyrazoles, fused pyrazolo[3,4-d]-pyrimidine and 1,2-dihydroimidazo-[2,1-c][1,2,4]triazin-6-one derivatives. Molecules 16:6549–6560. https://doi.org/10.3390/MOLECULES16086549
Trivedi AR, Dholariya BH, Vakhariya CP et al (2012) Synthesis and anti-tubercular evaluation of some novel pyrazolo[3,4-d] pyrimidine derivatives. Med Chem Res 21:1887–1891. https://doi.org/10.1007/S00044-011-9712-3/TABLES/1
Rashad AE, Hegab MI, Abdel-Megeid RE et al (2008) Synthesis and antiviral evaluation of some new pyrazole and fused pyrazolopyrimidine derivatives. Bioorg Med Chem 16:7102–7106. https://doi.org/10.1016/J.BMC.2008.06.054
Bakhotmah DA, Ali TE, Assiri MA, Yahia IS (2020) Synthesis of some novel 2-{pyrano[2,3-c]pyrazoles-4-ylidene}malononitrile fused with pyrazole, pyridine, pyrimidine, diazepine, chromone, pyrano[2,3-c]pyrazole and pyrano[2,3-d]pyrimidine systems as anticancer agents. Polycycl Aromat Compd 42:2136–2150. https://doi.org/10.1080/10406638.2020.1827445
Al-Ghorbani M, Gouda MA, Baashen M et al (2022) Piperazine heterocycles as potential anticancer agents: a review. Pharm Chem J 56:29–37. https://doi.org/10.1007/S11094-022-02597-Z
Patel RV, Mistry B, Syed R et al (2016) Chrysin-piperazine conjugates as antioxidant and anticancer agents. Eur J Pharm Sci 88:166–177. https://doi.org/10.1016/J.EJPS.2016.02.01
Sullivan DW, Gad SC, Laulicht B et al (2015) Nonclinical safety assessment of PER977: a small molecule reversal agent for new oral anticoagulants and heparins. Int J Toxicol 34(4):308–317. https://doi.org/10.1177/1091581815590667
Zala AR, Rajani DP, Kumari P (2022) Design, synthesis, molecular docking and antimicrobial and antimycobacterial activities of novel hybrid of coumarin-cinnamic acids. Chem Data Collect. https://doi.org/10.1016/j.cdc.2022.100862
Wang SF, Yin Y, Wu X et al (2014) Synthesis, molecular docking and biological evaluation of coumarin derivatives containing piperazine skeleton as potential antibacterial agents. Bioorg Med Chem 22:5727–5737. https://doi.org/10.1016/J.BMC.2014.09.048
Bassetto M, Leyssen P, Neyts J et al (2017) In silico identification, design and synthesis of novel piperazine-based antiviral agents targeting the hepatitis C virus helicase. Eur J Med Chem 125:1115–1131. https://doi.org/10.1016/J.EJMECH.2016.10.043
Thamban Chandrika N, Shrestha SK, Ngo HX et al (2018) Alkylated piperazines and piperazine-azole hybrids as antifungal agents. J Med Chem 61:158–173. https://doi.org/10.1021/ACS.JMEDCHEM.7B01138
Batista DC, Silva DPB, Florentino IF et al (2018) Anti-inflammatory effect of a new piperazine derivative: (4-methylpiperazin-1-yl)(1-phenyl-1H-pyrazol-4-yl)methanone. Inflammopharmacology 26:217–226. https://doi.org/10.1007/S10787-017-0390-8
El-Faham A, Armand-Ugón M, Esté JA, Albericio F (2008) Use of N-methylpiperazine for the preparation of piperazine-based unsymmetrical bis-ureas as anti-HIV agents. ChemMedChem 3:1034–1037. https://doi.org/10.1002/CMDC.200800059
Devine R, Kelada M, Leonard S et al (2020) Design, synthesis, and biological evaluation of aryl piperazines with potential as antidiabetic agents via the stimulation of glucose uptake and inhibition of NADH:ubiquinone oxidoreductase. Eur J Med Chem 202:112416. https://doi.org/10.1016/J.EJMECH.2020.112416
Mendoza A, Pérez-Silanes S, Quiliano M et al (2011) Aryl piperazine and pyrrolidine as antimalarial agents. Synthesis and investigation of structure–activity relationships. Exp Parasitol 128:97–103. https://doi.org/10.1016/J.EXPPARA.2011.02.025
Prashanth MK, Revanasiddappa HD, Lokanatha Rai KM, Veeresh B (2012) Synthesis, characterization, antidepressant and antioxidant activity of novel piperamides bearing piperidine and piperazine analogues. Bioorg Med Chem Lett 22:7065–7070. https://doi.org/10.1016/J.BMCL.2012.09.089
Shaquiquzzaman M, Verma G, Marella A et al (2015) Piperazine scaffold: a remarkable tool in generation of diverse pharmacological agents. Eur J Med Chem 102:487–529. https://doi.org/10.1016/J.EJMECH.2015.07.026
Pal R, Jawaid Akhtar M, Raj K et al (2022) Design, synthesis and evaluation of piperazine clubbed 1,2,4-triazine derivatives as potent anticonvulsant agents. J Mol Struct 1257:132587. https://doi.org/10.1016/J.MOLSTRUC.2022.132587
Zala AR, Rajani DP, Kumari P (2022) Synthesis, molecular docking, ADME study, and antimicrobial potency of piperazine based cinnamic acid bearing coumarin moieties as a DNA gyrase inhibitor. J Biochem Mol Toxicol. https://doi.org/10.1002/jbt.23231
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428. https://doi.org/10.1021/AC60147A030
Frisch A (1996) Gaussian 09W reference
Morris GM, Huey R, Olson AJ (2008) Using autodock for ligand–receptor docking. Curr Protoc Bioinformatics 24:8.14.1-8.14.40. https://doi.org/10.1002/0471250953.BI0814S24
Zala AR, Rajani DP, Ahmad I et al (2023) Synthesis, characterization, molecular dynamic simulation, and biological assessment of cinnamates linked to imidazole/benzimidazole as a CYP51 inhibitor. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2170918
Tian W, Chen C, Lei X et al (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res 46:W363–W367. https://doi.org/10.1093/NAR/GKY473
Pandey R, Dubey I, Ahmad I et al (2022) In silico study of some dexamethasone analogs and derivatives against SARs-CoV-2 target: a cost-effective alternative to remdesivir for various COVID phases. Current Chin Sci 2:294–309. https://doi.org/10.2174/2210298102666220404102217
Puri S, Ahmad I, Patel H et al (2023) Evaluation of oxindole derivatives as a potential anticancer agent against breast carcinoma cells: in vitro, in silico, and molecular docking study. Toxicol Vitro 86:105517. https://doi.org/10.1016/J.TIV.2022.105517
Benjamin I, Louis H, Ekpen FO et al (2022) Modeling the anti-methicillin-resistant Staphylococcus aureus (MRSA) Activity of (E)-6-chloro-N2-phenyl-N4-(4-phenyl-5-(phenyl diazinyl)-2λ3, 3 λ2-thiazol-2-yl)-1, 3, 5-triazine-2,4-diamine. Polycycl Aromat Compd 1:28. https://doi.org/10.1080/10406638.2022.2160773
Patel KB, Mukherjee S, Bhatt H et al (2023) Synthesis, docking, and biological investigations of new coumarin-piperazine hybrids as potential antibacterial and anticancer agents. J Mol Struct 1276:134755. https://doi.org/10.1016/J.MOLSTRUC.2022.134755
Mathew B, Ravichandran V, Raghuraman S et al (2022) Two dimensional-QSAR and molecular dynamics studies of a selected class of aldoxime- and hydroxy-functionalized chalcones as monoamine oxidase-B inhibitors. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2146198
Wang T, Liu X, Hao M et al (2016) Design, synthesis and evaluation of pyrrolo[2,3-d]pyrimidine-phenylamide hybrids as potent Janus kinase 2 inhibitors. Bioorg Med Chem Lett 26:2936–2941. https://doi.org/10.1016/J.BMCL.2016.04.027
Kim JT, Hamilton AD, Bailey CM et al (2006) FEP-guided selection of bicyclic heterocycles in lead optimization for non-nucleoside inhibitors of HIV-1 reverse transcriptase. J Am Chem Soc 128:15372–15373. https://doi.org/10.1021/JA066472G
Paul RK, Ahmad I, Patel H et al (2023) Phytochemicals from Amberboa ramosa as potential DPP-IV inhibitors for the management of type-II diabetes mellitus: Inferences from in-silico investigations. J Mol Struct 1271:134045. https://doi.org/10.1016/J.MOLSTRUC.2022.134045
Halder SK, Ahmad I, Shathi JF et al (2022) A comprehensive study to unleash the putative inhibitors of serotype2 of dengue virus: insights from an in silico structure-based drug discovery. Mol Biotechnol. https://doi.org/10.1007/S12033-022-00582-1
Ayipo YO, Alananzeh WA, Ahmad I et al (2022) Structural modelling and in silico pharmacology of β-carboline alkaloids as potent 5-HT1A receptor antagonists and reuptake inhibitors. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2104376
Haque MA, Hossain MS, Ahmad I et al (2022) Unveiling chlorpyrifos mineralizing and tomato plant-growth activities of Enterobacter sp. strain HSTU-ASh6 using biochemical tests, field experiments, genomics, and in silico analyses. Front Microbiol. https://doi.org/10.3389/FMICB.2022.1060554/FULL
Ayipo YO, Yahaya SN, Babamale HF et al (2021) ß-Carboline alkaloids induce structural plasticity and inhibition of SARS-CoV-2 nsp3 macrodomain more potently than remdesivir metabolite GS-441524: computational approach. Turk J Biol 45:503–517. https://doi.org/10.3906/biy-2106-64
Ayipo YO, Ahmad I, Alananzeh W et al (2022) Computational modelling of potential Zn-sensitive non-β-lactam inhibitors of imipenemase-1 (IMP-1). J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2153168
Acknowledgements
The authors thank CUG, Gujarat, India, for providing spectral data. The authors are thankful to the Department of Chemistry of S.V. National Institute Technology, Surat, Gujarat, India, for providing all the facilities for the research work.
Author information
Authors and Affiliations
Contributions
ARZ: Conceptualization, methodology, writing original draft, and visualization. RGT: methodology and visualization. HNN: biological evaluation. AI: molecular dynamic simulation. HP, SJ, PK: supervision, writing review and editing. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zala, A.R., Tiwari, R., Naik, H.N. et al. Design and synthesis of pyrrolo[2,3-d]pyrimidine linked hybrids as α-amylase inhibitors: molecular docking, MD simulation, ADMET and antidiabetic screening. Mol Divers (2023). https://doi.org/10.1007/s11030-023-10683-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-023-10683-x