Abstract
Post-translational modifications of proteins such as protein ubiquitination are crucial for regulating conformation, stability and localization of the modified protein. Ubiquitin-specific protease 2 (USP2), a multifunctional cysteine protease is reported to be a key regulator of ubiquitylation events in numerous oncogenic proteins e.g., fatty acid synthetase, Mdm2, EGFR, cyclin A1, and cyclin-D1, etc. Thus targeting USP2 is a promising strategy for cancer therapy. USP2 is characterized by a catalytic triad comprising of cysteine, histidine and aspartic acid residues. Five residues including three from the catalytic triad and two from outside of the catalytic triad have been reported as a catalytic site of USP2 that catalyze hydrolysis and stabilizes the oxyanion formed in the intermediate step of catalysis. Here, we report two more novel residues (L269 and Y558) on USP2 involved in the catalysis of Ubiquitin using computational alanine scanning (CAS) followed by molecular dynamic simulation studies. The results obtained from CAS were further validated by a highly reliable, time- and cost-effective SDS-PAGE-based kinetics assay using UBA52 which is a natural substrate of USP2. Our results showed that mutating L269 and Y558 significantly compromised the catalytic efficiency of USP2 in hydrolyzing UBA52 which can further be extended to rational drug design of USP2 selective inhibitors and to explore the catalytic sites of other USPs.
Graphical abstract
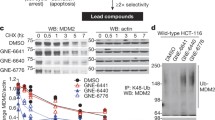
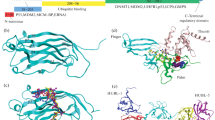
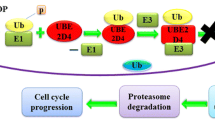
Two novel residues take part in catalytic activity of USP2 which were depicted by MD Simulations and were further validated by novel SDS-PAGE-based reliable time- and cost-effective kinetics assay.






Similar content being viewed by others
References
Hershko A, Ciechanover A, Varshavsky A (2000) The ubiquitin system. Nat Med 6(10):1073–1081. https://doi.org/10.1038/80384
Hoeller D, Dikic I (2009) Targeting the ubiquitin system in cancer therapy. Nature 458(7237):438–444. https://doi.org/10.1038/nature07960
Kerscher O, Felberbaum R, Hochstrasser M (2006) Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol 22:159–180. https://doi.org/10.1146/annurev.cellbio.22.010605.093503
Harrigan JA, Jacq X, Martin NM, Jackson SP (2018) Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat Rev Drug Discov 17(1):57–78. https://doi.org/10.1038/nrd.2017.152
Myung J, Kim KB, Crews CM (2001) The ubiquitin-proteasome pathway and proteasome inhibitors. Med Res Rev 21(4):245–273. https://doi.org/10.1002/med.1009
Peng J, Schwartz D, Elias JE, Thoreen CC, Cheng D, Marsischky G, Gygi SP (2003) A proteomics approach to understanding protein ubiquitination. Nat biotech 21(8):921–926. https://doi.org/10.1038/nbt849
Meierhofer D, Wang X, Huang L, Kaiser P (2008) Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J proteome Res 7(10):4566–4576. https://doi.org/10.1021/pr800468j
Komander D, Clague MJ, Urbé S (2009) Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol 10(8):550–563. https://doi.org/10.1038/nrm2731
Edelmann MJ, Kessler BM (2008) Ubiquitin and ubiquitin-like specific proteases targeted by infectious pathogens: Emerging patterns and molecular principles. Biochim Biophys Acta-Mol Basis Dis 1782(12):809–816. https://doi.org/10.1016/j.bbadis.2008.08.010
Zhang W, Sulea T, Tao L, Cui Q, Purisima EO, Vongsamphanh R, Li Y (2011) Contribution of active site residues to substrate hydrolysis by USP2: insights into catalysis by ubiquitin specific proteases. Biochemistry 50(21):4775–4785. https://doi.org/10.1021/bi101958h
Renatus M, Parrado SG, D’Arcy A, Eidhoff U, Gerhartz B, Hassiepen U, Worpenberg S (2006) Structural basis of ubiquitin recognition by the deubiquitinating protease USP2. Structure 14(8):1293–1302. https://doi.org/10.1016/j.str.2006.06.012
Tencer AH, Liang Q, Zhuang Z (2016) Divergence in ubiquitin interaction and catalysis among the ubiquitin-specific protease family deubiquitinating enzymes. Biochemistry 55(33):4708–4719. https://doi.org/10.1021/acs.biochem.6b00033
Stevenson LF, Sparks A, Allende NV, Xirodimas DP, Lane DP, Saville MK (2007) The deubiquitinating enzyme USP2a regulates the p53 pathway by targeting Mdm2. EMBO J 26(4):976–986. https://doi.org/10.1038/sj.emboj.7601567
Graner E, Tang D, Rossi S, Baron A, Migita T, Weinstein LJ, Signoretti S (2004) The isopeptidase USP2a regulates the stability of fatty acid synthase in prostate cancer. Cancer Cell 5(3):253–261. https://doi.org/10.1016/S1535-6108(04)00055-8
Buckley D, Duke G, Heuer TS, O’Farrell M, Wagman AS, McCulloch W, Kemble G (2017) Fatty acid synthase–modern tumor cell biology insights into a classical oncology target. Pharmacol Ther 177:23–31. https://doi.org/10.1016/j.pharmthera.2017.02.021
Shan J, Zhao W, Gu W (2009) Suppression of cancer cell growth by promoting cyclin D1 degradation. Mol Cell 36(3):469–476. https://doi.org/10.1016/j.molcel.2009.10.018
Davis MI, Pragani R, Fox JT, Shen M, Parmar K, Gaudiano EF, Hall MD (2016) Small molecule inhibition of the ubiquitin-specific protease USP2 accelerates cyclin D1 degradation and leads to cell cycle arrest in colorectal cancer and mantle cell lymphoma models. J Biolog Chem 291(47):24628–24640. https://doi.org/10.1074/jbc.M116.738567
Pushpakom S, Iorio F, Eyers PA, Escott KJ, Hopper S, Wells A, Doig A, Guilliams T, Latimer J, McNamee C, Norris A (2019) Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov 18:41–58. https://doi.org/10.1038/nrd.2018.168
Chuang SJ, Cheng SC, Tang HC, Sun CY, Chou CY (2018) 6-Thioguanine is a noncompetitive and slow binding inhibitor of human deubiquitinating protease USP2. Sci Rep 8(1):3102. https://doi.org/10.1038/s41598-018-21476-w
Wang ZL, Xie W, Zhu M, Zhou H (2017) Development of a highly reliable assay for ubiquitin-specific protease 2 inhibitors. Bioorg Med Chem Lett 27(17):4015–4018. https://doi.org/10.1016/j.bmcl.2017.07.059
Rehman AU, Rafiq H, Rahman MU, Li J, Liu H, Luo S, Chen HF (2019) Gain-of-function SHP2 E76Q mutant rescuing autoinhibition mechanism associated with juvenile myelomonocytic leukemia. J Chem Inf Model 59(7):3229–3239. https://doi.org/10.1021/acs.jcim.9b00353
Riaz M, Rehman AU, Shah SA, Rafiq H, Lu S, Qiu Y, Wadood A (2021) Predicting Multi-Interfacial Binding Mechanisms of NLRP3 and ASC Pyrin Domains in Inflammasome Activation. ACS Chem Neurosci 12(4):603–612. https://doi.org/10.1021/acschemneuro.0c00519
Salomon RF, Case DA, Walker RC (2013) An overview of the Amber biomolecular simulation package. Wiley Interdiscip Rev Comput Mol Sci 3(2):198–210. https://doi.org/10.1021/acs.jcim.0c00613
Zwanzig R (1973) Nonlinear generalized Langevin equations. J Stat Phys 9(3):215–220. https://doi.org/10.1007/BF01008729
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: An N⋅ log (N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092. https://doi.org/10.1063/1.464397
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103(19):8577–8593. https://doi.org/10.1063/1.470117
Ryckaert JP, Ciccotti G, Berendsen HJ (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Salomon RF, Case DA, Walker RC (2013) Routine microsecond molecular dynamics simulations with AMBER on GPUs: Explicit solvent particle mesh Ewald. J Chem Theory Comput 9(9):3878–3888. https://doi.org/10.1021/ct400314y
Gotz AW, Williamson MJ, Xu D, Poole D, Le Grand S, Walker RC (2012) Routine microsecond molecular dynamics simulations with AMBER on GPUs 1. Generalized born. J Chem Theory Comput 8(5):1542–1555. https://doi.org/10.1021/ct200909j
Wang Z, Liu Y, Zhang J, Ullah S, Kang N, Zhao Y, Zhou H (2020) Benzothiophene-2-carboxamide derivatives as SENPs inhibitors with selectivity within SENPs family. Eur J Med Chem 204:112553. https://doi.org/10.1016/j.ejmech.2020.112553
Reverter D, Lima CD (2006) Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates. Nat Struct Mol Biol 13(12):1060–1068. https://doi.org/10.1038/nsmb1168
Junaid M, Khan MT, Malik SI, Wei DQ (2018) Insights into the mechanisms of the pyrazinamide resistance of three pyrazinamidase mutants N11K, P69T, and D126N. J Chem Inf Model 59(1):498–508. https://doi.org/10.1021/acs.jcim.8b00525
Khan A, Zia T, Suleman M, Khan T, Ali SS, Abbasi AA, Wei DQ (2021) Higher infectivity of the SARS-CoV-2 new variants is associated with K417N/T, E484K, and N501Y mutants: an insight from structural data. J Cell Physiol 236(10):7045–7057. https://doi.org/10.1002/jcp.30367
Acknowledgements
The principal author is grateful to China Scholarship Council (CSC) for funding this project, School of Pharmacy, SJTU, China and CADD lab, Abdul Wali Khan University, Pakistan for providing all relevant facilities. We are grateful to Professor Huchen Zhou for providing all the technical support in project design and facilities to conduct kinetics studies at her lab.
Funding
The lead author Shafi Ullah acknowledges China Scholarship Council (CSC) for funding this project.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest with anyone.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ullah, S., Junaid, M., Liu, Y. et al. Validation of catalytic site residues of Ubiquitin Specific Protease 2 (USP2) by molecular dynamic simulation and novel kinetics assay for rational drug design. Mol Divers 27, 1323–1332 (2023). https://doi.org/10.1007/s11030-022-10499-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10499-1