Abstract
Skeletal muscle satellite cells (SMSCs), also known as a multipotential stem cell population, play a crucial role during muscle growth and regeneration. In recent years, numerous miRNAs have been associated with the proliferation and differentiation of SMSCs in a number of mammalian species; however, the regulatory mechanisms of miR-194-5p in rabbit SMSCs still remain scarce. In this study, miR-194-5p was first observed to be highly expressed in the rabbit leg muscle. Furthermore, both the mimics and inhibitor of miR-194-5p were used to explore its role in the proliferation and differentiation of rabbit SMSCs cultured in vitro. Results from both EdU and CCK8 assays showed that miR-194-5p inhibited the proliferation of SMSCs. Meanwhile, Mef2c was identified as a target gene of miR-194-5p based on the dual-luciferase reporter assay results. In addition, upregulation of miR-194-5p decreased the expression levels of Mef2c and MyoG during rabbit SMSCs differentiation on Days 3 and 7 of in vitro culture. Taken together, these data demonstrated that miR-194-5p negatively regulates the proliferation and differentiation of rabbit SMSCs by targeting Mef2c.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Skeletal muscle satellite cells (SMSCs), also known as a multipotential stem cell population, mainly exist on the surface of myofiber but beneath the basement membrane. Usually, adult SMSCs are quiescent, but they are prone to be activated by physical trauma or growth signals [1]. It has been widely accepted that SMSCs function as the myogenic precursors and give rise to myoblasts which eventually differentiate into multinucleated myotubes [2]. The myogenic basic-helix-loop-helix (bHLH) family of transcription factors, including MyoD, MyoG, Myf5, and Mrf4, is known as key regulators of myogenic differentiation of myoblasts. Silencing of MyoG resulted in a reduction of skeletal muscle tissue of mice by impeding the differentiation of myoblasts [3]. In addition, the members of the myocyte enhancer factor 2(Mef2) gene family, which recognize a conserved A/T-rich element of muscle-specific genes, are highly expressed in myoblasts. Previous studies have suggested that Mef2c and MyoD can regulate the differentiation of myoblast by co-activating their downstream target genes [4, 5].
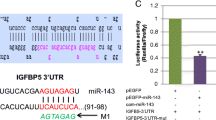
MiRNAs, a class of non-coding RNAs, serve as posttranscriptional regulators of gene functions through base pairing with the seed sequence of 3′UTR region of genes, thereby interfering genes expression [6]. Over the last few decades, many studies have proved that miRNAs play important roles in a variety of cellular activities, including cell proliferation, differentiation, and apoptosis [7, 8]. Nowadays, lots of studies have indicated that miRNAs also play a role in myogenesis. For instance, miR-1 and miR-206 were upregulated during the differentiation of SMSCs, and upregulation of their expression promoted both the proliferation and differentiation of SMSCs [9]. miR-34a was shown to positively regulate smooth muscle cell differentiation by increasing the expression of Mef2c, SRF and SirT1 [10]. And levels of miR-133a and miR-133b increased during the differentiation of human myoblasts [11]. Recently, increasing studies have focused on the regulation of satellite cells in domestic animals because of the major contribution of muscle to meat production. For example, the levels of Bta-miR-378 increased significantly during the differentiation of bovine skeletal muscle-derived satellite cells (bMDSC) and the over-expression of bta-miR-378 enhanced bMDSC differentiation [12]. Zhang et al. [13] found miR-143 promoted the differentiation of bovine SMSCs by targeting IGFBP5. miR-27b was identified as an activator of differentiation but an inhibitor of proliferation by targeting MDFI in porcine pig satellite cells [14]. Miao et al. [15] proved that miR-194 functioned as a suppressor of the proliferation and migration of osteosarcoma cells. Similarly, upregulation of miR-194 had decreased cell viability and increased apoptosis in prostatic cancer cells [16]. Apart from this, miR-194 family was found to associate with PI3K/Akt signaling pathway [17], which has been proved to be necessary for the proliferation and differentiation of myoblasts [18,19,20].
In this study, we aimed to investigate the expression of miR-194-5p in the leg muscle of rabbit and reveal the possible role of miR-194-5p in the proliferation and differentiation of rabbit SMSCs through gain and loss function of miR-194-5p. Our results showed that miR-194-5p negatively regulated the proliferation or differentiation by acting on Mef2c.
Materials and methods
Ethics statement
All animals used in this experiment were treated properly according to the “Guidelines for Experimental Animals” enacted by Ministry of Science and Technology (Beijing, China). This study was reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of Sichuan Agricultural University, under the permit No. DKY-B20141401.
Sample collection and cell culture
Three New Zealand rabbits, aged 84 days, had free access to food and water. After being slaughtered, tissues (including heart, liver, spleen, lung, kidney, and muscle) were immediately collected and frozen in liquid nitrogen. SMSCs were isolated and purified as described in our laboratory [21]. SMSCs were cultured in growth media (GM) consisting of DMEM/HIGH GLUCOSE (DMEM) supplemented with 10% Fetal bovine serum (Gibco, Australia) and 2% penicillin streptomycin (Hyclone, USA). For differentiation induction, SMSCs were cultured in differentiation media (DM) consisting of DMEM with 2% horse serum (Beyotime, Shanghai, China) and 2% penicillin streptomycin (Hyclone, USA). All media had been changed every other day.
Cell transfection
Transfection was conducted when SMSCs reached 70–80% confluence in 24 well or 96 cell. SMSCs were transfected with miR-194-5p mimics (50 nM; 5′–3′ Sense: UGUAACAGCAACUCCAUGUGGA; Antisense: CACAUGGAGUUGCUGUUACAUU), negative control (NC, 50 nM; Sense: UUCUCCGAACGUGCUACGUTT; Antisense: AGCUGACACGUUCGGAGAATT), inhibitor (100 nM; UCCACAUGGAGUUGCUGUUACA), and inhibitor negative control (inNC, 100 nM; CAGUACUUUUGUGUAGUACAA) using Lipofectamine 3000 (Invitrogen, USA), respectively. Cells were exposed to transfection regents for 24 h or 36 h and then incubated in GM or DM as needed.
RT-qPCR
Total RNA was extracted from cells or tissues using RNAiso Reagent (Takara, Japan) according to the manufacturers’ protocol. The RNA concentration and purity were estimated using NanoDrop 2000 UV–Vis spectrophotometer (Thermo, Waltham, MA). The RNA quality was assessed using 1.5% agarose gel electrophoresis. The cDNA was synthesized using PrimeScript™ RT reagent Kit (Takara, Japan). RT-qPCR reaction was performed in triplicate using SYBR Premix EX Taq™II (Takara, Japan) and run on CFX96™ Real-Time PCR Detection System (Bio-Rad, USA). No-template controls and negative controls without cDNA template were also included in all qPCR runs. The relative expression levels of genes were normalized to the reference gene GAPDH using the 2−△△Ct method [22].
The synthesis and quantification of miRNA were conducted using Mir-X™ miRNA qRT-PCR SYBR® Kit (Takara, Japan) according to the manufactures’ protocol. The relative expression of miRNA was normalized to reference gene U6 snRNA using the 2−△△Ct method. All the corresponding primer sets are listed in Table S1.
Cell proliferation assay
When cells seeded at 2 × 103cells/well in 96-well plates reached about 70–80% density, transfection was conducted. 10 μL of Cell-Counting Kit-8 (CCK-8) reagent was added into wells for 2-h incubation at a 24-h interval. Each time the absorbance of cells was measured at a test wavelength of 450 nm using an automatic microplate reader.
In addition, EdU analysis was also conducted to explore the function of miR-194-5p in SMSCs proliferation. Cell preparation and transfection were also done as described previously. After transfection, cells were cultured in GM for 24 h. Then, SMSCs were incubated in medium containing 50 μM EdU for 2 h. Thereafter, cells were fixed, permeabilized, and stained by Hoechst 33,342 (1:1000) for 10 min at room temperature. In the end, images were obtained using a fluorescence microscope and the number of nuclei and EdU incorporation were analyzed using Image J software (National Institute of Health, Bethesda, MD). The percentage of EdU-positive cells was calculated by dividing the number of nuclei incorporating EdU by the number of total nuclei.
Luciferase reporter analysis
The target genes of miR-194-5p were predicted using online tools miRbase and TargetScan (https://www.mirbase.org/earch.shtml/https://www.targetscan.org/vert_71/). For luciferase assays, psiCHECK2-Mef2c-3′UTR (wild plasmid) and psiCHECK2-Mut-Mef2c-3′UTR (mutated plasmid) were constructed. Then wild plasmid or mutated plasmid were co-transfected with miR-194-5p mimics or NC into 293 T cells. After 36 h of incubation, cells were harvested and firefly luciferase (luc2) activity was measured and normalized to the Renilla luciferase (hRluc-neo) activity (luc2/hRluc-neo) using the Dual-Luciferase reporter assay system (TransGen Biotech, China).
Target gene prediction and gene ontology enrichment analysis
Online software miRWalk (https://mirwalk.umm.uni-heidelberg.de/) was used to predict the target genes of miR-194-5p. As there were no complete data of rabbit in the current version of miRWalk, prediction was conducted based on the miRNA-mRNA interaction of Mus musculus. Then the target genes were analyzed through gene ontology (GO) term enrichment analysis and KEGG pathway analysis using DAVID 6.8 (https://david.ncifcrf.gov/).
Statistical analysis
All data are expressed as mean ± SEM and were analyzed using GraphPad Prism 6.07 (GraphPad Software, San Diego, CA, USA). A p-value less than 0.05 indicates statistically significant difference.
Results
Bioinformatic analysis of the predicted target genes of miR-194-5p
A total of 16,190 target genes of miR-194-5p were predicted using the miRWalk online software and then used for GO term analysis (Fig. 1). First, these targets were mainly enriched in 9 biology progresses, including RNA polymerase II core promoter proximal region sequence-specific DNA binding, chromatin binding, ligase activity, ubiguitin-protein transferase activity, zinc ion binding, sequence-specific DNA binding, transcription factor activity/sequence-specific DNA binding, DNA binding, and protein binding. Second, these target genes were related to several cellular components consisting of Golgi membrane, cytoskeleton, membrane, cell junction, Golgi apparatus, cytosol, nucleus, cytoplasm, and nucleoplasm. Third, the analysis of molecular functions of these target genes showed that they mainly play a role in covalent chromatin modification, neural tube closure, multicellular organism development, nervous system development, regulation of transcription/DNA-templated, negative regulation of transcription from RNA polymerase II promoter, positive regulation of transcription/DNA-templated, transcription/DNA-templated, and positive regulation of transcription from RNA polymerase II promoter. Furthermore, the results of KEGG pathway analysis showed that these targets were mainly enriched in several pathways, including ubiquitin-mediated proteolysis, signaling pathways regulating pluripotency of stem cells, TGF-beta signaling pathway, Wnt signaling pathway, MAPK signaling pathway, Axon guidance, protein processing in endoplasmic reticulum, ErbB signaling pathway, Rap 1 signaling pathway, and Hippo signaling pathway (Table 1).
miR-194-5p expressed highly in rabbit leg muscle
The expression levels of miR-194-5p in various tissues (heart, liver, spleen, lung, kidney, and muscle) were detected using RT-qPCR. The result showed that miR-194-5p expressed significantly higher in the liver and leg muscle of rabbits than that in other tissues (Fig. 2).
miR-194-5p negatively regulates the proliferation of rabbit SMSCs
To investigate whether miR-194-5p plays a role in the proliferation of rabbit SMSCs, miR-194-5p mimics, inhibitor, and their negative controls were introduced into rabbit SMSCs, respectively. Next, both CCK-8 and EdU assays were used to detect the effects of miR-194-5p during the proliferation of rabbit SMSCs. The results of EdU assay showed that although there were no significant differences in proliferation of SMSCs between miR-194-5p mimics and NC groups, inhibition of miR-194-5p dramatically promoted the proliferation rate of SMSCs (Fig. 3a, b). On the other hand, the CCK-8 assay showed that miR-194-5p mimics significantly inhibited the proliferation of SMSCs at 120 h after transfection (Fig. 3c, d).
SMSCs were transfected with miR-194-5p mimics, negative control (NC), inhibitor negative control (inNC), and inhibitor for 48 h. The proliferation rate was represented by the percentage of EdU-positive cells a-b and CCK8 assay (c-d). The ratio of EdU-positive cells was calculated using the formula: (EdU-positive cells/ Hoechst stained cells) × 100% and the scale bar length was 200 μm. Values are presented as mean ± SEM of 3 pooled cells per group. *p < 0.05 or **p < 0.01 means the difference is significant
Mef2c is identified as one of the target genes of miR-194-5p
According to the KEGG pathway analysis, Mef2c was one of the potential target genes of miR-194-5p and a key gene in MAPK signaling pathway (Table 1; Fig. 4a). To confirm this relationship, recombined plasmids (wild plasmid and mutated plasmid) were constructed. Then 293 T cells were co-transfected by wild or mutated plasmid with the mimics or NC of miR-194-5p and the relative activity of firefly luciferase was analyzed using dual-luciferase reporter system. The result showed that luc2/hRluc-neo relative activity was significantly reduced after co-transfection of miR-194-5p mimics and wild plasmid but remained unchanged in other treatments. (Fig. 4b). To further verify the relationship between Mef2c and miR-194-5p, the expression of Mef2c in rabbit SMSCs was measured after transfection of miR-194-5p mimics, NC, inhibitor, and inNC, and the results showed that over-expression of miR-194-5p significantly suppressed the Mef2c expression, but there was no significant difference between the treatments of miR-194-5p inhibitor and inNC (Fig. 5).
Mef2c is a target gene of miR-194-5p. The binding sites of miR-194-5p are completely conservative in Homo sapiens, Mus musculus, Oryctolagus cuniculus, Pan troglodytes, and Sus scrofa (a). psiCHECK2-Mef2c and psiCHECK2-Mut-Mef2c plasmids were co-transfected with miR-194-5p mimics or NC into 293 T cells and the activity of firefly luciferase normalized by Renilla luciferase activity (luc2/hRluc-neo) was measured for 6 replicates (b). *p < 0.05 or **p < 0.01 means the difference is significant
miR-194-5p negatively regulated the differentiation of rabbit SMSCs
In order to define the role of miR-194-5p in the differentiation of rabbit SMSCs, differentiation of SMSCs were induced by replacing 10% FBS with 2% horse serum after 24-h transfection. During the differentiation of SMSCs, the expression levels of MyoG on Days 1, 3, 5, 7, and 9 were measured. The results showed that over-expression of miR-194-5p significantly suppressed the expression of MyoG on Days 3 and 7 during SMSCs differentiation. In contrast, inhibiting miR-194-5p expression had no significant effect on MyoG expression (Fig. 6).
Discussion
In recent years, a growing number of studies have demonstrated that miRNAs play a vital role in myogenesis by targeting muscle-specific genes [23, 24]. For instance, Bjorkman et al. [25] showed that miR-206 and miR-1 could downregulate Srsf9 expression to promote the differentiation of C2C12 cells. Over-expression of miR-675-3p and miR-675-5p promoted the differentiation of mouse myoblasts [26]. Apart from myogenesis-associated genes and miRNAs, a lot of signaling pathways, including PI3K/AKT, are also involved in regulating myogenic differentiation [27]. To date, many researchers have identified miR-194-5p as a regulator of tumorigenesis. For example, over-expression of miR-194 significantly inhibited the proliferation and migration of gastric cancer cells [28], and the enhanced expression of miR-194-5p promoted Gallbladder cancer cell proliferation by targeting AKT2 gene [29]. These researches indicated that the effect of miR-194-5p on cell proliferation are cell specific. A previous study showed Zfb609 circular RNA negatively regulated mouse myoblast differentiation by sponging miR-194-5p [30], suggesting miR-194-5p may play a positive role in the myogenesis of mouse myoblasts. However, it is unclear whether miR-194-5p play a role in the proliferation and differentiation of rabbit SMSCs.
Bioinformatics analysis of miR-194-5p displayed the potential functions of its target genes and several enriched signal pathways, like Wnt and MAPK signaling pathways, playing a role in myogenesis. Wnt/PCP pathway has been shown to play a role in the symmetric expansion of satellite cells [31]. Besides, activation of Wnt-β-catenin signaling induced follistatin and myogenin to promote myoblasts differentiation [32]. It has been proved that p38α promotes myoblasts differentiation by preventing cell proliferation [33]. Moreover, the activation of p38-MAPK pathway could promote the fusion of myoblasts to myotubes [34]. In the study, the expression of miR-194-5p was detected in various tissues of rabbits for the first time. RT-qPCR analysis showed miR-194-5p expressed the highest in leg muscle of rabbits, indicating that miR-194-5p might play a role in rabbit skeletal muscle development. It has been known that activated satellite cell pool served as a cell reservoir for the maintenance, hypertrophy, and repair of adult muscle [2]. The classic view proposed by Moss and Leblond [35] pointed that on account of the asymmetrical cell division, the population of SMSCs is heterogeneous containing myoblast precursor cells and quiescent satellite cells, which made SMSCs self-renew possible. Thus, it seems proliferation is the precondition of the formation of myoblast precursor cells. In our study, the EdU and CCK-8 assays showed that inhibiting miR-194-5p expression significantly promoted the proliferation rate of SMSCs, whereas over-expression of miR-194-5p suppressed the proliferation of SMSCs at 120 h after transfection, suggesting that miR-194-5p negatively regulated the proliferation of SMSCs. Until now, a lot of myogenesis-associated genes have been well defined, among which MEF-2 was widely expressed in muscle cells (cardiac, skeletal, and smooth muscle cells) [36]. It has also been reported that MEF-2 acts as a downstream target of the bHLH proteins (MyoD, MyoG, Myf5, etc.), and co-activation of MEF-2 and these bHLH proteins can activate the majority of muscle-specific genes [37]. Furthermore, Mef2c has been identified to repress gene expression by interacting with histone deacetylases and responding to various signaling pathways to activate gene expression after calcium influx, activation of calcineurin, and activation of MAPK signaling pathway [38,39,40]. In this study, Mef2c was first identified as one of the target genes of miR-194-5p using dual-luciferase assay. Upregulation of miR-194-5p in rabbit SMSCs significantly decreased the expression of Mef2c. Besides, the expression levels of MyoG on Day 3 and Day 7 during myogenesis was reduced by over-expression of miR-194-5p. Studies have proposed that Mef2c worked as a signal mediator of MAPK signaling pathway in cells [41, 42]. So, the results suggested that miR-194-5p inhibited the differentiation of rabbit SMSCs via inhibition of MAPK signaling pathway by acting on Mef2c.
In summary, miR-194-5p was revealed to be highly expressed in the leg muscle of rabbits. At the cellular level, it negatively regulated the proliferation of rabbit SMSCs. Moreover, the over-expression of miR-194-5p decreased the expression of MyoG, suggesting that miR-194-5p inhibited the differentiation of rabbit SMSCs by inactivating the MAPK signaling pathway through targeting Mef2c.
References
Dumont NA, Bentzinger CF, Sincennes MC, Rudnicki MA (2015) Satellite cells and skeletal muscle regeneration. Compr Physiol 5:1027–1059. https://doi.org/10.1002/cphy.c140068
Zammit PS, Partridge TA, Yablonka-Reuveni Z (2006) The skeletal muscle satellite cell: the stem cell that came in from the cold. J Histochem Cytochem 54:1177–1191. https://doi.org/10.1369/jhc.6R6995.2006
Klein WH, Hasty P, Bradley A, Morris JH, Edmondson DG, Venuti JM, Olson EN (1993) Muscle deficiency and neonatal death in mice with a targeted mutation in the myogenin gene. Nature 364:501–506
Arnold HH, Winter B (1998) Muscle differentiation: more complexity to the network of myogenic regulators. Curr Opin Genet Dev 8:539–544. https://doi.org/10.1016/s0959-437x(98)80008-7
Black BL, Olson EN (1998) Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins. Annu Rev Cell Dev Biol 14:167–196. https://doi.org/10.1146/annurev.cellbio.14.1.167
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. https://doi.org/10.1038/nature02871
Hwang H, Mendell J (2006) MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 94:776
Shivdasani RA (2006) MicroRNAs: regulators of gene expression and cell differentiation. Blood 108:3646–3653
Chen JF, Tao Y, Li J, Deng Z, Yan Z, Xiao X, Wang DZ (2010) microRNA-1 and microRNA-206 regulate skeletal muscle satellite cell proliferation and differentiation by repressing Pax7. J Cell Biol 190:867–879. https://doi.org/10.1083/jcb.200911036
Yu X, Zhang L, Wen G, Zhao H, Luong L, Chen Q, Huang Y, Zhu J, Ye S, Xu Q (2015) Upregulated sirtuin 1 by miRNA-34a is required for smooth muscle cell differentiation from pluripotent stem cells. Cell Death Differ 22:1170
Koutsoulidou A, Mastroyiannopoulos NP, Furling D, Uney JB, Phylactou LA (2011) Expression of miR-1, miR-133a, miR-133b and miR-206 increases during development of human skeletal muscle. BMC Dev Biol 11:34
Tong H, Jiang R, Liu T, Wei Y, Li S, Yan Y (2018) bta-miR-378 promote the differentiation of bovine skeletal muscle-derived satellite cells. Gene 668:246–251. https://doi.org/10.1016/j.gene.2018.03.102
Zhang WR, Zhang HN, Wang YM, Dai Y, Liu XF, Li X, Ding XB, Guo H (2017) miR-143 regulates proliferation and differentiation of bovine skeletal muscle satellite cells by targeting IGFBP5. Vitro Cell Dev Biol Anim 53:265–271. https://doi.org/10.1007/s11626-016-0109-y
Hou L, Xu J, Jiao Y, Li H, Pan Z, Duan J, Gu T, Hu C, Wang C (2018) MiR-27b promotes muscle development by inhibiting MDFI expression. Cell Physiol Biochem 46:2271–2283. https://doi.org/10.1159/000489595
Miao J, Wang W, Wu S, Zang X, Li Y, Wang J, Zhan R, Gao M, Hu M, Li J, Chen S (2018) miR-194 suppresses proliferation and migration and promotes apoptosis of osteosarcoma cells by targeting CDH2. Cell Physiol Biochem 45:1966–1974. https://doi.org/10.1159/000487973
Gao S, Zhao Z, Wu R, Wu L, Tian X, Zhang Z (2018) MicroRNA-194 regulates cell viability and apoptosis by targeting CDH2 in prostatic cancer. Onco Targets Ther 11:4837–4844. https://doi.org/10.2147/ott.S169101
Wang T, Cheng Y, Han H, Liu J, Tian B, Liu X (2019) miR-194 accelerates apoptosis of Aβ(1-42)-transduced hippocampal neurons by inhibiting Nrn1 and decreasing PI3K/Akt signaling pathway activity. Genes (Basel). https://doi.org/10.3390/genes10040313
Huang W, Guo L, Zhao M, Zhang D, Xu H, Nie Q (2019) The Inhibition on MDFIC and PI3K/AKT pathway caused by miR-146b-3p triggers suppression of myoblast proliferation and differentiation and promotion of apoptosis. Cells. https://doi.org/10.3390/cells8070656
Sun W, Hu S, Hu J, Yang S, Hu B, Qiu J, Gan X, Liu H, Li L, Wang J (2019) miR-365 inhibits duck myoblast proliferation by targeting IGF-I via PI3K/Akt pathway. Biosci Rep. https://doi.org/10.1042/bsr20190295
Fu S, Yin L, Lin X, Lu J, Wang X (2018) Effects of cyclic mechanical stretch on the proliferation of L6 myoblasts and its mechanisms: PI3K/Akt and MAPK signal pathways regulated by IGF-1 receptor. Int J Mol Sci. https://doi.org/10.3390/ijms19061649
Liu B, Shi Y, He H, Cai M, Xiao W, Yang X, Chen S, Jia X, Wang J, Lai S (2018) miR-221 modulates skeletal muscle satellite cells proliferation and differentiation. In Vitro Cell Dev Biol Anim 54:147–155
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C T method. Nat Protoc 3:1101
Güller I, Russell AP (2010) MicroRNAs in skeletal muscle: their role and regulation in development, disease and function. J Physiol 588:4075–4087
Williams AH, Liu N, van Rooij E, Olson EN (2009) MicroRNA control of muscle development and disease. Curr Opin Cell Biol 21:461–469
Bjorkman KK, Buvoli M, Pugach EK, Polmear MM, Leinwand LA (2019) miR-1/206 downregulates splicing factor Srsf9 to promote C2C12 differentiation. Skelet Muscle 9:31. https://doi.org/10.1186/s13395-019-0211-4
Dey BK, Pfeifer K, Dutta A (2014) The H19 long noncoding RNA gives rise to microRNAs miR-675-3p and miR-675-5p to promote skeletal muscle differentiation and regeneration. Genes Dev 28:491–501. https://doi.org/10.1101/gad.234419.113
Briata P, Lin W-J, Giovarelli M, Pasero M, Chou C-F, Trabucchi M, Rosenfeld MG, Chen C-Y, Gherzi R (2012) PI3K/AKT signaling determines a dynamic switch between distinct KSRP functions favoring skeletal myogenesis. Cell Death Differ 19:478–487
Zhao X, Hou Y, Tuo Z, Wei F (2018) Application values of miR-194 and miR-29 in the diagnosis and prognosis of gastric cancer. Exp Ther Med 15:4179–4184. https://doi.org/10.3892/etm.2018.5931
Wang S-H, Wu X-C, Zhang M-D, Weng M-Z, Zhou D, Quan Z-W (2016) Long noncoding RNA H19 contributes to gallbladder cancer cell proliferation by modulated miR-194–5p targeting AKT2. Tumour Biol 37:9721–9730
Wang Y, Li M, Wang Y, Liu J, Zhang M, Fang X, Chen H, Zhang C (2019) A Zfp609 circular RNA regulates myoblast differentiation by sponging miR-194-5p. Int J Biol Macromol 121:1308–1313. https://doi.org/10.1016/j.ijbiomac.2018.09.039
Le Grand F, Jones AE, Seale V, Scimè A, Rudnicki MA (2009) Wnt7a activates the planar cell polarity pathway to drive the symmetric expansion of satellite stem cells. Cell Stem Cell 4:535–547. https://doi.org/10.1016/j.stem.2009.03.013
Jones AE, Price FD, Le Grand F, Soleimani VD, Dick SA, Megeney LA, Rudnicki MA (2015) Wnt/β-catenin controls follistatin signalling to regulate satellite cell myogenic potential. Skelet Muscle 5:14. https://doi.org/10.1186/s13395-015-0038-6
Perdiguero E, Ruiz-Bonilla V, Serrano AL, Muñoz-Cánoves P (2007) Genetic deficiency of p38alpha reveals its critical role in myoblast cell cycle exit: the p38alpha-JNK connection. Cell Cycle 6:1298–1303. https://doi.org/10.4161/cc.6.11.4315
Lluís F, Perdiguero E, Nebreda AR, Muñoz-Cánoves P (2006) Regulation of skeletal muscle gene expression by p38 MAP kinases. Trends Cell Biol 16:36–44. https://doi.org/10.1016/j.tcb.2005.11.002
Moss FP, Leblond CP (1971) Satellite cells as the source of nuclei in muscles of growing rats. Anat Rec 170:421–435. https://doi.org/10.1002/ar.1091700405
Arnold MA, Kim Y, Czubryt MP, Phan D, McAnally J, Qi X, Shelton JM, Richardson JA, Bassel-Duby R, Olson EN (2007) MEF2C transcription factor controls chondrocyte hypertrophy and bone development. Dev Cell 12:377–389. https://doi.org/10.1016/j.devcel.2007.02.004
Naidu PS, Ludolph DC, To RQ, Hinterberger TJ, Konieczny SF (1995) Myogenin and MEF2 function synergistically to activate the MRF4 promoter during myogenesis. Mol Cell Biol 15:2707–2718
Blaeser F, Ho N, Prywes R, Chatila TA (2000) Ca(2+)-dependent gene expression mediated by MEF2 transcription factors. J Biol Chem 275:197–209. https://doi.org/10.1074/jbc.275.1.197
Zhao M, New L, Kravchenko VV, Kato Y, Gram H, di Padova F, Olson EN, Ulevitch RJ, Han J (1999) Regulation of the MEF2 family of transcription factors by p38. Mol Cell Biol 19:21–30. https://doi.org/10.1128/mcb.19.1.21
Youn HD, Grozinger CM, Liu JO (2000) Calcium regulates transcriptional repression of myocyte enhancer factor 2 by histone deacetylase 4. J Biol Chem 275:22563–22567. https://doi.org/10.1074/jbc.C000304200
Khiem D, Cyster JG, Schwarz JJ, Black BL (2008) A p38 MAPK-MEF2C pathway regulates B-cell proliferation. Proc Natl Acad Sci USA 105:17067–17072. https://doi.org/10.1073/pnas.0804868105
Jiang K, Teng GD, Chen YQ (2020) MicroRNA-23 suppresses osteogenic differentiation of human bone marrow mesenchymal stem cells by targeting the MEF2C-mediated MAPK signaling pathway. J Gene Med. https://doi.org/10.1002/jgm.3216
Acknowledgements
This research was funded by the earmarked fund for China Agricultural Research System [Grant No. CARS-43-A-2].
Author information
Authors and Affiliations
Contributions
YS and XM designed this study. XM and YS performed the experiments and analyzed the data. YS, MC, and SH drafted the paper. JW, XJ, XL, and SC participated in animal manipulation, data collection, and analysis. SL supervised this study and reviewed this manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shi, Y., Mao, X., Cai, M. et al. miR-194-5p negatively regulates the proliferation and differentiation of rabbit skeletal muscle satellite cells. Mol Cell Biochem 476, 425–433 (2021). https://doi.org/10.1007/s11010-020-03918-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-020-03918-0