Abstract
Context
Land use change reduced calcareous grasslands throughout Europe during the last decades. Subsequent fragmentation and habitat deterioration led, moreover, to a massive biodiversity decline. To counteract this alarming development, a clear understanding of genetic variation patterns, as fundamental level of biodiversity, becomes inevitable.
Objectives
The aim of our study was to identify the drivers of genetic variation in common calcareous grassland plant species. More specifically, we tested whether genetic diversity or differentiation of Asperula cynanchica, Campanula rotundifolia, and Linum catharticum depend on habitat age, landscape structure, habitat quality, and/or population size.
Methods
We investigated 912 individuals, 304 per study species, from 19 calcareous grasslands across the Swabian Alb in Baden-Württemberg (Germany) using AFLP analyses.
Results
We observed no significant influence of habitat age on genetic diversity and differentiation. Habitat quality also had no impact on genetic diversity and population size only showed weak effects. However, genetic diversity strongly depended on landscape structure represented by distance to the nearest settlement, total area of surrounding calcareous grasslands, and their connectivity.
Conclusions
Migratory sheep herding is considered as main land use in calcareous grasslands on the Swabian Alb and thus, landscape structures in the study region may describe movement patterns of grazing livestock. In this study, genetic variation in calcareous grassland populations was strongly affected by surrounding landscape structures and subsequent grazing patterns. Therefore, we assume that moderate grazing intensities over the long term may increase levels of genetic diversity, whereas periods of overgrazing or abandonment could lower genetic diversity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Central European calcareous grasslands represent local biodiversity hotspots due to their long existence, habitat diversity, and species richness (Steffan-Dewenter and Tscharntke 2002; Karlik and Poschlod 2009; Poschlod 2017). More precisely, they constitute valuable habitats for many specialised, rare, and endangered plant or insect species and are, therefore, considered as key areas for biodiversity conservation in agricultural landscapes (Raatikainen et al. 2009; Rosengren et al. 2013).
The shift from traditional to modern (animal) husbandry caused a drastic decline of calcareous grasslands during the last 150 years (WallisDeVries et al. 2002; Poschlod 2017). Due to abandonment and intensification more than 70% of the calcareous grasslands on the Swabian Alb in south-west Germany disappeared by the 1990s (Mattern et al. 1992; Steffan-Dewenter and Tscharntke 2002). Remnant calcareous grasslands are often highly fragmented and small in size. Populations in these habitat patches may consequently suffer from reduced probabilities of gene flow and increased genetic drift (Aguilar et al. 2008). Therefore, habitat loss affects not only biodiversity at the species level, but also the genetic variation of local plant populations (Ouborg et al. 2006). Following May (1994), genetic variation represents the most fundamental level of biodiversity. Levels of genetic variation are shaped by changing environmental conditions driving natural selection, adaptation, gene flow, genetic drift, and stochastic processes (McKay et al. 2005; Rosengren et al. 2013). To protect biodiversity fundamentally, we need to identify the key variables influencing genetic variation.
Calcareous grasslands are characterized by a diverse land use history as well as management continuity and could, therefore, be found either on historically old (‘ancient’) or historically young (‘recent’) sites. High levels of gene flow at the time of founding and afterwards may result in identical levels of genetic variation between sites of different age (Vandepitte et al. 2010). Nevertheless, the genetic variation of populations on recent sites seems to depend on both the number and origin of colonists (Wade and McCauley 1988) as well as the rate of gene flow and selection after colonization (Barrett et al. 2008). These populations may, therefore, show reduced genetic variation by bottlenecks and increased divergence among populations by selection (Wade and McCauley 1988; Dlugosch and Parker 2008). Due to potential past and present bottleneck, selection, or gene flow events, we would expect an impact of habitat age on the genetic variation of typical calcareous grassland species.
Past and present landscape structures provide valuable information about potential gene flow and further dispersal processes (Prentice et al. 2006; Purschke et al. 2012). The impact of both habitat size and area of surrounding habitats on biodiversity was analysed for many species groups and habitats, since MacArthur and Wilson (1967) established the theory of island biogeography. Hence, various studies reported that plant populations on small and isolated calcareous grasslands, with reduced gene flow, increased inbreeding as well as genetic drift, showed reduced seed set (Kéry et al. 2000), genetic erosion (Honnay et al. 2007), and finally higher extinction risks (Spielman et al. 2004). Besides habitat size, habitat connectivity and the kind of grazing management supply essential information about possible gene flow and seed dispersal in networks of (fragmented) habitat patches (Reitalu et al. 2010). Grazing, e.g. by sheep (typically for calcareous grasslands), ensures propagule dispersal over large distances and improves habitat quality by trampling and browsing (Fischer et al. 1996; Willerding and Poschlod 2002). Such highly connected sites are expected to show increased colonization and reduced extinction rates due to rescue effects (Brown and Kodric-Brown 1977). Thus, it can be hypothesized that surrounding landscape structures and resulting gene flow mechanisms are important determinants for genetic variation in highly diverse calcareous grasslands.
The abandonment of migratory sheep farming and, in turn, lower grazing pressure on calcareous grasslands led to deteriorated habitat conditions in the last decades (Zulka et al. 2014). Biomass is not removed in the absence of grazing animals, which results in litter accumulation, eutrophication, and thus, increasing vegetation height with grasses dominating (Jacquemyn et al. 2011). The germination of calcareous grassland species depends on the availability of light and open soil (Grubb 1977). Moreover, thick litter layers acting as seed traps (Ruprecht and Szabó 2012) and high vegetation causing ground shadowing (Jensen and Gutekunst 2003) inhibit germination and establishment of these species. Therefore, an impact of the local vegetation structure, which is also an indicator for habitat quality, on genetic variation can be expected.
Despite intact habitat quality, habitat fragmentation could lead to isolated populations with decreased population size. Small populations are more sensitive to demographic and environmental changes due to the fixation of deleterious alleles by genetic drift (Young et al. 1996). These populations will show lower genetic variability, consequently increased levels of inbreeding (Van Treuren et al. 2005), and therefore, face a higher risk of extinction (Spielman et al. 2004; Ouborg et al. 2006). Many empirical studies observed a positive impact of population size on the genetic variation of calcareous grassland species (Leimu et al. 2006) and thus, we predict a positive association between population size and genetic variation.
Considering all these aspects, the aim of this study was to disentangle the relative impact of biotic and abiotic factors on the genetic variation of common calcareous grassland species. In changing environments, gene flow, migration, and/or dispersal potential of species may be represented by different levels of genetic variation (Holderegger et al. 2006). Thus, we asked the following questions: (i) Is genetic diversity influenced by habitat age? Are populations of different habitat age genetically differentiated? (ii) What is the impact of past and/or present landscape structure on genetic diversity? (iii) Is genetic diversity affected by the present habitat quality and/or population size?
Materials and methods
Study design
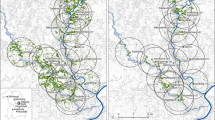
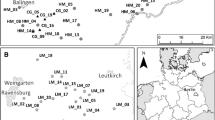
For our study, we selected 19 calcareous grasslands all over the Swabian Alb in south-west Germany (Fig. 1, Table S1). This region belongs to the largest Jurassic low mountain range in Central Europe (Park 2017). The climate is characterized by cool, humid westerly winds with an annual average temperature between 6.7 and 8.0 °C and an average precipitation from 750 to 1050 mm/year (Jooß 2014).
In order to study the impact of habitat age on genetic diversity and differentiation of common calcareous grassland species, we sampled populations on sites with different habitat age (Reitalu et al. 2010). Selected patches were currently calcareous grassland habitats with comparable habitat quality. We selected ten historically old patches (‘ancient sites’), which are calcareous grasslands since before the 1820s, and nine historically young patches (‘recent sites’), which developed from arable fields during the 1900s (Fig. 1, Table S1). The habitat age was determined using historical cadastral maps from 1820 to 1850 as well as 1902 to 1914. Further, topographical maps from 1951 to 1953 and actual aerial photographs were examined using the software ArcGIS® 10.3.1 (Esri, Redlands, CA, USA) (Table S2).
At each site, we analysed genetic variation of three typical calcareous grassland species: Asperula cynanchica L., Campanula rotundifolia L. s. str., and Linum catharticum L. Asperula cynanchica (Rubiaceae; 2n = 22, (44)) is flowering from June to September (Kühn et al. 2004). Main pollinators are insects, e.g. bees, bumblebees, wasps, bombylides, or syrphids, but occasionally A. cynanchica is self-pollinated (Kühn et al. 2004). The bell-shaped flowers of C. rotundifolia (Campanulaceae; 2n = 34, 68) are mostly pollinated by bees between June and October (Kühn et al. 2004). The annual L. catharticum (Linaceae; 2n = 16), which can sometimes live longer, flowers from May to July (Kühn et al. 2004). It is generally self-pollinated, but could also be insect-pollinated by bees, bumblebees, wasps, bombylides, or syrphids (Kühn et al. 2004). All three species can be dispersed ecto- and/or endozoochorously (Poschlod et al. 2003).
To analyse the impact of landscape structure on genetic diversity, we digitized historical cadastral maps (1820–1850) as well as actual topographical maps (2014–2018) in a circle with 3 km radius around the center of each study site (Table S2). As potential explanatory variables, we identified the area of each study site (AREA_S) and measured the past and present distance to the nearest settlement (DIST_1820; DIST_2018) (Table S3). Additionally, we calculated the past and present total area of surrounding calcareous grasslands per 3 km radius (AREA_1820; AREA_2018) as well as the past and present connectivity (CON_1820; CON_2018) per circle (Table S3). The connectivity was determined according to Hanski (1994) as Si = ∑j≠i exp (− αdij) Aj where Si is the connectivity of the patch i, dij is the distance (km) between patches i and j, Aj is the area (ha) of the patch j, and α is the parameter of the exponential distribution setting the influence of distance on connectivity (Helm et al. 2006). Following Lindborg and Eriksson (2004) and Reitalu et al. (2010) α was set to one and not weighted by the dispersal abilities of the plant species in the community.
Data about the cover of vascular plants, mosses, litter, and open soil were incorporated per study site to investigate the influence of habitat quality on genetic diversity (Table S4). Furthermore, population size was determined by counting the number of individuals in 10–15 1 m2 randomly placed plots in the field. The average number of individuals per square metre was then multiplied with AREA_S (Reisch et al. 2018) (Table S4). For those study sites, where no individual could be found in the 1 m2 plots although plant material was collected, the total number of individuals was set from 0 to 1 before multiplying.
For molecular analyses we took leaf samples from 16 individuals per population and species to cover more than 90% of the total genetic diversity (Leipold et al. 2020).
Molecular analyses
DNA extraction was conducted following the CTAB protocol from Rogers and Bendich (1994) modified by Reisch (2007). DNA quality and concentration were determined with a spectrophotometer. All DNA samples were diluted to the same level of 7.8 ng DNA per µl H2O. Genetic variation within populations was determined for 912 individuals using genome-wide genotyping with amplified fragment length polymorphisms (AFLP; Vos et al. 1995). The AFLP analyses were performed following the standardized protocol of Beckmann Coulter (Bylebyl et al. 2008; Reisch 2008). We screened 36 primer combinations per species to choose three appropriate primer combinations for the selective amplification (Table S5). An automated capillary electrophoresis machine (GeXP, Beckmann Coulter) was used to separate the fluorescence-labelled DNA fragments by capillary gel electrophoresis. Fragment data were analysed manually applying the software Bionumerics 4.6 (Applied Maths, Kortrijk, Belgium). Only strong and clearly defined fragments were taken into account for further analyses and thus, samples without clear banding pattern were repeated.
The reproducibility of the AFLP analyses was tested by calculating the genotyping error rate (Bonin et al. 2004). Therefore, 10% of all analysed samples were replicated twice and the percentage of fragments with differences between original and replicate was evaluated. The genotyping error rates of A. cynanchica, C. rotundifolia, and L. catharticum were 2.6%, 4.2%, and 2.5% respectively.
Statistical analyses
Binary (0/1) matrices were created applying Bionumerics 4.6. Using this matrices, genetic diversity within each population was calculated as Nei’s gene diversity (GD) H = 1 − ∑(pi)2, with pi representing the allele frequency, in PopGene 32 (Yeh et al. 1997). We calculated a Kruskal–Wallis test with a post hoc Dunn’s test and a Bonferroni p-adjustment in R (R Core Team 2017) to compare Nei’s gene diversity on species level and to test the dependence of Nei’s gene diversity on habitat age.
Hierarchical analyses of molecular variance, AMOVA, based on pairwise Euclidian distances among samples, were conducted applying the software GenAlEx 6.41 (Peakall and Smouse 2006). Thus, the genetic variation within and among populations as well as among populations of different habitat age was analysed.
Mantel tests with 999 permutations were calculated using GenAlEx 6.41 (Peakall and Smouse 2006) to test the correlation between geographic and genetic distances (ΦPT values calculated in the AMOVA) among populations (Mantel 1967).
We then built a starting model with the full set of scaled and centred explanatory variables (except for habitat age) to analyse the relationship between genetic diversity and potential explanatory variables. More specifically, we formulated multiple linear regression models for each species in R (R Core Team 2017) to describe the variation of Nei’s gene diversity related to (i) habitat age, (ii) area of each study site, (iii) past and (iv) present total area of surrounding calcareous grasslands, (v) past and (vi) present distance to the nearest settlement, and (vii) past and (viii) present connectivity, which were described above. Further data about the coverage of (ix) vascular plants, (x) mosses, (xi) litter, and (xii) open soil, as well as the (xiii) population size of each species were included per study site. We then ranked all potential linear models according to AICc values (Akaike Information Criterion corrected for small sample sizes) to detect the models with the highest information content (Burnham and Anderson 2002). Differences between past and present landscape variables were tested by calculating Wilcoxon–Mann–Whitney tests.
Correlation tests (Pearson correlation coefficients) were conducted to show potential collinearity between the explanatory variables (ii–xiii) and to avoid, therefore, false interpretation of the linear regression models (Table S7).
Results
AFLP analyses resulted in 148, 151, and 146 fragments for A. cynanchica, C. rotundifolia, and L. catharticum. 69.59%, 68.49%, and 44.81% of these fragments were polymorphic. No identical genotypes were detected.
Nei’s gene diversity of A. cynanchica populations ranged between 0.21 and 0.30 (GDmean = 0.27) (Table 1). Campanula rotundifolia populations showed with 0.24 a lower mean Nei’s gene diversity than A. cynanchica (p = 0.054) (Fig. 2a). It ranged from 0.22 to 0.26 (Table 1). Linum catharticum populations indicated a significantly lower mean Nei’s gene diversity (GDmean = 0.16) than A. cynanchica (p < 0.001) and C. rotundifolia (p < 0.001) (Fig. 2a). The lowest value was 0.13 and the highest 0.19 (Table 1). AMOVAs (Table 2) indicated only weak levels of differentiation among populations of A. cynanchica (ΦPT = 0.072), C. rotundifolia (ΦPT = 0.048), and L. catharticum (ΦPT = 0.078).
Moreover, AMOVAs revealed no differentiation among populations on ancient and recent grasslands (Table 2) Furthermore, genetic diversity did not differ significantly among populations on ancient and recent sites (Fig. 2b).
Mantel tests revealed significant correlations between pairwise genetic and geographic distances for A. cynanchica (r = 0.41; p = 0.001) (Fig. 3a) and C. rotundifolia (r = 0.37; p = 0.001) (Fig. 3b), but not for L. catharticum (r = 0.06; p = 0.263) (Fig. 3c).
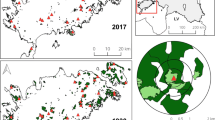
Our study indicated a significant decline of the total area of calcareous grasslands, their connectivity, and their distance to the nearest settlement between the 1820s and 2018 (Table S6).
The AICc model selection generated significant linear models for all studied species (Table 3a–c). Nei’s gene diversity of A. cynanchica populations was positively associated with present connectivity (p < 0.001). Nevertheless, past total area of surrounding calcareous grasslands (p = 0.019), present distance to the nearest settlement (p < 0.001), and the species’ population size (p < 0.001) displayed a negative impact (Table 3a). Genetic diversity of C. rotundifolia populations increased with rising past distance to the nearest settlement (p = 0.022) (Table 3b), while Nei’s gene diversity of L. catharticum populations was positively linked to present total area of surrounding calcareous grasslands (p = 0.021) (Table 3c).
Discussion
Genetic variation
Mean genetic diversity of our study species A. cynanchica, C. rotundifolia, and L. catharticum complied with the genetic diversity previously reported for common grassland species (Reisch and Bernhardt-Römermann 2014). Generally, genetic diversity depends on a species’ pollination and mating system (Schoen and Brown 1991). Therefore, the insect pollinated, outcrossing species A. cynanchica and C. rotundifolia revealed significantly higher genetic diversity levels than L. catharticum, which is mostly self-pollinated (Kühn et al. 2004).
Weak levels of differentiation and comparatively low ΦPT values among populations led to the assumption that the spatial distance among populations (< 100 km) still seems to allow sufficient gene flow (Neel 2008), since seed dispersal by animals may exceed distances of 100 km or more (Fischer et al. 1996; Manzano and Malo 2006).
Habitat age
Our study revealed similar levels of genetic diversity concerning habitat age. Following Rosengren et al. (2013) the genetic diversity of recent sites may be increased if they are connected to continuously grazed ancient sites with a diverse gene pool. Therefore, sufficient gene flow at the time of founding and afterwards might reduce the effects of habitat age (Vandepitte et al. 2010).
Furthermore, we observed no significant differentiation among populations on ancient and recent grassland sites. Genetic differentiation is often described as a direct function of dispersal (Oostermeijer et al. 1996). More than 50% of a local species pool could be transported by one sheep during a vegetation period (Fischer et al. 1996). Thus, especially dispersal by sheep is thought to have a detectable effect on the genetic variation of grazed calcareous grassland populations (Willerding and Poschlod 2002; Rico et al. 2014a, b). The suggested dispersal rate of 660,000 diaspores per 400-head sheep flock (Willerding and Poschlod 2002) results in a substantial gene flow over long time periods and large distances (Fischer et al. 1996; Poschlod et al. 1998; Poschlod 2017). Furthermore, hayseed of populations on ancient sites was used to establish calcareous grasslands artificially on abandoned arable fields until the 20th century (Poschlod and WallisDeVries 2002). According to the migrant pool model (Wade and McCauley 1988), genetic divergence could only occur if the number of colonists is less than twice the number of migrants.
Anthropogenic land use allows comparatively high levels of gene flow (Neel 2008) and could, therefore, establish viable populations in a relatively short time by overcoming pronounced founder effects (Helsen et al. 2013). By this means, anthropogenic land use may have led to similar diversity levels as well as undetectable genetic differentiation among populations of different habitat age.
Landscape structure
During the last century, land use change caused a quantitative decline of semi-natural grasslands, especially of calcareous grasslands (Poschlod et al. 2005). More particularly, settlement expansion (Poschlod 2017) or abandonment of migratory sheep farming caused massive habitat loss (WallisDeVries et al. 2002). The present study revealed a significant decline of the total area of calcareous grasslands, their connectivity, and their distance to the nearest settlement between the 1820s and 2018 as well.
However, present connectivity appeared as the only positive explanatory variable for the genetic diversity of the analysed A. cynanchica populations. This result is corroborated by the findings of Raatikainen and Heikkinen (2009), although other studies revealed only an influence of the past connectivity on grassland species (Lindborg and Eriksson 2004; Helm et al. 2006). The model indicates the presence of a rescue effect (Brown and Kodric-Brown 1977). Thus, populations in small habitat patches could not only persist with a high probability (Helm et al. 2006), they even show increased genetic diversity if they are well connected.
In general, the distance to the nearest settlement and the area of surrounding calcareous grasslands may describe the movement patterns of livestock (Reitalu et al. 2010) because migratory sheep herding was the main land use in calcareous grasslands of the study region. Migratory sheep herding represents both an important vector for seed dispersal (Fischer et al. 1996; Willerding and Poschlod 2002) and ecological disturbance by grazing and trampling (Olff and Ritchie 1998). Thus, overgrazing may lead to increased levels of gene flow and disturbance, while abandonment of migratory sheep herding is expected to reduce levels of gene flow as well as the probability of seedling establishment due to a denser litter and vegetation layer (Ruprecht and Szabó 2012). Intermediate levels of gene flow may reveal a positive impact on genetic diversity levels, while ‘too low’ and even ‘too high’ levels of gene flow may promote outbreeding depression and/or genetic ‘swamping’ (Bradshaw 1984). Overgrazing and abandonment of migratory sheep herding may, therefore, decrease both species (Klimek et al. 2007) and genetic diversity.
Unexpectedly, C. rotundifolia populations showed a positive impact of past distance to the nearest settlement on Nei’s gene diversity, resulting in decreased levels of genetic diversity around settlements. Although the relationship was not very strong, we assume that grasslands close to those settlements may reflect the impact of periodic overgrazing with increased levels of gene flow and disturbance during the 1820s.
Nevertheless, the linear model for A. cynanchica displayed a negative impact of present distance to the nearest settlement. Thus, the highest levels of genetic diversity occurred in populations near settlements in 2018. Despite nowadays ongoing decline of livestock grazing (Poschlod 2017), we suspect that grazing intensity and associated gene flow are still at an intermediate level around present settlements. Based on the model of Reitalu et al. (2010), an unimodal association between genetic diversity and the distance to the nearest settlement could be considered as possible explanation. The authors interpreted this result in terms of the intermediate disturbance hypothesis of Connell (1978) on a landscape scale. In calcareous grasslands, the landscape scale corresponds to the intensity of grazing and associated disturbance. Therefore, these results could also be explained in terms of a classic intermediate disturbance hypothesis (Connell 1978).
Unexpectedly, the linear model for A. cynanchica displayed a negative impact of the past total area of surrounding calcareous grasslands on genetic diversity. This idiosyncratic result is inconsistent with the generally accepted expectation that populations, which are embedded in a landscape matrix containing a large proportion of grasslands, are more likely to reveal high levels of genetic diversity (Rosengren et al. 2013). Therefore, we suggest that comparatively high levels of gene flow in the past may have led to a highly unified and impoverished gene pool. Moreover, A. cynanchica populations could have also been affected by periodic overgrazing with increased levels of disturbance during the 1820s. Thus, A. cynanchica still seems to suffer from a kind of ‘over-connection’ with comparatively high measures of gene flow and/or increased levels of disturbance by grazing animals during the 1820s.
Nei’s gene diversity of L. catharticum populations was positively associated with the present total area of surrounding calcareous grasslands. On the one hand, small and isolated habitat fragments may show reduced fitness levels and finally extinction if they suffer from edge effects and the invasion of generalist species (Leimu et al. 2006). On the other hand, a large patch size and a high proportion of surrounding grasslands may increase the variability of the incoming gene flow (Prentice et al. 2006). Thus, the total area of surrounding grassland patches has not only a positive effect on species richness and presence (Raatikainen et al. 2009), it could also increase the genetic diversity of species (Dahlström et al. 2006). Although the relationship was not very strong, levels of genetic diversity seem to come up with an intermediate gene flow level, since significantly lower values of the present total area of surrounding calcareous grasslands (compared to the past total area of surrounding calcareous grasslands) positively affected genetic diversity in L. catharticum. Therefore, high genetic diversity seems to depend on an intermediate size level of surrounding calcareous grasslands or rather gene flow.
However, following Jacquemyn et al. (2006), pollination as well as dispersal vectors determine gene flow over great geographic distances. The analysed species revealed different isolation by distance patterns, although all three species are dispersed ecto- and/or endozoochorously (Poschlod et al. 2003), e.g. by grazing sheep. The insect pollinated perennials, A. cynanchica and C. rotundifolia, showed lower gene flow over increasing distances (isolation by distance) because pollinating insects may rarely travel distances larger than 1 km (Kwak et al. 1998; Steffan-Dewenter and Tscharntke 2002). The mainly self-pollinated L. catharticum did, therefore, not reveal any isolation by distance.
Habitat quality and population size
The habitat quality showed no impact on the genetic diversity of the analysed calcareous grassland species. However, the linear model for A. cynanchica displayed an influence of the species’ population size. Although correlations between population size and genetic diversity are generally positive (Leimu et al. 2006), the genetic diversity of A. cynanchica decreased with increasing population size. Grassland plant species with comparatively large population size, long life cycles, and slow intrinsic dynamics may occur as remnant populations in modern landscapes (Maurer et al. 2003). Additionally, Piqueray et al. (2011) observed that the present occurrence of species can be influenced by past habitat configuration. These species often show a time lag between habitat loss, fragmentation, and their consequences on genetic diversity (Helm et al. 2006). Various studies revealed a significant relationship between the genetic diversity and the linkage of the studied populations in the past landscape. Thus, they indicated a delayed response of genetic diversity to habitat fragmentation (Honnay et al. 2007). The total area of calcareous grasslands as well as the connectivity of the study sites significantly decreased since the 1820s. Thus, especially the huge populations of A. cynanchica seem to suffer from a kind of extinction debt today. We, therefore, assume that the slow response of A. cynanchica populations to previous habitat loss events led to decreased genetic diversity levels although the present population size is high.
Conclusions
From our study it can be concluded that habitat age seems to have no impact on genetic variation within and among populations if a sufficient number of source populations is nearby and gene flow is high. Therefore, our results support the assumption that the populations of the study species have previously been or are still connected by gene flow.
However, our study revealed a significant impact of the surrounding landscape structure and related land use patterns. Thus, we suspect that moderate grazing intensities over a long period of time may lead to increased levels of genetic diversity by intermediate levels of gene flow, while periods of overgrazing or abandonment seem to result in genetically less variable plant populations.
Finally, neither habitat quality nor population size appeared as crucial variables for genetic diversity patterns in our study. These findings provide evidence that surrounding landscape patterns are more important to preserve the genetic variation of typical calcareous grassland species than local site conditions.
Data availability
Datasets generated during this study are available from the authors upon request.
References
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego YV, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17:5177–5188
Barrett SCH, Colautti RI, Eckert CG (2008) Plant reproductive systems and evolution during biological invasion. Mol Ecol 17:373–383
Bonin A, Bellemain E, Bronken Eidesen PB, Pompanon F, Brochmann C, Taberlet P (2004) How to track and assess genotyping errors in population genetics studies. Mol Ecol 13:3261–3273
Bradshaw AD (1984) Ecological significance of genetic variation between populations. In: Dirzo RJS (ed) Perspectives on plants population ecology. Sinauer Associates Inc, Sunderland, MA, pp 213–228
Brown JH, Kodric-Brown A (1977) Turnover rates in insular biogeography: effect of immigration on extinction. Ecology 58:445–449
Burnham KP, Anderson DR (2002) Model selection and multimodel inference: a practical information-theoretic approach, 2nd edn. Springer, New York
Bylebyl K, Poschlod P, Reisch C (2008) Genetic variation of Eryngium campestre L. (Apiaceae) in Central Europe. Mol Ecol 17:3379–3388
Connell JH (1978) Diversity in tropical rain forests and coral reefs. Science 199:1302–1310
Dahlström A, Cousins SAO, Eriksson O (2006) The history (1620–2003) of land use, people and livestock, and the relationship to present plant species diversity in a rural landscape in Sweden. Environ Hist Camb 12:191–212
Dlugosch KM, Parker IM (2008) Invading populations of an ornamental shrub show rapid life history evolution despite genetic bottlenecks. Ecol Lett 11:701–709
Fischer SF, Poschlod P, Beinlich B (1996) Experimental studies on the dispersal of plants and animals on sheep in calcareous grasslands. J Appl Ecol 33:1206–1222
Grubb PJ (1977) The maintenance of species-richness in plant communities: the importance of the regeneration niche. Biol Rev 52:107–145
Hanski I (1994) Patch-occupancy dynamics in fragmented landscapes. Trends Ecol Evol 9:131–135
Helm A, Hanski I, Pärtel M (2006) Slow response of plant species richness to habitat loss and fragmentation. Ecol Lett 9:72–77
Helsen K, Jacquemyn H, Hermy M, Vandepitte K, Honnay O (2013) Rapid buildup of genetic diversity in founder populations of the gynodioecious plant species Origanum vulgare after semi-natural grassland restoration. PLoS ONE 8:e67255
Holderegger R, Kamm U, Gugerli F (2006) Adaptive vs. neutral genetic diversity: implications for landscape genetics. Landsc Ecol 21:797–807
Honnay O, Adriaens D, Coart E, Jacquemyn H, Roldan-Ruiz I (2007) Genetic diversity within and between remnant populations of the endangered calcareous grassland plant Globularia bisnagarica L. Conserv Genet 8:293–303
Jacquemyn H, Honnay O, Van Looy K, Breyne P (2006) Spatiotemporal structure of genetic variation of a spreading plant metapopulation on dynamic riverbanks along the Meuse River. Heredity (Edinb) 96:471–478
Jacquemyn H, Van MC, Brys R, Honnay O (2011) Management effects on the vegetation and soil seed bank of calcareous grasslands: an 11-year experiment. Biol Conserv 144:416–422
Jensen K, Gutekunst K (2003) Effects of litter on establishment of grassland plant species: the role of seed size and successional status. Basic Appl Ecol 4:579–587
Jooß R (2014) Schwäbische Alb (Swabian Alb) biosphere reserve. J Prot Mt Areas Res Manag 5:43–48
Karlik P, Poschlod P (2009) History or abiotic filter: which is more important in determining the species composition of calcareous grasslands? Preslia 81:321–340
Kéry M, Matthies D, Spillmann HH (2000) Reduced fecundity and offspring performance in small populations of the declining grassland plants Primula veris and Gentiana lutea. J Ecol 88:17–30
Klimek S, Richter gen. Kemmermann A, Hofmann M, Isselstein J (2007) Plant species richness and composition in managed grasslands: the relative importance of field management and environmental factors. Biol Conserv 134:559–570
Kühn I, Durka W, Klotz S (2004) BiolFlor: a new plant-trait database as a tool for plant invasion ecology. Divers Distrib 10:363–365
Kwak MM, Velterop O, van Andel J (1998) Pollen and gene flow in fragmented habitats. Appl Veg Sci 1:37–54
Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94:942–952
Leipold M, Tausch S, Hirtreiter M, Poschlod P, Reisch C (2020) Sampling for conservation genetics: how many loci and individuals are needed to determine the genetic diversity of plant populations using AFLP? Conserv Genet Resour 12:99–108
Lindborg R, Eriksson O (2004) Historical landscape connectivity affects present plant species diversity. Ecology 85:1840–1845
MacArthur RH, Wilson EO (1967) The theory of island biogeography. Princeton University Press, Princeton
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Manzano P, Malo JE (2006) Extreme long-distance seed dispersal via sheep. Front Ecol Environ 4:244–248
Mattern H, Mauk J, Kübler R (1992) Die Entwicklung der Heiden im Regierungsbezirk Stuttgart während des letzten Jahrzehnts (1980/1990). Veröffentlichungen Naturschutz Landschaftspfl Baden-württemb 67:127–135
Maurer K, Durka W, Stöcklin J (2003) Frequency of plant species in remnants of calcareous grassland and their dispersal and persistence characteristics. Basic Appl Ecol 4:307–316
May RM (1994) Biological diversity: differences between land and sea. Philos Trans R Soc London Ser B 343:105–111
McKay JK, Christian CE, Harrison S, Rice KJ (2005) “How local is local?”—a review of practical and conceptual issues in the genetics of restoration. Restor Ecol 13:432–440
Neel MC (2008) Patch connectivity and genetic diversity conservation in the federally endangered and narrowly endemic plant species Astragalus albens (Fabaceae). Biol Conserv 141:938–955
Olff H, Ritchie ME (1998) Effects of herbivores on grassland plant diversity. Trends Ecol Evol 13:261–265
Oostermeijer G, Berholz A, Poschlod P (1996) Genetical aspects of fragmented plant populations. In: Settele J, Margules C, Poschlod P, Henle K (eds) Species survival in fragmented landscapes. Kluwer Academic Publisher, Dordrecht, pp 93–101
Ouborg NJ, Vergeer P, Mix C (2006) The rough edges of the conservation genetics paradigm for plants. J Ecol 94:1233–1248
Park G (2017) Geologie Europas, 2nd edn. Wissenschaftliche Buchgesellschaft, Darmstadt
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Piqueray J, Bisteau E, Cristofoli S, Palm R, Poschlod P, Mahy G (2011) Plant species extinction debt in a temperate biodiversity hotspot: community, species and functional traits approaches. Biol Conserv 144:1619–1629
Poschlod P (2017) Geschichte der Kulturlandschaft, 2nd edn. Eugen Ulmer KG, Stuttgart
Poschlod P, WallisDeVries MF (2002) The historical and socioeconomic perspective of calcareous grasslands—lessons from the distant and recent past. Biol Conserv 104:361–376
Poschlod P, Bakker JP, Kahmen S (2005) Changing land use and its impact on biodiversity. Basic Appl Ecol 6:93–98
Poschlod P, Kiefer S, Tränkle U, Fischer S, Bonn S (1998) Plant species richness in calcareous grasslands as affected by dispersability in space and time. Appl Veg Sci 1:75–91
Poschlod P, Kleyer M, Jackel AK, Dannemann A, Tackenberg O (2003) BIOPOP—a database of plant traits and internet application for nature conservation. Folia Geobot 38:263–271
Prentice HC, Lönn M, Rosquist G, Ihse M, Kindström M (2006) Gene diversity in a fragmented population of Briza media: grassland continuity in a landscape context. J Ecol 94:87–97
Purschke O, Sykes MT, Reitalu T, Poschlod P, Prentice HC (2012) Linking landscape history and dispersal traits in grassland plant communities. Oecologia 168:773–783
R Core Team (2017) A language and environment for statistical computing. Vienna, Austria: R foundation for statistical computing
Raatikainen KM, Heikkinen RK, Luoto M (2009) Relative importance of habitat area, connectivity, management and local factors for vascular plants: spring ephemerals in boreal semi-natural grasslands. Biodivers Conserv 18:1067–1085
Reisch C (2007) Genetic structure of Saxifraga tridactylites (Saxifragaceae) from natural and man-made habitats. Conserv Genet 8:893–902
Reisch C (2008) Glacial history of Saxifraga paniculata (Saxifragaceae): molecular biogeography of a disjunct arctic-alpine species from Europe and North America. Biol J Linn Soc 93:385–398
Reisch C, Bernhardt-Römermann M (2014) The impact of study design and life history traits on genetic variation of plants determined with AFLPs. Plant Ecol 215:1493–1511
Reisch C, Schmid C, Hartig F (2018) A comparison of methods for estimating plant population size. Biodivers Conserv 27:2021–2028
Reitalu T, Johansson LJ, Sykes MT, Hall K, Prentice HC (2010) History matters: village distances, grazing and grassland species diversity. J Appl Ecol 47:1216–1224
Rico Y, Boehmer HJ, Wagner HH (2014a) Effect of rotational shepherding on demographic and genetic connectivity of calcareous grassland plants. Conserv Biol 28:467–477
Rico Y, Holderegger R, Boehmer HJ, Wagner HH (2014b) Directed dispersal by rotational shepherding supports landscape genetic connectivity in a calcareous grassland plant. Mol Ecol 23:832–842
Rogers SO, Bendich AJ (1994) Extraction of total cellular DNA from plants, algae and fungi. In: Gelvin SB, Schilperoort RA (eds) Plant molecular biology manual. Kluwer Academic Publishers, Dordrecht, pp 183–190
Rosengren F, Cronberg N, Reitalu T, Prentice HC (2013) Genetic variation in the moss Homalothecium lutescens in relation to habitat age and structure. Botany 91:431–441
Ruprecht E, Szabó A (2012) Grass litter is a natural seed trap in long-term undisturbed grassland. J Veg Sci 23:495–504
Schoen DJ, Brown AH (1991) Intraspecific variation in population gene diversity and effective population size correlates with the mating system in plants. Proc Natl Acad Sci USA 88:4494–4497
Spielman D, Brook BW, Frankham R (2004) Most species are not driven to extinction before genetic factors impact them. Proc Natl Acad Sci USA 101:15261–15264
Steffan-Dewenter I, Tscharntke T (2002) Insect communities and biotic interactions on fragmented calcareous grasslands—a mini review. Biol Conserv 104:275–284
Van Treuren R, Bas N, Goossens PJ, Jansen J, Van Soest LJ (2005) Genetic diversity in perennial ryegrass and white clover among old Dutch grasslands as compared to cultivars and nature reserves. Mol Ecol 14:39–52
Vandepitte K, Honnay O, Jacquemyn H, Roldán-Ruiz I (2010) Effects of outcrossing in fragmented populations of the primarily selfing forest herb Geum urbanum. Evol Ecol 24:1353–1364
Vos P, Hogers R, Bleeker M, Reijans M, Lee TV, Hornes M, Friters A, Pot J, Paleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Wade MJ, McCauley DE (1988) Extinction and recolonization: their effects on the genetic differentiation of local populations. Evolution (N Y) 42:995–1005
WallisDeVries MF, Poschlod P, Willems JH (2002) Challenges for the conservation of calcareous grasslands in northwestern Europe: integrating the requirements of flora and fauna. Biol Conserv 104:265–273
Willerding C, Poschlod P (2002) Does seed dispersal by sheep affect the population genetic structure of the calcareous grassland species Bromus erectus? Biol Conserv 104:329–337
Yeh FC, Yang R-C, Boyle TBJ, Ye Z-H, Mao JX (1997) POPGENE, the user-friendly shareware for population genetic analysis. Edmonton, Canada: molecular biology and biotechnology centre, University of Alberta
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Zulka KP, Abensperg-Traun M, Milasowszky N, Bieringer G, Gereben-Krenn BA, Holzinger W, Hölzler G, Rabitsch W, Reischütz A, Querner P, Sauberer N (2014) Species richness in dry grassland patches of eastern Austria: a multi-taxon study on the role of local, landscape and habitat quality variables. Agric Ecosyst Environ 182:25–36
Acknowledgements
The authors thank Petra Schitko for assistance in the lab and Sven Rubanschi for support in statistical approaches. We further thank Cornelia Straubinger providing the vegetation cover data. The study was financially supported by the Federal Agency for Agriculture and Food (BLE).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
CR and PP conceived and designed the study. TAL and EP collected the data and performed the genetic analyses. TAL analysed the data and wrote the paper. EP, PP, and CR revised the article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lehmair, T.A., Pagel, E., Poschlod, P. et al. Surrounding landscape structures, rather than habitat age, drive genetic variation of typical calcareous grassland plant species. Landscape Ecol 35, 2881–2893 (2020). https://doi.org/10.1007/s10980-020-01120-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-020-01120-7