Abstract
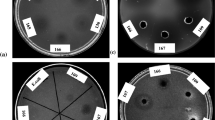
Biological control to prevent fungal plant diseases offers an alternative approach to facilitate sustainable agriculture. Since the chitin in fungal cell walls is a target for biocontrol agents, chitinases are one of the important antifungal molecules. In this study, the aim was to investigate a new chitinase isolated from a fluvial soil bacterium and to show the antifungal activity of the characterized chitinase by comparing the three common methods. The bacterium with the highest chitinase activity was identified as Aeromonas sp. by 16 S rRNA sequence analysis. Following the determination of the optimum enzyme production time, the enzyme was partially purified, and the physicochemical parameters of the enzyme were investigated. In the antifungal studies, direct Aeromonas sp. BHC02 cells or partially purified chitinase were used. As a result, in the first method in which the Aeromonas sp. BHC02 cells were spread on the surface of petri dishes, no zone formation was observed around the test fungi spotted on the surface. However, zone formation was observed in the methods in which the antifungal activity was investigated using the partially purified chitinase enzyme. For example, in the second method, the enzyme was spread on the surface of PDA, and zone formation was observed only around Penicillum species among the test fungi spotted on the surface. In the third method, in which the necessary time was given for the formation of mycelium of the test fungi, it was observed that the growth of Fusarium solani, Alternaria alternata and Botrytis cinerea was inhibited by the partially purified chitinase. This study concludes that the results of the antifungal activities depend on the method used and all fungal chitins cannot be degraded with one strain’s chitinase. Depending on the variety of chitin, some fungi can be more resistant.






Similar content being viewed by others
Abbreviations
- SDS-PAGE:
-
Sodium dodecyl sulfate- polyacrylamide gel electrophoresis
- NAG:
-
N-acetylglucosamine
- CHD:
-
Chitinase-detection
- NCBI:
-
National Centre for Biotechnology Information
- DNS:
-
3,5-Dinitrosalicylic acid
- BSA:
-
Bovine serum albumin
- V max :
-
Maximum rate of reaction
- K M :
-
Michaelis Constant
- PCR:
-
Polymerase chain reaction
- PDA:
-
Potato dextrose agar
References
Fisher MC, Gurr SJ, Cuomo CA, Blehert DS, Jin H, Stukenbrock EH, Stajich JE, Kahmann R, Boone C, Denning DW, GowNAR,Klein BS, Kronstad JW, Sheppard DC, Taylor JW, Wright GD, Heitman J, Casadevall A, Cowen LE (2020) Threats posed by the fungal kingdom to humans, wildlife, and agriculture. mBio 11:e00449–e00420
Cd D, Lb V, Ma M, Md B (2021) Extracellular antifungal activity of chitinase-producing Bacteria isolated from Guano of Insectivorous Bats. Curr Microbiol 78:2787–2798
Elieh Ali KD, Sharma L, Dela Cruz CS (2018) Chitin and its Effects on Inflammatory and Immune responses. Clin Rev Allerg Immunol 54:213–223
Basu D, Chaudhuri A (2013) Purification and characterization of chitinase from Thermophilic Staphylococcus sp. Internatıonal J Envıronmental Scıences 4:4
Zarei M, Aminzadeh S, Zolgharnein H, Safahieh A, Daliri M, Noghabi KA, Ghoroghi A, Motallebi A (2011) Characterization of a chitinase with antifungal activity from a native Serratia marcescens B4A. Braz J Microbiol 42(3):1017–1029
Thakur N, Nath AK, Chauhan A, Gupta R (2021) Purification, characterization, and antifungal activity of Bacillus cereus strain NK91 chitinase from rhizospheric soil samples of Himachal Pradesh, India. Biotechnol. Appl. Biochem
Ekundayo FO, Folorunsho AE, Ibisanmi TA (2022) Antifungal activity of chitinase produced by Streptomyces species isolated from grassland soils in Futa Area, Akure. Bull Natl Res Cent 46:95
Hueso-Rodríguez JA, Jimeno ML, Rodríguez B, Savona G, Bruno M (1983) Abietane diterpenoids from the root of Salvia phlomoides. Phytochemistry 22(9):2005–2009
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences (USA) 101:11030–11035
Toharisman A, Suhartono MT, Spindler-Barth M, Hwang JK, Pyun YR (2005) Purification and characterization of a thermostable chitinase from Bacillus licheniformis Mb-2. World J Microbiol Biotechnol 21:733–738
Wang D, Li A, Han H, Liu T, Yang Q (2018) A potent chitinase from Bacillus subtilis for the efficient bioconversion of chitin-containing wastes. Int J Biol Macromol 116:863–868
Wang D, Shih L, Liang W, Wangn CH (2002) Purification and Characterization of Two Antifungal Chitinases Extracellularly Produced by Bacillus amyloliquefaciens V656
Stoscheck CM (1990) Quantitation of protein. Methods Enzymol 182:50–68
Smith BJ (1994) SDS polyacrylamide gel electrophoresis of proteins. Basic protein and peptide protocols 23–34
Ernst O, Zor T (2010) Linearization of the Bradford Protein Assay. J. Vis. Exp 38:1918
Yuli PE, Suhartono TM, Rukayadi Y, Hwang JK, Pyun YR (2004) Characteristics of thermostable chitinase enzymes from the indonesian Bacillus sp. Enzym Microb Technol 35:147–115
Choi B, Rempala GA, Kim JK (2017) Beyond the Michaelis-Menten equation: Accurate and efficient estimation of enzyme kinetic parameters. Sci Rep 7(1):1–11
Ozyigit C (2020) Evaluation of effectiveness of propolis extracts, collected from different regions of Turkey, against mold agents Master Thesis, Department of Crop Protection, Tokat Gaziosmanpasa University
Vaidya RJ, Shah IM, Vyas PR, Chhatpar HS (2001) Production of chitinase and its optimization from a novel isolate Alcaligenes xylosoxydans: potential in antifungal biocontrol. World J Microbiol Biotechnol 17:691–696
Kirubakaran SI, Sakthivel N (2007) Cloning and overexpression of antifungal barley chitinase gene in Escherichia coli. Protein Exp Purific 52:159–166
Sarker KN, Das Mohapatra PK, Dutta S (2019) Use of low-cost natural resources for enhanced chitinase production and optimization using CCD and RSM: a new initiative for bio-control of plant pathogen. Indian Phytopathol 72(2):281–300
Smith AC, Hussey M (2005) Gram stain protocols. Am Soc Microbiol 1:14
English BP, Min W, Van OAM, Lee KT, Luo G, Sun H, Xie XS (2006) Ever-fluctuating single enzyme molecules: Michaelis-Menten equation revisited. Nat Chem Biol 2(2):87–94
Vijayanandraj VR, Nagendra Prasad D, Mohan N, Gunasekaran M (2006) Effect of ozone on Aspergillus niger causing black rot disease in onion. Ozone: Sci Eng 28(5):347–350
Karima HEH, Nadia GE (2012) In vitro study on Fusarium solani and Rhizoctonia solani isolates causing the damping off and root rot diseases in tomatoes. Nat Sci 10(11):16–25
Zhu L, Ni W, Liu S, Cai B, Xing H, Wang S (2017) Transcriptomics analysis of apple leaves in response to Alternaria alternata apple pathotype infection. Front Plant Sci 8:22
Petrasch S, Knapp SJ, Van Kan JA, Blanco-Ulate B (2019) Grey mould of strawberry, a devastating disease caused by the ubiquitous necrotrophic fungal pathogen Botrytis cinerea. Mol Plant Pathol 20(6):877–892
Latorre BA, Rioja ME, Wilcox WF (2001) Phytophthora species associated with crown and root rot of apple in Chile. Plant Dis 85(6):603–606
Bajpai A, Agnihotri R, Prakash A, Johri BN (2022) Biosurfactant from Bacillus sp. A5F reduces Disease incidence of Sclerotinia sclerotiorum in soybean crop. Curr Microbiol 79(7):1–11
Donegan KK, Schaller DL, Stone JK, Ganio LM, Reed G, Hamm PB, Seidler RJ (1996) Microbial populations, fungal species diversity and plant pathogen levels in field plots of potato plants expressing the Bacillus thuringiensis var. Tenebrionis endotoxin. Transgenic Res 5(1):25–35
Lim YL, Ee R, Yin WF, Chan KG (2014) Quorum sensing activity of Aeromonas caviae strain YL12, a bacterium isolated from compost. Sensors 14(4):7026–7040
Mehmood MA, Gai Y, Zhuang Q, Wang F, Xiao X, Wang F (2010) Aeromonas caviae CB101 contains four chitinases encoded by a single gene chi1. Mol Biotechnol 44(3):213–220
Cardozo FA, Gonzalez JM, Feitosa VA, Pessoa A, Rivera ING (2017) Bioconversion of α-chitin into N-acetyl-glucosamine using chitinases produced by marine-derived Aeromonas caviae isolates. World J Microbiol Biotechnol 33(11):1–11
Saima MK, Roohi IZA (2013) Isolation of novel chitinolytic bacteria and production optimization of extracellular chitinase. J Genetic Eng Biotechnol 11(1):39–46
Stumpf AK, Vortmann M, Dirks-Hofmeister ME, Moerschbacher BM, Philipp B (2019) Identification of a novel chitinase from Aeromonas hydrophila AH-1 N for the degradation of chitin within fungal mycelium. FEMS Microbiol Lett 366(1):fny294
Zhang Y, Zhou Z, Liu Y, Cao Y, He S, Huo F, Qin C, Yao B, Ringo E (2014) High-yield production of a chitinase from Aeromonas veronii B565 as a potential feed supplement for warm-water aquaculture. Appl Microbiol Biotechnology 98:1651–1662
Chang WT, Chen CS, Wang SL (2003) An antifungal chitinase produced by Bacillus cereus with shrimp and crab Shell Powder as a Carbon source. Curr Microbiol 47:0102–0108
Pandya U, Saraf M (2015) Purification and characterization of antifungal chitinase from Bacillus safensis MBCU6 and its application for production of chito-oligosaccharides. Biologia 70:863–868
Pentekhina I, Hattori T, Tran DM, Shima M, Watanabe T, Sugimoto H, Suzuki K (2020) Chitinase system of Aeromonas salmonicida, and characterization of enzymes involved in chitin degradation. Biosci Biotechnol Biochem 84:1347–6947
Zhang W, Ma J, Yan Q, Jiang Z, Yang S (2021) Biochemical characterization of a novel acidic chitinase with antifungal activity from Paenibacillus xylanexedens Z2–4. Int J Biol Macromol 182:1528–1536
Okay S, Ozdal M, Kurbanoglu EB (2013) Characterization, antifungal activity, and cell immobilization of a chitinase from Serratia marcescens MO-1. Turkish J Biology 37:639–644
Bhushan B (1998) Isolation, purification, characterization and scale up production of a thermostable chitinase from an alkalophilic microorganism. Ph.D. thesis, Department of Microbiology Chandigarh: Punjab University
Author information
Authors and Affiliations
Contributions
Bilge Hilal CADIRCI established the hypothesis of the research, wrote and edit the manuscript. Gulesme Yilmaz did the experiments and prepared almost most of the figures.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cadirci, B.H., Yilmaz, G. Comparison of in vitro Antifungal Activity Methods Using Extract of Chitinase-producing Aeromonas sp. BHC02. Protein J 42, 125–134 (2023). https://doi.org/10.1007/s10930-023-10098-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-023-10098-5