Abstract
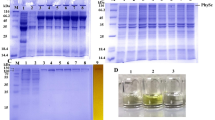
Regulator of the H+-ATPase of the vacuolar and endosomal membranes (RAVE) is essential for the reversible assembly of H+-ATPase. RAVE primarily consists of three subunits: Rav1p, Rav2p and Skp1p. To characterize these subunits, in this study, four strains derived from Saccharomyces cerevisiae BY4742 were constructed with a FLAG tag on the Rav1p and Rav2p subunits. Then, the corresponding RAVE containing complex was isolated by affinity purification. Western blot and MALDI-TOF mass spectrometry analyses showed that the RAVE complex contains not only the known V1-ATPase subunits (Vma1p and Vma2p) but also a newly found Leu1p that interacts with the RAVE subunit. Furthermore, we constructed rav1−/rav2−/vma2−/leu1-deficient recombinants by fusion PCR and homologous recombination and demonstrated that leu1 is indispensable in adjusting the microbial cell to adverse environments and that the function is similar to that of rav1/rav2 but significantly differs from that of vma2. Leu1p probably plays an important role in RAVE regulation of V-ATPase activity in conjunction with RAVE.







Similar content being viewed by others
Abbreviations
- G418:
-
Geneticin
- HRP:
-
Horseradish peroxidase
- IgG:
-
Immunoglobulin G
- PVDF:
-
Polyvinylidene difluoride
- TBS:
-
Tris buffered saline
- TBS/T:
-
TBS, 0.1% Tween-20
- BSA:
-
Bovine serum albumin
- DAB:
-
Diaminobenzidine
- TFA:
-
Tallow fatty acid
- ACN:
-
Acetonitrile
References
Baars TL, Petri S, Peters C, Mayer A (2007) Role of the V-ATPase in regulation of the vacuolar fission-fusion equilibrium. Mol Biol Cell 18(10):3873–3882
Boettner DR, Chi RJ, Lemmon SK (2012) Lessons from yeast for clathrin-mediated endocytosis. Nat Cell Biol 14(1):2–10
Dettmer J, Hong-Hermesdorf A, Stierhof YD, Schumacher K (2006) Vacuolar H+-ATPase activity is required for endocytic and secretory trafficking in Arabidopsis. Plant Cell 18(3):715–730
Forgac M (2007) Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat Rev Mol Cell Biol 8(11):917–929
Galan JM, Wiederkehr A, Seol JH et al (2001) Skp1p and the F-box protein Rcy1p form a non-SCF complex involved in recycling of the SNARE Snc1p in yeast. Mol Cell Biol 21(9):3105–3117
Graham L, Flannery A, Stevens T (2003) Structure and assembly of the yeast V-ATPase. J Bioenerg Biomembr 35(4):301–312. doi:10.1023/A:1025772730586
Haber JE (2012) Mating-type gene switching in Saccharomyces cerevisiae. Trends Genet 191(32):561–599
Hunt SR, Hernandez R, Brown DT (2010) Role of the vacuolar-ATPase in Sindbis virus infection. J Virol 85(3):1257–1266. doi:10.1128/jvi.01864-10
Inoue T, Wikens S, Forgac M (2003) Subunit structure, function, and arrangement in the yeast and coated vesicle V-TTPases. J Bioenerg Biomembr 35(4):291–299
Jefferies KC, Cipriano DJ, Forgac M (2008) Function, structure and regulation of the vacuolar (H+)-ATPases. Arch Biochem Biophys 476(1):33–42
Jacq C, AltMorbe J, Andre B et al (1997) The nucleotide sequence of Saccharomyces cerevisiae chromosome IV. Nature 387(6632):75–78
Kane PA (2006) The where, when, and how of organelle acidification by the yeast vacuolar H+-ATPase. Microbiol Mol Biol Rev 70(1):177–191. doi:10.1128/mmbr.70.1.177-191.2006
Kane PM (2012) Targeting reversible disassembly as a mechanism of controlling V-ATPase activity. Curr Protein Pept Sci 13(2):117–123
Lloyd SJ, Lauble H, Prasad GS, Stout CD (1999) The mechanism of aconitase: 1.8 angstrom resolution crystal structure of the S642A:citrate complex. Protein Sci 8(12):2655–2662
Marchler-Bauer A, Zheng C, Chitsaz F et al (2013) CDD: conserved domains and protein three-dimensional structure. Nucleic Acids Res 41(D1):348–352
Mario PS, Abel GG, Maria DRL, Pilar GV (2009) Role of V-ATPases in solid tumors: importance of the subunit C (review). Int J Oncol 34(6):1513–1520. doi:10.3892/ijo_00000280
Morel N (2003) Neurotransmitter release: the dark side of the vacuolar-H(+)ATPase. Biol Cell 95(7):453–457
Oka T, Futai M (2000) Requirement of V-ATPase for ovulation and embryogenesis in Caenorhabditis elegans. J Biol Chem 275(38):29556–29561. doi:10.1074/jbc.M002756200
Oluwatosin YE, Kane PM (1998) Mutations in the yeast KEX2 gene cause a Vma—like phenotype: a possible role for the Kex2 endoprotease in vacuolar acidification. Mol Cell Biol 18(3):1534–1543
Sadowski M, Mawson A, Baker R, Sarcevic B (2007) Cdc34 C-terminal tail phosphorylation regulates Skp1/cullin/F-box (SCF)-mediated ubiquitination and cell cycle progression. Biochem J 405:569–581
Smardon AM, Kane PM (2007) RAVE is essential for the efficient assembly of the C subunit with the vacuolar H+−ATPase. J Biol Chem 282(36):26185–26194
Smardon AM, Tarsio M, Kane PM (2002) The RAVE complex is essential for stable assembly of the yeast V-ATPase. J Biol Chem 277(16):13831–13839
Sennoune SR, Bakunts K, Martinez GM, Chua-Tuan JK, Kebir Y, Attaya MN, Martinez-Zaguilan R (2004) Vacuolar H+-ATPase in human breast cancer cells with distinct metastatic potential: distribution and functional activity. Am J Physiol Cell Physiol 286(6):1443–1452
Seol JH, Shevchenko A, Shevchenko A, Deshaies RJ (2001) Skp1 forms multiple protein complexes, including RAVE, a regulator of V-ATPase assembly. Nat Cell Biol 3(4):384–391
Solutions S. Western Immunoblotting — Standard Protocol 3–8
Toei M, Saum R, Forgac M (2010) Regulation and isoform function of the V-ATPases. Biochemistry 49(23):4715–4723
Wang YL, Addess KJ, Chen J et al (2007) MMDB: annotating protein sequences with Entrez's 3D-structure database. Nucleic Acids Res 35:298–300
Wendland J (2003) PCR-based methods facilitate targeted gene manipulations and cloning procedures. Curr Genet 44(3):115–123
Williamson WR, Hiesinger PR (2010) On the role of V-ATPase V0a1-dependent degradation in Alzheimer disease. Commun Integr Biol 3(6):604–607
Xu XL, Song Y, Li Y, Chang JF, Zhang H, An L (2010) The tandem affinity purification method: an efficient system for protein complex purification and protein interaction identification. Protein Expr Purif 72(2):149–156. doi:10.1016/j.pep.2010.04.009
Yamashiro CT, Kane PM, Wolczyk DF, Preston RA, Stevens TH (1990) Role of vacuolar acidification in protein sorting and zymogen activation: a genetic analysis of the yeast vacuolar proton-translocating ATPase. Mol Cell Biol 10(7):3737–3749
Zhang ZY, Charsky C, Kane PM, Wilkens S (2003) Yeast V1-ATPase-affinity purification and structural features by electron microscopy. J Biol Chem 278(47):47299–47306
Zhang ZY, Zheng Y, Mazon H, Milgrom E, Kitagawa N, Kish-Trier E, Heck AJR, Kane PM, Wilkens S (2008) Structure of the yeast vacuolar ATPase. J Biol Chem 283(51):35983–35995
Acknowledgements
The work was supported by the National Natural Science Foundation of China (30970058; 21176106) and Natural Science Foundation of Jiangsu Province (BK2012554), together with the Project Funded by China Postdoctoral Science Foundation (2015 M571666) and Fundamental Research Funds for the Central Universities (JUSRP51635B). Authors also would like to give the thanks to the Priority Academic Program Development of Jiangsu Higher Education Institutions, the 111 Project (No. 111-2-06), and the Jiangsu province “Collaborative Innovation Center for Advanced Industrial Fermentation” industry development program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOCX 1194 kb)
Rights and permissions
About this article
Cite this article
Zhang, Z., Wang, X., Gao, T. et al. Characterization of the complex involved in regulating V-ATPase activity of the vacuolar and endosomal membrane. J Bioenerg Biomembr 49, 347–355 (2017). https://doi.org/10.1007/s10863-017-9712-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-017-9712-1