Abstract
Bacterial communities present in the host digestive tract are important for the breakdown and absorption of nutrients required by the host. Changes in diet and the environment are major factors affecting the composition and diversity of the fecal microbiome. In addition to changes in ambient temperature and rainfall, primates living in seasonal temperate environments also need to adapt to seasonal changes in food resource quantity and quality. However, there is a lack of information about the fecal microbiome in African strepsirrhines relative to other primate taxa. We examined the effects of seasonal dietary and environmental changes on fecal microbial alpha diversity and composition in wild greater thick-tailed galagos (Otolemur crassicaudatus) at Lajuma Research Centre, South Africa. We collected fecal samples and assessed food availability and weather in summer and winter across 1 year and used 16S rRNA next-generation sequencing to characterise the fecal microbiome of 49 animals. We found significant increases in rainfall, ambient temperature, and food availability in summer compared with winter. However, we found no significant changes in body mass or in the overall diversity of bacterial species present in fecal samples between the two seasons. We found significant decreases in the abundance of certain bacterial families in winter, suggesting a change in diet. Our findings suggest that greater thick-tailed galagos can find food resources to maintain their body mass throughout the year. Our insights into the seasonal fecal microbiome of greater thick-tailed galagos add to the growing knowledge and understanding of fecal microbiomes in primates and how they help primates cope with changes to their environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Primates have adapted to living in a variety of different habitats, such as forests, mountains, and savannahs, and show an array of dietary preferences, such as insectivory, folivory, gummivory, and frugivory (MacKinnon & MacKinnon, 1980). Primates adjust their activity patterns and feeding behaviour in response to changes in their environment (Strier, 2009; Reyna-Hurtado et al., 2018). These developments may include changes to their social structure or habitat use (Chevalier et al., 2015). Changes also may occur in the gut microbiome, which consists of a symbiotic community of microbial organisms residing in the gut of the host individual (Amato et al., 2014). Bacterial communities present in the host digestive tract are linked to changes in the host diet, health, and the extrinsic environment (David et al., 2014; Bjork et al., 2019; Orkin et al., 2019). These communities are important for the breakdown and absorption of nutrients required by the host (Cabana et al., 2018). For example, carbohydrates are fermented and broken down into branched- and short-chain fatty acids (SCFAs; Scheppach, 1994; Nicholson et al., 2012), which can be absorbed in the gut and are available as an energy source for the host (Hildebrandt et al., 2009; De Filippo et al., 2010; Wu et al., 2011; Claesson et al., 2012). Carbohydrate fermentation is a critical function of the gut in the catabolism of these important nutrient sources in the metabolic homeostasis of the host (Scheppach, 1994; Nicholson et al., 2012; Yuan et al., 2020).
Insects and fruit are common food preferences for primate species as they are high in carbohydrates and nutrients (O’Malley & Power, 2012). Gum exudate is low in nutrients and rare (Irani & Khaled, 2020) but is a common dietary component in many primate species (Caton et al, 2000). Gummivory may be beneficial for an insectivorous primate, providing calcium to complement the phosphorous taken from insects (Génin et al., 2010; Bearder & Martin, 1980). Both gum exudates and insect exoskeletons are comprised of insoluble β-linked complex polysaccharides, which the mammalian gut is incapable of digesting without the assistance of specialized gut microbes and fermentation (O’Malley & Power, 2012). The presence of gut microbes in the gut allows primates to digest and absorb the carbohydrates found in gum and insects exoskeletons (O’Malley & Power, 2012). As a result, fermentation and the digestive abilities of gut microbes are required in the gut for the primate to absorb and use these carbohydrates (Caton et al., 2000; O’Malley & Power, 2012). However, the gut microbiome is sensitive to both the diet of an organism, as well as the prevailing environmental conditions (Xu & Knight, 2015; Kers et al., 2018). Thus, any change in either component will result in a microbiome response to assist an individual in maintaining digestive health (Manor et al., 2020).
Most studies of the influence of seasonal shifts in climate and resource use on the gut microbial composition of primates have been conducted on haplorrhines (Amato et al., 2014; Sun et al., 2016). Very few studies have examined the effect of changing environment on the gut microbiome of African strepsirrhines. Filling this taxonomic gap is important in helping us understand the nutritional and metabolic requirements and physiology of primates.
Two nocturnal strepsirrhine primates are found in South Africa, the southern lesser galago (G. moholi) and the thick-tailed greater galago (Otolemur crassicaudatus). While research on G. moholi has increased considerably over the past decade (Nowack et al., 2010, 2013; Scheun et al., 2015, 2016), including a study that found significant differences in seasonal bacterial profiles of the fecal microbiome (Long, 2018), very little has been attempted on thick-tailed greater galagos. These galagos inhabit highly seasonal environments in South Africa (Limpopo, Mpumalanga, and KwaZulu-Natal provinces; Hill, 1953), where changes in rainfall and temperature across the four seasons (summer, autumn, winter, spring) result in large changes in resource availability (Hill, 1953). They feed primarily on insects in summer and fruits in winter with gum-feeding occurring throughout the year (Bearder, 1974; Harcourt, 1986; Happold & Happold, 1992). Both insects and fruits provide a range of nutrients, such as phosphorous, sugars, and fats (Scheun et al., 2014; Rothman et al., 2014; O’Malley & Power, 2012). The foregut of thick-tailed greater galagos has evolved to allow fermentation of complex polysaccharides (Cabana et al., 2018). They show morphological and gut adaptations for both accessing and digesting gums and are designated as obligate gum feeders (Smith, 2010; Cabana et al., 2018).
To contribute to our understanding of the effect of seasonality on the gut microbiome in strepsirrhines, we aimed to identify the main microbial taxa found in the fecal microbiome of thick-tailed greater galagos inhabiting an isolated temperate environment in two contrasting seasons. Under the hypothesis that the fecal microbiome is affected by changes in diet, we predict a decrease in microbial taxonomy and relative abundance during winter when a considerable shift in diet occurs in comparison to summer. We also explore potential influences on the fecal microbiome, including seasonal changes in food availability, rainfall, and ambient temperature.
Methods
Study Site
Lajuma Research Centre (23°02’16.2”S, 29°26’35.4”E) is situated in the Soutpansberg Mountains in the Limpopo Province in the North-Eastern region of South Africa. It has a highly seasonal, temperate environment, comprised of montane forests and savannah woodland habitats (Van Wyk & Smith, 2001; Von Maltitz et al., 2003). Summer is from October to March, and winter is from late April to September. Temperatures fluctuate from 0 °C to 22 °C in the dry, winter months when overnight frost is common in more open areas and 12 °C to 36 °C in the wet, summer months (Mostert, 2010; Durowoju et al., 2019). While rainfall generally peaks in January and February, almost no rainfall is seen in June and July (median = 0 insects per capture,). The vegetation grows from October to April, whereas the less favourable season for vegetation growth is from May to September. Five of the six species of primates found in South Africa live sympatrically at this location while several natural predators exist for galagos (Cuozzo et al., 2021; Allan et al., 2022).
Animal Captures and Faecal Sampling
We trapped thick-tailed greater galagos in a 3-km2 area at Lajuma along transects from a northwest to southeast direction in June 2017, December 2017/January 2018, March 2018, and May/June 2018. We aimed to collect samples from at least 20 adults (10 males and 10 females; Table A1) in each trapping period. Each night, we focused on one area of the trapping range of approximately 50–100 m2. In each sampling area, we set between four and nine traps, depending on the density of galagos and the proximity of the traps to one another. A licenced veterinarian was present during all animal capture and sampling events. We set Havahart® traps (Woodstream Corporation Inc., Lancaster, PA) baited with small amounts of peanut butter and banana before sunset (between 17h00 and 18h30, depending on the season) and checked them at sunrise (between 04h30 and 06h00, depending on the season). To digest gum, galagos use gut fermentation, a process that requires prolonged gut retention times (Caton et al., 2000). With a gut retention time of >10 hours (Long et al., 2021) and their preference for being most active late at night, it is unlikely that the samples we collected from trapped individuals changed due to the ingestion of the bait that we set out. We cleaned empty traps with a solution of 99% rubbing alcohol and water and closed them until the next trapping session. If an animal was present in the trap, we used a capture bag to collect it, identified it by using a microchip transponder reader (ID100 Trojan, EURO I.D., Weilerswist, Germany), then placed it into a pet travel box and translocated it to the field laboratory. We collected fecal samples for seed availability assessment from the trap once the animal was safely placed in the travel box.
Once at the field lab, we anesthetized the animal with an intramuscular injection of zolazepam and tiletamine (Zoletil®, Virbac, South Africa). We maintained the level of anesthesia with inhaled isoflurane in oxygen, administered intermittently via a face mask. Once anesthetized, we weighed each animal using a table-top scale. If we did not detect a microchip, we inserted a transponder microchip subcutaneously, and recorded the transponder identity number as the animal’s ID. The vet checked reproductive status for females (pregnant, lactating, not pregnant). We recorded male reproductive status as either mating (June to August) or not mating (summer months) depending on the time of year. We inserted a FLOQ® fecal swab (FLOQSwabs®, Copan Group, Brescia, Italy) into the rectum to retrieve a sufficient volume of fecal matter to discolour the swab. Once removed, we cut off the head of the swab and placed it immediately into a 2-ml sterile tube, then stored it immediately in a −4 °C freezer. Once transported back to the University of Pretoria, approximately one week after the start of sampling, we stored the samples in a −80 °C freezer until DNA isolation.
We released captured animals where we caught them, on the same day we removed from the traps, as soon as they were completely conscious, and the veterinarian was satisfied with their health condition.
Seasonal Food Availability
We collected insect, gum, and seed samples to reflect the annual food availability for galagos. We collected insect and gum data from the environment and ascertained seed data from ingested fruit. We collected insects from September 2017 to June 2018. We set up two light traps at the field site before sunset approximately 15-m apart on consecutive days before and after trapping sessions in areas frequented by galagos. We placed traps before sunset. We set one light trap on the ground to estimate the abundance of ground dwelling insects, and the second 2 m above the ground to assess the abundance of flying insects. We set each light above a collecting chamber with a hole in the lid preventing most insects from escaping from the trap. We checked traps every morning at sunrise and sprayed all trapped insects with 70% ethanol, counted, and, where possible, identified them. We stored all specimens in 90% ethanol until we analysed food availability. For the analysis, we took the cumulative value of insects from both light traps.
We collected gum exudate samples monthly from 50 trees (10 trees per sampling section). We selected trees after we identified trees frequented by galagos or with gum exudates (personal observations by J.B. Millette and C. Long). We scraped gum exudates off the tree bark using a multitool and placed the sample in a sample bag. We stored these samples in a −20 °C freezer until the food availability analysis.
We collected all fecal matter excreted by trapped individuals from July 2017 to June 2018 (winter: N = 12, summer: N = 25, winter 2: N = 32). We used a subset of each fecal sample to determine the presence and number of seeds. We dried the feces and extracted seeds by using tweezers. We took each seed to represent one fruit except for multiple Ficus seeds, which we identified as one fruit per fecal sample (Long et al., 2021). We counted the seeds retrieved and identified them to genus if possible.
Seasonal Weather Differences
We captured rainfall and ambient temperature data by using the on-site weather station (Davis Instruments sensor suite linked via wireless to a Davis Pro 2™ console, Measurement and Console Systems CC, Cape Town, South Africa), where conditions were analogous to those experienced by the sampled individuals. The Ndlovu Node of the Northeastern Mountain Observatories project of the South African Environmental Observation Network (SAEON) made these data available. We used data collected every hour to calculate the mean (± standard deviation; SD) daily ambient temperature (°C) and daily cumulative rainfall (mm) total from June 2017 to August 2017 (winter 1), November 2017 to February 2018 (summer), and May to July 2018 (winter 2).
DNA Isolation and Next-Generation Sequencing
We extracted the total fecal bacterial DNA from 49 individuals (males: N = 28; females: N = 21) using the DNeasy® PowerSoil® Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. We assessed the quality of DNA using a QuBit 2.0 Fluorometer (Invitrogen, Carlsbad, CA). Inqaba Biotec Industries (Pty) Ltd (Muckleneuk, Pretoria) conducted PCR amplification, library preparation, next-generation sequencing, and data process and analysis. They created bacterial gene libraries by using the PacBio Full-Length 16S Amplification, SMRTbell® Library Preparation and Sequencing protocol. Briefly, the workflow implements two PCR rounds, the first with universal primer-tailed 16S primers (27F; 1492R) and the second with PacBio Barcoded Universal Primers. They performed the first round of amplification with an initial denaturation at 95 °C for 30 s, 23 cycles of denaturing (95 °C for 30 s), annealing (55 °C for 30 s), and elongation (72 °C for 60 s). At each PCR step they generated amplicons using a KAPA HiFi PCR kit (Kapa Biosystems, Wilmington, MA) and quantified them on a Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA). The second round of amplification used the same thermocycler instructions. They prepared libraries by using SMRTbell Template Prep kit (Pacific Biosciences, Menlo Park, CA) and purified them using AMPure® PB beads (Pacific Biosciences).
We sequenced the prepared libraries on the PacBio Sequel system (PacBio, http://www.pacbio.com). We processed the raw subreads through the SMRTlink (v7) Circular Consensus Sequences (CCS) algorithm to produce highly accurate reads (>QV40). The minimum sequence read length was set at >150 bp. We identified and removed the chimeric sequences and, finally, generated FASTQ files through vsearch (https://github.com/torognes/vsearch). We identified taxa using QIIME2 (Bolyen et al., 2019). We assigned the taxonomy of each gene sequence by using the Silva SSU NR database (v132.1; Quast et al., 2013), picking operational taxonomic units (OTUs) using a 97% similarity cutoff. We normalized the data by reducing the sequence reads to 10,000 per sample.
Statistical Analyses
We performed statistical analyses in R (R Core Team, 2020). We set significance at p < 0.05, except where we corrected p-values for multiple testing by using the Bonferroni correction. We report means plus SD for parametric data and medians plus interquartile ranges for nonparametric data.
Seasonal Analysis
We split the data for food availability into the three sampling seasons: June 2017 to August 2017 (winter 1); November 2017 to February 2018 (summer); and May to July 2018 (winter 2). We assessed seasonal differences in insect count, gum mass, and seed count data using Kruskal-Wallis tests with a chi-square distribution as the data were not normally distributed. We used Dunn’s post-hoc tests to test for differences between seasons. For insects, we analysed the cumulative number of insects from both light traps per month. For gum mass, we analysed the sum of the gum mass from all trees per month. We also tested for seasonal differences (between winter 1, summer, and winter 2) in body mass for both sexes by using Kruskal-Wallis tests with chi-squared distribution.
If the Kruskal-Wallis analysis found seasonal differences, we used post-hoc Dunn’s tests to test for differences between the three seasons. Before analysis, we used the Levene’s test to determine that variances were significantly different for both rainfall and ambient temperature. If seasonal differences were found after the Kruskal-Wallis analysis, we performed post hoc Dunn’s tests to determine which seasons experienced the highest ambient temperature and rainfall.
We compared relative abundances of the phyla present across the three seasons (winter 1, summer, winter 2) using a Kruskal-Wallis test and post hoc Dunn’s tests. We assessed the seasonal presence of the most predominant microbial families by performing Kruskal-Wallis tests with post hoc Dunn’s tests. We created Venn diagrams by using Venny 2.1 (Olivieros, 2015) to illustrate the unique and shared bacterial families between seasons. As there was little variation between the bacterial families present between winter 1 and winter 2, we combined the data. We used rarefaction curves to evaluate the depth of sequencing (Fig. A1). We used Analysis of Similarities (ANOSIM) to compare gut microbial abundance (the total number of bacteria present) between seasons using the anosim function in the “vegan” package. We tested the effects of sex and seasonal variation (winter 1, summer, winter 2) on the microbial composition using a permutational multivariate analysis of variance (PERMANOVA) using the adonis2 function of the “vegan” package. We set the number of permutations at 9999. We used a Mann-Whitney U test to test for seasonal differences in bacterial diversity (the number of bacterial taxa present). We used the Kruskal-Wallis test to assess differences in the relative abundance (log10 transformed) of genera between winter 1, summer, and winter 2 and Dunn’s test to identify which seasons significantly differed.
We used linear mixed models (LMMs) to identify factors contributing to gut microbial composition. The fixed parameters were daily insect count, gum volume, seed count, rainfall, ambient temperature, sex, and reproductive status. We set Animal ID as a random variable to control for nonindependence of samples from the same animal. We ran the linear model by using the lmer function in the “lme4” package (Bates, 2010). Before modelling, we normalized the dataset by using the preProcess function. After assessing the variance inflation factor (VIF > 3; Zuur & Ieno, 2016), we removed ambient temperature as a fixed variable owing to collinearity. Finally, we investigated Spearman’s correlations between the most abundant phyla and specific environmental factors to assess their effects on the fecal microbial community composition within galagos.
Ethical Note
We received ethical clearance to conduct the study from the Research and Ethical Sciences Committee at the South African National Biodiversity Institute’s (SANBI) National Zoological Gardens (NZG; Project 18/26), the Animal Ethics Committee of the University of Pretoria (Project V037-17), and Animal Protocol Approval 181 Assurance D16-00388 from the IACUC office of the University of Colorado, which reviewed our research protocols. The authors declare there is no conflict of interest.
Data Availability Statement
Data can be viewed or uploaded from the National Centre of Biotechnology Information https://ncbi.nlm.nih.gov by using the reference SUB13752167.
Results
Seasonal Food Availability
We captured 2414 insects from June 2017 to June 2018. The number of insects captured varied significantly with season (Kruskal-Wallis: χ2 = 23.12, degree of freedom [df] = 2, p < 0.001). Insects were significantly more abundant during summer (102 capture events) than winter 1 (56 capture events) and winter 2 (48 capture events) with the highest number of insects found in December 2017 and the lowest in June 2018 (Fig. 1a). Winter 2 also had significantly higher abundance of insects than winter 1 (Fig. 1a).
Seasonal differences in a) insect count, b) gum volume, and c) seed count in winter 1 (June – August 2017), summer (November 2017 – February 2018), and winter 2 (May 2018 – July 2018) at Lajuma Research Centre in South Africa. Asterisks show significant differences between seasons (p < 0.05 based on Kruskal-Wallis analyses followed by post hoc tests). Boxplots show the median and interquartile range, whiskers show the standard deviation, and points show the outliers.
We found significant variation in gum availability between seasons (χ2 = 14.69, df = 2, p < 0.0001) with the highest gum mass in summer (n = 69, median = 0.5 ml, interquartile range = 0.4–0.7 ml) and the lowest in winter 1 (n = 48; median = 0.2 ml, interquartile range = 0.1–0.6 ml; Fig. 1b). Gum volume in summer was significantly higher than winter 2. We also found significantly lower gum volumes in winter 1 compared with winter 2 (Fig. 1b).
Most of the seeds originated from fig (Ficus sp) and jojoba (Simondsdia chinensis) fruits. We found no significant variability between seasons (χ2 = 1.43, df = 2, p = 0.489); however, winter 2 had the highest count of seeds present in the feces (Fig. 1c).
There was no significant difference in body mass between seasons for both males (Kruskal-Wallis: χ2 = 4.34, df = 2, p = 0.11) and females (χ2 = 0.31, df = 2, p = 0.86; Table I).
Seasonal Weather Analysis
Rainfall was significantly higher in the summer season (χ2 = 7.03, df = 2, p = 0.029; post hoc Tukey HSD: p = 0.044), with peak levels observed in February 2018 (mean: 8.2 mm per day; Fig. 2) compared with winter 2, which received <1 mm of precipitation per day. Summer was not significantly higher than winter 1 (mean: 1.2 mm per day; Tukey HSD: p = 0.281). Ambient temperatures were significantly higher in summer (range = 9.9–35.6 °C; mean = 20.58 ± 4.59 °C; χ2 = 24.72, df = 2, p < 0.001) than in winter 1 (range = 4.4–29.6 °C; mean = 14.85 ± 0.59 °C; Tukey HSD: p < 0.001) and winter 2 (Tukey HSD: p < 0.001; Fig. 2).
Diversity of Bacterial Taxa
We sequenced 75 fecal samples. After excluding samples with low read numbers, we used 49 samples in analyses, from 15 adult males and 16 adult females (mean: females = 1.38 ± 0.62; males = 1.40 ± 0.83 samples from each individual). In total, we identified 17,269 sequences, ranging from 100–376 bp, and 295 unique OTUs with a 97% sequence identity threshold. We detected 12 phyla and 53 families in the fecal samples. We identified all taxa to at least family level, 136 taxa to genus level, and 72 to species level. Neither season (PERMANOVA: F = 0.984, R2 = 0.021, p = 0.444) nor sex (F = 0.593, R2 = 0.013, p = 0.695) had a significant effect on the diversity of the bacterial populations in the galago feces.
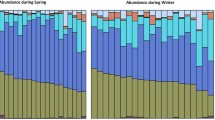
At phylum level, the fecal microbiome of the galago population was dominated by Actinobacteria (65%), Bacteroidetes (15%), Firmicutes (14%), Proteobacteria (3%), with unclassified (>1%; Fig. A2). We detected several other phyla, including Cyanobacteria, Fusobacteria, Planctomycetes, and Tenericutes, which represented less than 1% of the total taxa present. We detected ten phyla in samples from winter 1 and winter 2 and eight in samples from summer (Fig. 3).
The relative abundance of Actinobacteria significantly differed between all seasons (winter 1 = 6%, summer = 69%, winter 2 = 82%; Kruskal-Wallis: χ2 = 47.08, df = 2, p < 0.001). Winter 2 had a significantly higher relative abundance of Actinobacteria than summer and winter 1 (post hoc Dunn’s test: p < 0.001) and made up 82% of the total bacterial population. Summer had a significantly higher relative abundance of Actinobacteria than winter 1 (Dunn’s test: p < 0.001). An inverse relationship was present between Bacteroidetes and Firmicutes; with Bacteroidetes increasing in summer (winter 1= 4%, summer = 19%, winter 2 = 7%) and Firmicutes relative abundance highest in winter 1 (winter 1 = 44%, summer = 5%; winter = 4%); however, season did not have a significant impact on the abundance of Bacteroidetes (χ2 = 1.90, df = 2, p = 0.387). Firmicutes showed significant differences between all seasons (χ2 = 52.96, df = 2, p < 0.001; Fig. 3).
The most prominent families identified were Bifidobacteriaceae (62%), Prevotellaceae (13%), Clostridiaceae (6%), and Micrococcaceae (3%; Fig. A3). Overall, galagos share 51 bacterial families between seasons. Bifidobacteriaceae was the most dominant family for each individual throughout each sampling period (Fig. A4). In summer, we observed 23 families, including Neisseriaceae, Gemellaceae, and Rhodospirillaceae; in winter, we observed 21 families, including Peptostreptococcaceae, Turicibacteraceae, and Lactobacillaceae (Fig. 4; Table A1).
Venn diagram demonstrating shared and unique bacterial families found in galagos (O. crassicaudatus) during summer and winter in the Lajuma Research Centre in South Africa. The top number is the number of unique families. The number in brackets is the percentage of the total families unique to the season.
The most abundant genera comprised >85% of the total. Bifidobacterium (55%), Prevotella (14%), Clostridium (6%) were the most dominant genera detected with lesser contributions from Staphylococcus, Arthrobacter, Gardnerella, and unclassified genera of Proteobacteria, Micrococcaceae, and Bacteria (Fig. 5). Bifidobacterium bacteria increased significantly during summer (Kruskal-Wallis: Bifidobacterium: χ2 = 23.66, df = 2; p < 0.001) compared with both winter 1 (Dunn’s test: Z = 4.79; p < 0.001) and winter 2 (Z = 0.78; p < 0.001). Prevotella also significantly increased in summer (χ2 = 22.02, df = 2; p < 0.001) compared with the relative abundance in winter 1 (Z = 4.70; p < 0.001) and winter 2 (Z = 1.78; p < 0.001), while unclassified Micrococcaceae decreased in summer (t = 2.81, df = 2; p = 0.011). Clostridium bacteria were only present in the winter 2 samples, whereas Gardnerella, unclassified Micrococcaceae, and Staphylococcus were absent from the winter 2 samples altogether.
Seasonal differences in the relative abundance (log10 transformed) of the ten most abundant genera in the fecal microbiome of galagos (O. crassicaudatus) at Lajuma Research Centre, South Africa, between June 2017 and July 2018. Lines show the median, boxes the interquartile range, whiskers the standard deviation, and points outliers.
In winter, the most prominent species found in females were Bifidobacterium sp. (50%), Clostridium perfringens (10%), and Prevotella unclassified (3%). In the males, the most prominent species were Bifidobacterium sp. (19%), Clostridium perfringens (17%), and P. ruminicola (5%). In summer, the most abundant species in both sexes were Bifidobacterium sp., Prevotella sp., and P. ruminicola.
The LMM we fitted to explore which predictors influenced bacterial composition had moderate variation (marginal R2 = 0.26). It showed that bacterial composition was most influenced by insects (β = 2.62, standard error (SE) = 6.40, t = 0.41), gum (β = −3.33, SE = 10.23, t = −0.33), rainfall (β = −21.56, SE = 42.56, t = −0.51), and seeds (β = 0.68, SE = 0.82, t = 0.82). We found significant effects of rainfall, insects, gum, and seeds (Table II). Reproductive status showed no significant effect (χ2 = 1.12, df = 4, p = 0.89; Table II). Ambient temperature, gum mass, and insect availability were positively correlated with Proteobacteria (Fig. 6). Rainfall and seed count were negatively correlated with Bacteroidetes (Fig. 6b, d).
Correlations between the most abundant phyla (Actinobacteria, Bacteroidetes, Firmicutes, Proteobacteria) in the fecal microbial community and environmental factors. a) ambient temperature, b) rainfall, c) insect availability, d) seed count, e) gum volume, in galagos (Otolemur crassicaudatus) sampled at Lajuma Research Centre in South Africa between June 2017 and July 2018. Each element of the matrix shows the correlation coefficient (the number situated in the blank box), a scatter plot and line graph of plotted variables. *p < 0.05; **p < 0.01; ***p < 0.001.
Discussion
We found that the fecal microbial abundance of galagos is dominated by a small selection of bacterial genera; however, we can see the effects of seasonal and dietary changes as several taxa are able to thrive in a certain season while not in another. We found seasonal changes in ambient temperature and rainfall, which matched our findings that insects and gum volumes were more abundant in summer but less so in winter. Our findings are not fully consistent with the hypothesis that changes in season (dietary and environmental) influence changes in the galago fecal microbiome. However, our results might indicate the plasticity of the microbiota in galagos to cope with seasonal changes. First, we found significant effects of rainfall, insects, and gum on the bacterial composition. Interestingly, we found no significant seasonal differences in body mass. Second, we did not find significant differences in bacterial abundance and diversity between seasons. Bifidobacterium was the most dominant genus throughout the study period. Nevertheless, we did identify several unique bacterial taxa present in each season, such as Clostridium. We found an inverse relationship between the population of Firmicutes and Bacteroidetes in seasons. whereby there was a lower abundance of Firmicutes in summer while Bacteroidetes thrived as a phylum.
The lack of significant variation in body mass between seasons may be linked to the consistent food reserves of gum and fruit during winter to sustain the galago population. The galago population also may reduce their activity in winter to preserve energy levels. Testing this hypothesis will require data on their activity budgets across seasons.
Rainfall was an important driver of the galago microbiome composition. This pattern is similar to that in geladas (Theropithecus gelada; Baniel et al., 2021), where increased rainfall had a positive effect on the presence of cellulolytic degraders necessary to breakdown cell walls and fermenters (such as Prevotellaceae and Bacteroidales) required for the breakdown of polysaccharides into simple sugars (Flint et al., 2008). We observed a significant increase in Prevotella during summer. Prevotella, which is important in carbohydrate metabolism, is associated with a carbohydrate and plant-based diet (Wu et al., 2011; Amato et al., 2014; Kovatcheva-Datchary et al., 2015; Gorvitovskaia et al., 2016). High densities of Prevotella are found in primate species, which have a largely plant-based diet, such as the common marmoset (Callithrix jacchus; Sheh et al., 2012) and golden snub-nosed monkey (Rhinopithecus roxellanae; Zeng et al., 2022). It is possible that the increase in Prevotella was a result of a shift to a high-fat and -sugar diet of insect and gum. As the presence of Prevotella is linked to carbohydrate consumption and glucose metabolism, the high numbers of this genus in the galago population are possibly linked to the nondigestible food matter that can be broken down by Prevotella to allow for galagos to extract nutrients.
Firmicutes, which are associated with producing high-energy SCFAs (Schleifer, 2009), increase in relative abundance when individuals increase their dietary dependence on fruits. In contrast, Bacteroidetes abundance increases when fiber consumption increases and the consumption of high-glycemic index sugars decreases (De Filippo et al., 2010; Ley, 2010; Wu et al., 2011; Nagpal et al., 2018). The increase in Firmicutes densities in this study occurred during the cold, winter months when fruit consumption increased. This increase may lead to a rise in SCFA production (Houtman et al., 2022; Kapoor et al., 2023). An increase in Bacteroidetes in summer may occur in response to an increase in insect consumption and the need to digest hard exoskeletons (Wu et al., 2011; Rothman et al., 2014). In winter, we also saw an increase in the phylum Firmicutes. The ratio of Firmicutes to Bacteroides we observed may lead to increased fermentation functionality. Such functionality, which would improve energy production and has been noted in wild, black howler monkeys (Alouatta pigra; Amato et al., 2014) and Javan lorises (Nycticebus javanicus; Cabana et al., 2018).
A major role of the anatomical and physiological aspects of the galago’s gut and the major taxa characterized in the fecal microbiome is to ferment food matter (Nash, 1986; Langer & Clauss, 2018). Gut fermentation is a critical mechanism in several galago species that allows for the breakdown and absorption of nutrients from low-quality and difficult-to-digest foods that are necessary for survival in the less ideal conditions of winter (Nash, 1986). Due to the high-energy demands placed on galagos in winter, when temperature and resource quality are low and mating activity occurs, it is important that gut microbiome health is optimized to ensure nutrient and energy absorption. Fermentation spaces are present in the foregut of G. moholi and galagos to elongate the gut passage time and increase the volume of food matter broken down (Caton et al., 2000). The phylum Firmicutes, the family Bifidobacteriaceae, and the genus Clostridium are primary fermenters and may contribute to the fermentation abilities in the gut of galagos. Interestingly, Clostridium is only present in winter. Clostridium can cause harm when an individual’s health is compromised (McClane et al., 2012). Clostridium perfringens, the most common species found in the fecal gut microbiota of galagos in this study, is an opportunistic pathogen that can form part of the mucosa lining in the gastrointestinal tract (Pluvinage et al., 2019). During the study, we found two animals with dental pathologies in winter, which may be associated with the presence of Clostridium during this season.
The presence of the genus Clostridium in winter, while absent in summer, could suggest a role in fermentation and the breakdown of cellulose and dietary fiber, as in Tibetan macaques (Macaca thibetana; Sun et al., 2016). Clostridium is commonly found in decaying vegetation, soil, and plays a role in the fermentation of polysaccharides. Similarly, an increase in Clostridiales and a decrease in Bacteroides was determined in rats (Rattus norvegicus) fed fruit pectin (Licht et al., 2010). We found a slight increase in unclassified Proteobacteria genera in winter. As Proteobacteria plays a role in energy acquisition, this increase may be a coping mechanism during cold weather when energy loss is inevitable (Koren et al., 2012; Amato et al., 2014).
Unlike in most other primate host species, such as black howler monkeys (Amato et al., 2014) and ring-tailed lemurs (Lemur catta; Bennett et al., 2016), Bifidobacterium was the most abundant genus in galagos over the entire sampling period. Bifidobacterium is a commensal genus that plays a key role in carbohydrate metabolism and also is beneficial to the host health by protecting the gut mucosal lining from harmful bacteria (Pinzone et al., 2012; Ghouri et al., 2014; Turroni et al., 2014). Bifidobacterium also has been associated with the degradation of arabinogalactan (Crociani et al., 1994; Grieshop et al., 2002; Bornbusch et al., 2019; Long, 2018) and was abundant in the fecal microbiome in the lesser galago (G. moholi; Long, 2018) during the winter when gum exudates were the main food source. Certain genes in the genome of B. aesculapii are necessary for the breakdown of β-arabinans in gum exudates in Acacia enegal trees that common marmosets (Callithrix jacchus) eat (Zhu et al., 2021). Bifidobacterium also was prevalent in the fecal microbiome of exudativorous emperor tamarins (Sanguinus imperator; Duranti et al., 2017). A deeper assessment of the function of the fecal microbiome may provide more insight about whether Bifidobacterium also plays a functional role in the degradation of gum in galagos.
Conclusions
We assessed the gut microbial community in thick-tailed greater galagos, nocturnal primates living in a temperate environment and depending on food resources generally found in low quantities. Our results show the variation and versatility in the microbial community in the gut of galagos and their symbiotic relationship with their host. The microbiota appears to be able to shift with the changes in diet to allow the galagos to gain the nutrients they need from their food resources. Food availability in the study site appears to be influenced by rainfall. With climate change progressing, this may lead to decreases in rainfall, which may weaken the food resources currently available to galagos. Approaches, such as shotgun metagenomics coupled with more refined characterization of bacterial diversity and metabolic function, would further enhance our understanding of the relationship between diet and microbial communities (Troci et al., 2022). Overall, by identifying the gut microbial profile for galagos, this study provides insight into the ecology and the relationship between host and microbiome and demonstrates the importance of this relationship during changes in the environment.
References
Allan, A. T., LaBarge, L. R., Howlett, C., Bailey, A. L., Jones, B., Mason, Z., & Hill, R. A. (2022). Patterns of predation and meat-eating by chacma baboons in an Afromontane environment. Folia Primatologica, 94(1), 13–36.
Amato, K. R., Yeoman, C. J., Kent, A., Righini, N., Carbonero, F., Estrada, A., Rex Gaskins, H., Stumpf, R. M., Yildirim, S., Torralba, M., & Gillis, M. (2013). Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. The ISME Journal, 7(7), 1344–1353.
Amato, K. R., Leigh, S. R., Kent, A., Mackie, R. I., Yeoman, C. J., Stumpf, R. M., Wilson, B. A., Nelson, K. E., White, B. A., & Garber, P. A. (2014). The role of gut microbes in satisfying the nutritional demands of adult and juvenile wild, black howler monkeys (Alouatta pigra). American Journal of Physical Anthropology, 155, 652–664.
Amato, K. R., Leigh, S. R., Kent, A., Mackie, R. I., Yeoman, C. J., Stumpf, R. M., Wilson, B. A., Nelson, K. E., White, B. A., & Garber, P. A. (2015). The gut microbiota appears to compensate for seasonal diet variation in the wild black howler monkey (Alouatta pigra). Microbial Ecology, 69, 434–443.
Baniel, A., Amato, K.R., Beehner, J.C., Berman, T.J., Mercer, A., Perlman, R.F., Petrullo, L., Reitsema, L., Sams, S., Lu, A., & Snyder-Mackler, N. (2021). Seasonal shifts in the gut microbiome indicate plastic responses to diet in wild geladas. Microbiome, 9(26). https://doi.org/10.1186/s40168-020-00977-9.
Barton K. (2018). MuMIn: multi-model inference. R package version 3.6 https://cran.r-project.org/web/packages/MuMIn/MuMIn.pdf. Accessed 13 Jul 2021.
Bates, D. M. (2010). lme4: Mixed-effects modeling with R. Springer.
Bates, D., Mächler, M., Bolker, B., & Walker, S. (2014). Fitting linear mixed-effects models using lme4. Journal of Statistical Software, 67, 1–48. https://doi.org/10.18637/jss.v067.i01
Bearder, S. K. (1974). Aspects of the ecology and behaviour of the thick-tailed bushbaby, Galago galagos (Doctoral dissertation). University of Witwatersrand.
Bearder, S., & Martin, R. (1980). Acacia gum and its uses by bushbabies, Galago senegalensis (Primates: Lorisidae). International Journal of Primatology, 1(2), 103–128.
Bennett, G., Malone, M., Sauther, M. L., Cuozzo, F. P., White, B., Nelson, K. E., Stumpf, R. M., Knight, R., Leigh, S. R., & Amato, K. R. (2016). Host age, social group, and habitat type influence the gut microbiota of wild ring-tailed lemurs (Lemur catta). American Journal of Primatology, 78(8), 883–892.
Björk, J. R., Dasari, M., Grieneisen, L., & Archie, E. A. (2019). Primate microbiomes over time: longitudinal answers to standing questions in microbiome research. American Journal of Primatology, 81(10–11), e22970.
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulch, N. A., Abnet, C. C., Al-Ghalith, G. A., Alexander, H., Alm, E. J., Arumugam, M., Asnicar, F., Bai, Y., Bisanz, J. E., Bittinger, K., Brejnrod, A., Brislawn, C. J., Brown, C. T., Callahan, B. J., Caraballo-Rodríguez, A. M., Chase, J., … Caporaso, J. G. (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology, 37, 852–857. https://doi.org/10.1038/s41587-019-0209-9
Bornbusch, S. L., Greene, L. K., McKenney, E. A., Volkoff, S. J., Midani, F. S., Joseph, G., Gerhard, W. A., Iloghalu, U., Granek, J., & Gunsch, C. K. (2019). A comparative study of gut microbiomes in captive nocturnal strepsirrhines. American Journal of Primatology, 81(10–11), e22986.
Cabana, F., Clayton, J. B., Nekaris, K. A. I., Wirdateti, W., Knights, D., & Seedorf, H. (2018). Nutrient-based diet modifications impact the gut microbiome of the Javan slow loris (Nycticebus javanicus). Scientific Reports, 9, 4078.
Caton, J. M., Lawes, M., & Cunningham, C. (2000). Digestive strategy of the south-east African lesser bushbaby, Galago moholi. Comparative Biochemistry & Physiology Part A: Molecular & Integrative Physiology, 127(1), 39–48.
Chevalier, C., Stojanović, O., Colin, D. J., Suarez-Zamorano, N., Tarallo, V., Veyrat-Durebex, C., Rigo, D., Fabbiano, S., Stevanović, A., Hagemann, S., & Montet, X. (2015). Gut microbiota orchestrates energy homeostasis during cold. Cell, 163(6), 1360–1374.
Claesson, M. J., Jeffery, I. B., Conde, S., Power, S. E., O’connor, E. M., Cusack, S., Harris, H., Coakley, M., Lakshminarayanan, B., O’sullivan, O., & Fitzgerald, G. F. (2012). Gut microbiota composition correlates with diet and health in the elderly. Nature, 488(7410), 178–184.
Crociani, F., Alessandrini, A., Mucci, M. M., & Biavati, B. (1994). Degradation of complex carbohydrates by Bifidobacterium spp. International Journal of Food Microbiology, 24(1–2), 199–210.
Cuozzo, F. P., Halajian, A., Sauther, M. L., Rampedi, K. M., & Millette, J. B. (2021). First report of the thick-tailed bushbaby (Otolemur crassicaudatus) being preyed upon by an endemic carnivore (Caracal caracal) in South Africa. African Zoology., 56(3), 231–235.
David, L. A., Maurice, C. F., Carmody, R. N., Gootenberg, D. B., Button, J. E., Wolfe, B. E., Ling, A. V., Devlin, A. S., Varma, Y., Fischbach, M. A., & Biddinger, S. B. (2014). Diet rapidly and reproducibly alters the human gut microbiome. Nature, 505(7484), 559–563.
De Filippo, C., Cavalieri, D., Di Paola, M., Ramazzotti, M., Poullet, J. B., Massart, S., Collini, S., Pieraccini, G., & Lionetti, P. (2010). Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proceeding of the National Academy of Sciences, 107(33), 14691–14696.
Duranti, S., Magifesta, M., Lugli, G. A., Turroni, F., Anzalone, R., Milani, C., Mancabelli, L., Ossiprandi, M. C., & Ventura, M. (2017). Bifidobacterium vansinderenii sp. nov., isolated from faeces of emperor tamarin (Saguinus imperator). International Journal of Systematic and Evolutionary Microbiology, 67(10), 3987–3995.
Durowoju, O. S., Butler, M., Ekosse, G. I., & Ogiyo, J. O. (2019). Hydrochemical processes and isotopic study of geothermal springs within Soutpansberg, Limpopo Province. South Africa. Applied Sciences, 9(8), 1688.
Engelbrecht, D. (2016). Galagos as avian nest predators in South Africa. Primates, 57, 455–458.
Flint, H. J., Bayer, E. A., Rincon, M. T., Lamed, R., & White, B. A. (2008). Polysaccharide utilization by gut bacteria: potential for new insights from genomic analysis. National Reviews in Microbiology, 6, 121–131.
Génin, F. G., Masters, J. C., & Ganzhorn, J. U. (2010). Gummivory in cheirogaleids: primitive retention or adaptation to hypervariable environments? In A. Burrows & L. Nash (Eds.), The evolution of exudativory in primates developments in primatology: Progress and prospects (pp. 123–140). Springer. https://doi.org/10.1007/978-1-4419-6661-2_6
Ghouri, Y. A., Richards, D. M., Rahimi, E. F., Krill, J. T., Jelinek, K. A., & DuPont, A. W. (2014). Systematic review of randomised controlled trials of probiotics, prebiotics, and synbiotics in inflammatory bowel disease. Clinical Experimental Gastroenterology, 9, 473–487.
Gomez, A., Petrzelkova, K., Yeoman, C. J., Vlckova, K., Mrázek, J., Koppova, I., Carbonero, F., Ulanov, A., Modry, D., Todd, A., & Torralba, M. (2015). Gut microbiome composition and metabolomic profiles of wild western lowland gorillas (Gorilla gorilla gorilla) reflect host ecology. Molecular Ecology, 24(10), 2551–2565.
Gorvitovskaia, A., Holmes, S. P., & Huse, S. M. (2016). Interpreting Prevotella and Bacteroides as biomarkers of diet and lifestyle. Microbiome, 4, 15.
Grieshop, C. M., Flickinger, E. A., & Fahey, G. C., Jr. (2002). Oral administration of arabinogalactan affects immune status and faecal microbial population in dogs. Journal of Nutrition, 132(3), 478–482.
Grueber, C., Nakagawa, S., Laws, R., & Jamieson, I. (2011). Multimodel inference in ecology and evolution: Challenges and solutions. Journal of Evolutionary Biology, 24, 699–711.
Hanya, G., Tackmann, J., Sawada, A., Lee, W., Pokharel, S. S., de Castro Maciel, V. G., Toge, A., Kuroki, K., Otsuka, R., Mabuchi, R., & Liu, J. (2020). Fermentation ability of gut microbiota of wild Japanese macaques in the highland and lowland Yakushima: In vitro fermentation assay and genetic analyses. Microbial Ecology, 80, 459–474.
Happold, D., & Happold, C. (1992). Termites as food for the thick-tailed bushbaby (Otolemur crassicaudatus) in Malawi. Folia Primatologica, 58, 118–120.
Harcourt, C. (1986). Seasonal variation in the diet of South African galagos. International Journal of Primatology, 7(5), 491–506.
Hildebrandt, M. A., Hoffmann, C., Sherrill-Mix, S. A., Keilbaugh, S. A., Hamady, M., Chen, Y. Y., Knight, R., Ahima, R. S., Bushman, F., & Wu, G. D. (2009). High-fat diet determines the composition of the murine gut microbiome independently of obesity. Gastroenterology, 137(5), 1716–1724.
Hill, W. C. O. (1953). Primates. Comparative anatomy and taxonomy. I: Strepsirhini. University Press.
Houtman, T. A., Eckermann, H. A., Smidt, H., & de Weerth, C. (2022). Gut microbiota and BMI thoughout childhood: The role of firmicutes, bacteroidetes, and short-chain fatty acid producers. Scientific Reports, 12, 3140.
Irani, R., & Khaled, K. L. (2020). Quantitative analysis of nutrients in the gum exudates of Acacia nilotica. International Journal of Current Research and Review, 12(8), 11–16.
Jayne, R. (2020). Gum. In: It’s what’s for dinner: The dietary significance and nutritional content of acacia gum for five sympatric primate species living in a high-altitude temperate environment. University of Colorado. Master’s Thesis.
Kapoor, P., Tiwari, A., Sharma, S., Tiwari, V., Sheoran, B., Ali, U., & Garg, M. (2023). Effect of anthocyanins on gut health markers, Firmicutes-Bacteroidetes ratio and short-chain fatty acids: a systematic review via meta-analysis. Scientific Reports, 13, 1729.
Kers, J. G., Velkers, F. C., Fischer, E. A., Hermes, G. D., Stegeman, J. A., & Smidt, H. (2018). Host and environmental factors affecting the intestinal microbiota in chickens. Frontiers in Microbiology, 9, 235.
Koren, O., Goodrich, J. K., Cullender, T. C., Spor, A., Laitinen, K., Bäckhed, H. K., Gonzalez, A., Werner, J. J., Angenent, L. T., Knight, R., & Bäckhed, F. (2012). Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell, 150(3), 470–480.
Kotze, A., Dalton, D. L., Strinden, M., Sauther, M. L., Cuozzo, F. P., & Stone, A. C. (2016). An evaluation of the oral microbiome and potential zoonoses of the Thick-Tailed or Greater Galago (Otolemur crassicaudatus). African Primates, 11, 19–26.
Kovatcheva-Datchary, P., Nilsson, A. C., Akrami, R., Lee, Y. S., De Vadder, F., Arora, T., Hallén, A., Martens, E. C., Björck, I. M., & Bäckhed, F. (2015). Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metabolism, 22(6), 971–982.
Langer, P., & Clauss, M. (2018). Morphological adaptation of the Eutherian gastrointestinal tract to diet. Vertebrate Zoology, 68, 237–252. https://doi.org/10.5167/uzh-157906
Ley, R. E. (2010). Obesity and the human microbiome. Current Opinions in Gastroenterology, 26(1), 5–11.
Licht, T. R., Hansen, M., Bergstrom, A., Poulsen, M., Krath, B. N., Markowski, J., & Wilcks, A. (2010). Effects of apples and specific apple components on the cecal environment of conventional rats: Role of apple pectin. BMC Microbiology, 10(1), 13. https://doi.org/10.1186/1471-2180-10-13
Long, C. (2018). A seasonal comparison of the gut microbiome of the Southern Lesser Galago, Galago moholi (A.Smith 1836). University of Pretoria. Master’s dissertation.
Long, C., Tordiffe, A. S. W., Sauther, M. L., Cuozzo, F. P., Millette, J. L., Ganswindt, A., & Scheun, J. (2021). Seasonal drivers of faecal glucocorticoid metabolite concentrations in an African strepsirrhine primate, the thick-tailed greater galago (Otolemur crassicaudatus). Conservation Physiology, 9(1), coa081. https://doi.org/10.1093/conphys/coab081
Macfarlane, G. T., & Macfarlane, S. (2012). Bacteria, colonic fermentation, and gastrointestinal health. Journal of AOAC International, 95(1), 50–60.
MacKinnon, J. R., & MacKinnon, K. S. (1980). Niche differentiation in a primate community. Malayan forest primates: Ten years’ study in tropical rain forest (pp. 167–190). Spinger US.
Mallott, E. K., Amato, K. R., Garber, P. A., & Malhi, R. S. (2018). Influence of fruit and invertebrate consumption on the gut microbiota of wild white-faced capuchins (Cebus capucinus). American Journal of Physical Anthropology, 165(3), 576–588.
Manor, O., Dai, C. L., Kornilov, S. A., Smith, B., Price, N. D., Lovejoy, J. C., Gibbons, S. M., & Magis, A. T. (2020). Health and disease markers correlate with gut microbiome composition across thousands of people. Nature Communications, 11, 5206.
McClane, B.A., Robertson, S.L. & Li, J. (2012). Clostridium perfringens. Food Microbiology: fundamentals and frontiers. 465–489.
Mostert, T. H. C. (2010). Vegetation ecology of the Soutpansberg and Blouberg area in the Limpopo Province. University of Pretoria. Doctoral dissertation.
Nagpal, R., Shively, C. A., Appt, S. A., Register, T. C., Michalson, K. T., Vitolins, M. Z., & Yadav, H. (2018). Gut microbiome composition in non-human primates consuming a Western or Mediterranean diet. Frontiers in Nutrition, 5, 28.
Nakagawa, S., Johnson, P. C., & Schielzeth, H. (2017). The coefficient of determination R 2 and intra-class correlation coefficient from generalized linear mixed-effects models revisited and expanded. Journal of the Royal Society Interface, 14, 20170213. https://doi.org/10.1098/rsif.2017.0213
Nash, L. T. (1986). Dietary, behavioral, and morphological aspects of gummivory in primates. American Journal of Physical Anthropology, 29(S7), 113–137.
Nicholson, J. K., Holmes, E., Kinross, J., Burcelin, R., Gibson, G., Jia, W., & Pettersson, S. (2012). Host-gut microbiota metabolic interactions. Science, 336, 1262–1267. https://doi.org/10.1126/science.1223813
Nowack, J., Mzilikazi, N., & Dausmann, K. H. (2010). Torpor on demand: heterothermy in the non-lemur primate Galago moholi. PLoS One, 5(5), e10797.
Nowack, J., Wippich, M., Mzilikazi, M., & N., Dausmann, K.H. (2013). Surviving the cold, dry period in Africa: behavioral adjustments as an alternative to heterothermy in the African lesser bushbaby (Galago moholi). International Journal of Primatology, 34, 49–64.
Olivieros, J. C. (2015). Venny. An interactive tool for comparing lists with Venn’s diagrams. https://bioinfogp.cnb.csic.es/tools/venny/index.html. Accessed 24 Jan 2023.
O’Malley, R. C., & Power, M. L. (2012). Nutritional composition of actual and potential insect prey for the Kasekela chimpanzees of Gombe National Park. Tanzania. American Journal of Physical Anthropology, 149(4), 493–503.
Orkin, J. D., Campos, F. A., Myers, M. S., Hernandez, S. E. C., Guadamuz, A., & Melin, A. D. (2019). Seasonality of the gut microbiota of free-ranging white-faced capuchins in a tropical dry forest. The ISME Journal, 13, 183–196.
Pinzone, M. R., Celesia, B. M., Di Rosa, M., Cacopardo, B., & Nunnari, G. (2012). Microbial translocation in chronic liver diseases. International Journal of Microbiology, 2012, 1–12. https://doi.org/10.1155/2012/694629
Pluvinage, B., Massel, P. M., Burak, K., & Boraston, A. B. (2019). Structural and functional analysis of four family 84 glycoside hydrolases from the opportunistic pathogen Clostridium perfringens. Glycobiology, 30(1), 49–57. https://doi.org/10.1093/glycob/cwz069
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., & Yarza, P. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research, 41, 590–596.
R Core Team. (2020). R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/
Reyna-Hurtado, R., Teichroeb, J. A., Bonnell, T. R., Hernández-Sarabia, R. U., Vickers, S. M., Serio-Silva, J. C., Sicotte, P., & Chapman, C. A. (2018). Primates adjust movement strategies due to changing food availability. Behavioral Ecology, 29(2), 368–376.
Rothman, J. M., Raubenheimer, D., Bryer, M. A. H., Takahashi, M., & Gilbert, C. C. (2014). Nutritional contributions of insects to primate diets: Implications for primate evolution. Journal of Human Evolution, 71, 59–69.
Said, H. M., & Mohammed, Z. M. (2006). Intestinal absorption of water-soluble vitamins: an update. Current Opinion in Gastroenterology, 22, 140–146. https://doi.org/10.1097/01.mog.0000203870.22706.52
Scheppach, W. (1994). Effects of short chain fatty acids on gut morphology and function. Gut, 35, 35–38. https://doi.org/10.1136/gut.35.1_suppl.s35
Scheun, J., Bennett, N. C., Ganswindt, A., & Nowack, J. (2014). Spicing up the menu: evidence of fruit feeding in Galago moholi. Primates, 55, 359–363.
Scheun, J., Bennett, N. C., Ganswindt, A., & Nowack, J. (2015). The hustle and bustle of city life: monitoring the effects of urbanisation in the African lesser bushbaby. Science of Nature, 102, 1–11.
Scheun, J., Nowack, J., Bennett, N. C., & Ganswindt, A. (2016). Female reproductive activity and its endocrine correlates in the African lesser bushbaby, Galago moholi. Journal of Comparative Physiology B, 186, 255–264.
Schleifer, K.-H. (2009). In Bergey’s Manual® of Systematic Bacteriology (pp. 19–1317). Springer.
Sheh, A., Artim, S. C., Burns, M. A., Molina-Mora, J. A., Lee, M. A., Dzink-Fox, J., Muthupalani, S., & Fox, J. G. (2022). Analysis of gut microbiome profiles in common marmosets (Callithrix jacchus) in health and intestinal disease. Scientific Reports, 12, 4430. https://doi.org/10.1038/s41598-022-08255-4
Smith, A. C. (2010). Exudativory in primates: Interspecific patterns. In A. M. Burrows & L. T. Nash (Eds.), The Evolution of exudativory in primates, developments in primatology: Progress and prospects. Springer.
Strier, K. B. (2009). Seeing the forest through the seeds: mechanisms of primate behavioral diversity from individuals to populations and beyond. Current Anthropology, 50(2), 213–228.
Sun, B., Wang, X., Bernstein, S., Huffman, M. A., Xia, D. P., Gu, Z., Chen, R., Sheeran, L. K., Wagner, R. S., & Li, J. (2016). Marked variation between winter and spring gut microbiota in free-ranging Tibetan Macaques. (Macaca thibetana). Scientific Reports, 6(1), 26035.
Troci, A., Rausch, P., Waschina, S., Lieb, W., Franke, A., & Bang, C. (2022). Long-term dietary effects on human gut microbiota composition employing shotgun metagenomics data analysis. Molecular Nutrition & Food Research, 2101098. https://doi.org/10.1002/mnfr.202101098
Turroni, F., Duranit, S., Bottacini, F., Guglielmetti, S., Van Sinderen, D., & Ventura, M. (2014). Bifidobacterium bifidum as an example of a specialised human gut commensal. Frontiers in Microbiology, 21(5), 437.
Van Wyk, A. E., & Smith, G. F. (2001). Regions of a review with emphasis on succulents. In A. E. Van Wyk & G. F. Smith (Eds.), Regions of floristic endemism in Southern Africa: a review with emphasis on succulents. Umdaus Press.
Von Maltitz, E. F., Kirsten, F., Malebana, P. S., Belmain, S. R., Sandmann, E. R. I. C., Lundall-Magnuson, E., Mosala, M., Hlangwani, K. F., Mavasa, M. R., Mugogovhali, T. V., & Nyamande, T. P. (2003). Developing a rodent management strategy for South Africa's Limpopo province. In G. R. Singleton, L. A., Hinds, C. J. Krebs, & D. M. Spratt (Eds.), Rats, mice and people: rodent biology and management, Australian Centre for International Agricultural Research (ACIAR) Monograph Series (Vol. 96, pp. 418–421). Canberra: ACIAR.
Wu, G. D., Chen, J., Hoffmann, C., Bittinger, K., Chen, Y. Y., Keilbaugh, S. A., Bewtra, M., Knights, D., Walters, W. A., Knight, R., & Sinha, R. (2011). Linking long-term dietary patterns with gut microbial enterotypes. Science, 334, 105–108.
Xu, Z., & Knight, R. (2015). Dietary effects on human gut microbiome diversity. British Journal of Nutrition, 133, 1–5.
Yuan, D., Li, C., You, L., Dong, H., & Fu, X. (2020). Changes of digestive and fermentation properties of Sargassum pallidum polysaccharide after ultrasonic degradation and its impacts on gut microbiota. International Journal of Biological Macromolecules, 164, 1443–1450.
Zeng, Y., Pu, Y., Niu, L. L., Deng, J. B., Zeng, D., Amato, K. R., Li, Y., Zhou, Y., Lin, Y. C., Wang, J., & Wu, L. Q. (2022). Comparison of gastrointestinal microbiota in golden snub-nosed monkey (Rhinopithecus roxellanae), green monkey (Chlorocebus aethiops sabaeus), and ring-tailed lemur (Lemur catta) by high throughput sequencing. Global Ecology and Conservation, 33, e01946.
Zhu, L., Yang, Q., Suhr Van Haute, M. J., Kok, C. R., Gomes-Neto, J. C., Pavlovikj, N., Pillai, R., Sinha, R., Hassenstab, H., Mustoe, A., & Moriyama, E. N. (2021). Captive common marmosets (Callithrix jacchus) are colonised throughout their lives by a community of Bifidobacterium species with species-specific genomic content that can support adaptation to distinct metabolic niches. Mbio, 12(4), e01153-21.
Zuur, A. F., & Ieno, E. N. (2016). A protocol for conducting and presenting results of regression-type analyses. Methods in Ecology and Evolution, 7(6), 636–645.
Acknowledgments
The authors thank the late Ian Gaigher and Bibi Linden and Jabu Linden for permission to work at the Lajuma Research Centre and for their assistance throughout the duration of this study. The authors would like to acknowledge the effort and guidance given by the editors and reviewers to ensure this manuscript is of publishable standards. This work was supported by The National Science Foundation, USA (grant number 1638833) and South African National Biodiversity Institute’s the National Zoological Gardens, South Africa.
Funding
Open access funding provided by University of Pretoria.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Luca Pozzi
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Long, C., Scheun, J., Sauther, M.L. et al. Seasonal Effects on the Fecal Microbial Composition of Wild Greater Thick-Tailed Galagos (Otolemur crassicaudatus). Int J Primatol (2023). https://doi.org/10.1007/s10764-023-00407-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10764-023-00407-1