Abstract
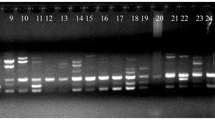
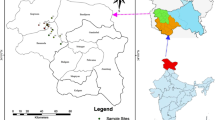
Wood-apple (Limonia acidissima L., Rutaceae) is an underutilized fruit tree species in the Indian subcontinent. The edible fruit and other parts of the wood-apple tree have immense medicinal and nutritional properties. In this study, we used 24 polymorphic simple sequence repeats (SSRs), developed de novo through RNA sequencing, to estimate the genetic diversity and population structure in 96 wood-apple samples collected from 16 states of India. We identified 11,401 SSRs from 215,859 unigenes. The dinucleotide repeat motifs were the most abundant SSRs (56.89%), followed by trinucleotide (37.61%) and tetranucleotide repeats (3.63%). SSR profiling detected 129 alleles, with an average of 5.37 per locus and an average PIC value of 0.62. The Jaccard’s genetic dissimilarity among 96 samples ranged from 0.39 to 1.0 (average 0.94 ± 0.03). Cluster analyses by Neighbour-joining (NJ) and principal coordinates analysis (PCoA) grouped all samples into three clusters. The Bayesian model-based structure analysis also identified three subpopulations. Population genetic analysis showed moderate gene diversity (0.226 ± 0.007) and Shannon’s information index (0.369 ± 0.009) in the species. Analysis of molecular variance (AMOVA) revealed higher genetic variance within the populations than among populations, while the FST statistics (0.134, p < 0.010) indicated low but significant genetic differentiation. The overall results indicated moderate to high genetic diversity among wood-apple samples. This study identified 14 diverse samples and nine highly informative SSRs that could be useful for developing breeding programs to improve the species and conserve its genetic resources.




Similar content being viewed by others
References
Ahmed S, Rattanpal HS, Singh G (2019) Diversity, characterization and evaluation of Pummelo (Citrus grandis Merr.) cultivars using SSR markers and quality parameters. Indian J Genet 79(3):594–605. https://doi.org/10.31742/IJGPB.79.3.9
Andrias DR, Fahmida U, Adi AC (2019) Nutritional potential of underutilized food crops to improve diet quality of young children in food insecure prone areas of Madura Island, Indonesia. Asia Pac J Clin Nutr 28(4):826–836
Annavaram V, Posa VR, Uppara VG, Jorepalli S, Somala AR (2015) Facile green synthesis of silver nanoparticles using Limonia acidissima leaf extract and its antibacterial activity. Bionanosci 5(2):97–103. https://doi.org/10.1007/s12668-015-0168-7
Arora RK (2014) Diversity in underutilized plant species—an Asia Pacific perspective. New Delhi, Biodiversity International, p 203
Asp T, Frei UK, Didion T, Nielsen KK (2007) Frequency, type, and distribution of EST-SSRs from three genotypes of Lolium perenne, and their conservation across orthologous sequences of Festuca arundinacea, Brachypodium distachyon, and Oryza sativa. BMC Plant Biol 7:36. https://doi.org/10.1186/1471-2229-7-36
Barbuiya AR, Khan ML, Dayanandan S (2016) Genetic structure and diversity in natural and domesticated populations of Citrus medica L. in the Eastern Himalayan region of Northeast India. Ecol Evol 6(12):3898–3911. https://doi.org/10.1002/ece3.2174
Bhargava R, Gurjar K, Singh AK, Singh S (2019) Genetic diversity of wood-apple (Feronia limonia L.) revealed by random amplified polymorphic DNA. Int J Chem Stud 7(1):208–212
Chaudhari BB, Annapure US (2021) Rheological, physicochemical, and spectroscopic characterizations of Limonia acidissima L. gum exudate with an application in extrusion processing. Carbohydr Polymer 2:100020. https://doi.org/10.1016/j.carpta.2020.100020
Chen LY, Cao YN, Yuan N, Nakamura K, Wang G-M, Qui W-X (2015) Characterization of transcriptome and development of novel EST-SSR makers based on next-generation sequencing technology in Neolitsea sericea (Lauraceae) endemic to East Asian land-bridge islands. Mol Breed 35:187. https://doi.org/10.1007/s11032-015-0379-1
Earl DA, Von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–366. https://doi.org/10.1007/s12686-011-9548-7
Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5:435–445. https://doi.org/10.1038/nrg1348
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Hofer T, Foll M (2009) Detecting loci under selection in a hierarchically structured population. Heredity 103:285–298. https://doi.org/10.1038/hdy.2009.74
Feng S, Zhao L, Liu Z, Liu Y, Yang T, Wei A (2017) De novo transcriptome assembly of Zanthoxylum bungeanum using Illumina sequencing for evolutionary analysis and simple sequence repeat marker development. Sci Rep 7:16754. https://doi.org/10.1038/s41598-017-15911-7
Grover A, Sharma PC (2016) Development and use of molecular markers: past and present. Crit Rev Biotechnol 36(2):290–302. https://doi.org/10.3109/07388551.2014.959891
Guo WW, Deng XX (2001) Wide somatic hybrids of Citrus with its related genera and their potential in genetic improvement. Euphytica 118:175–183. https://doi.org/10.1023/A:1004147208099
Haque MA, Hossain MS, Akanda MR, Haque MA, Naher S (2019) Procedure optimization of Limonia acidissima leaf extraction and silver nanoparticle synthesis for prominent antibacterial activity. Chem Select 4(48):14276–14280. https://doi.org/10.1002/slct.201904019
Hemalatha C, Parameshwari S (2019) Development and standardization of wood-apple (Limonia acidissima) incorporated novel food products. Int J Pharm Sci Res 10:5087–5093. https://doi.org/10.13040/IJPSR.0975-8232.10(11).5087-93
Hu Y, Tian L, Shi J, Tian J, Zhao L, Feng S, Wei A (2018) Genetic structure of cultivated Zanthoxylum species investigated with SSR markers. Tree Genet Genomes 14:89. https://doi.org/10.1007/s11295-018-1300-y
Huang X, Yan HD, Zhang XQ, Zhang J, Frazier TP, Huang DJ, Lu L, Huang LK, Liu W, Peng Y, Ma X, Yan YH (2016) De novo transcriptome analysis and molecular marker development of two Hemarthria species. Front Plant Sci 7:496. https://doi.org/10.3389/fpls.2016.00496
Hung TH, Wu ML, Su HA (2008) Identification of alternative hosts of the fastidious bacterium causing citrus greening disease. Phytopthol 148:321–326. https://doi.org/10.1046/j.1439-0434.2000.00506.x
Kalia RK, Rai MK, Kalia S, Singh R (2011) Microsatellite markers: an overview of the recent progress in plants. Euphytica 177(3):309–334. https://doi.org/10.1007/s10681-010-0286-9
Kaushik P, Kumar S (2018) Transcriptome analysis of bael (Aegle marmelos (L.) Corr.), a member of family Rutaceae. Forests 9:450. https://doi.org/10.3390/f9080450
Kerkar SP, Patil S, Arya SS, Dabade A, Sonawane SK (2020) Limonia acidissima: Versatile and nutritional fruit of India. Int J Fruit Sci 20(sup2):S405–S413. https://doi.org/10.1080/15538362.2020.1737302
Lamani S, Anu-Appiah KA, Murthy HN, Dewir YH (2021) Fatty acid profile, tocopherol content of seed oil, and nutritional analysis of seed cake of Wood-apple (Limonia acidissima L.), an underutilized fruit-yielding tree species. Horticulturae 7:275. https://doi.org/10.3390/horticulturae7090275
Liang M, Yang X, Li H, Su S, Yi H, Chai L, Deng X (2015) De Novo transcriptome assembly of pummelo and molecular marker development. PLoS ONE 10:0120615. https://doi.org/10.1371/journal.pone.0120615
Liu K, Muse SV (2005) Power Marker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Ma J, Zhao M, Zhang C, Wu X, Yang G (2020) Synthesis of L. acidissima mediated tin oxide nanoparticles for cervical carcinoma treatment in nursing care. J Drug Deliv Sci Technol 57:101745. https://doi.org/10.1016/j.jddst.2020.101745
Mali KK, Sutar GV, Dias RJ, Devade OA (2021) Evaluation of nootropic activity of Limonia acidissima against scopolamine-induced amnesia in rats. Turkish J Pharm Sci 18(1):3–9. https://doi.org/10.4274/tjps.galenos.2019.30316
Mao AA, Dash SS (2020) Flowering Plants of India — an Annotated Checklist (Dicotyledons), Botanical Survey of India, Kolkata, 1: 1–970
Martienssen RA, Colot V (2001) DNA methylation and epigenetic inheritance in plants and filamentous fungi. Science 293(5532):1070–1074. https://doi.org/10.1126/science.293.5532.1070
Murthy HN, Dalawai D (2020) Bioactive compounds of wood-apple (Limonia acidissima L.). In: Murthy H, Bapat V (eds) Bioactive compounds in underutilized fruits and nuts, reference series in phytochemistry. Springer, Cham, pp 543–569. https://doi.org/10.1007/978-3-030-30182-8_39
Patil BN, Taranath TC (2016) Limonia acidissima L. leaf mediated synthesis of zinc oxide nanoparticles: a potent tool against Mycobacterium tuberculosis. Int J Mycobacteriol 5(2):197–204. https://doi.org/10.1016/j.ijmyco.2016.03.004
Patil BN, Taranath TC (2018) Limonia acidissima L. leaf mediated synthesis of silver and zinc oxide nanoparticles and their antibacterial activities. Microb Pathog 115:227–232. https://doi.org/10.1016/j.micpath.2017.12.035
Pawar O, Deshpande N, Dagade S, Waghmode S, Nigam JP (2016) Green synthesis of silver nanoparticles from purple acid phosphatase apoenzyme isolated from a new source Limonia acidissima. J Exp Nanosci 11:28–37. https://doi.org/10.1080/17458080.2015.1025300
Peakall R, Smouse PE (2012) GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research- an update. Bioinformatics 28:2537–2539. http://bioinformatics.oxfordjournals.org/content/28/19/2537
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Perrier X, Flori A, Bonnot F (2003) Data analysis methods. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science Publishers Montpellier, Enfield, pp 43–76
Ponnuraj S, Jaganathan D, Kanagarajan M, Castro J (2015) Influence of Limonia acidissima L. against the biofilm-forming Aeromonas hydrophila isolated from freshwater fishes. J Biochem Technol 6(1):910–921
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep 15:8–15. https://doi.org/10.1007/BF02772108
POWO (2023) Plants of the World Online. Facilitated by the Royal Botanic Gardens, Kew. Published on Internet; http://www.plantsoftheworldonline.org/ Retrieved 27 July 2023
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Ratnayake SS, Kumar L, Kariyawasam CS (2020) Neglected and underutilized fruit species in Sri Lanka: Prioritisation and understanding the potential distribution under climate change. Agronomy 10(1):34. https://doi.org/10.3390/agronomy10010034
Rodrigues S, de Brito ES, de Oliveira SE (2018) Wood-apple—Limonia acidissima. In: Rodrigues S, de Oliveira SE, de Brito ES (eds) Exotic fruits reference guide. Academic Press, Cambridge, pp 443–446. https://doi.org/10.1016/B978-0-12-803138-4.00060-5
Samarina LS, Kulyan RV, Koninskaya NG, Gorshkov VM, Ryndin AV, Hanke MV, Flachowsky H, Reim S (2021) Genetic diversity and phylogenetic relationships among citrus germplasm in the Western Caucasus assessed with SSR and organelle DNA markers. Sci Hortic 288:110355. https://doi.org/10.1016/j.scienta.2021.110355
Schlötterer C (2000) Evolutionary dynamics of microsatellite DNA. Chromosoma 109:365–371. https://doi.org/10.1007/s004120000089
Sekhar EC, Rao KSVK, Rao KM, Kumar SP (2016) A green approach to synthesize controllable silver nanostructures from Limonia acidissima for inactivation of pathogenic bacteria. Cogent Chem 2:1. https://doi.org/10.1080/23312009.2016.1144296
Slatkin M (1987) Gene flow and geographic structure of natural populations. Science 236:787–792
Swingle WT, Reece PC (1967) The botany of Citrus and its wild relatives. In: Reuther W, Webber HJ, Batchelor LD (eds) The citrus industry, vol 1. University of California, Berkeley, pp 190–430
Tautz D, Renz M (1984) Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res 12(10):4127–4138. https://doi.org/10.1093/nar/12.10.4127
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422. https://doi.org/10.1007/s00122-002-1031-0
Tóth G, Gáspári Z, Jurka J (2000) Microsatellites in different eukaryotic genomes: Survey and analysis. Genome Res 10(7):967–981. https://doi.org/10.1101/gr.10.7.967
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23:48–55. https://doi.org/10.1016/j.tibtech.2004.11.005
Veeraiyan S, Gayathri SK, Pavithradevi G, Deepa S (2019) Limonia acidissima leaf interceded phytosynthesis of iron nanoparticles—A latent implement against A549 cell lines at SSRN.https://doi.org/10.2139/ssrn.3342984
Vieira MLC, Santini L, Diniz AL, Munhoz CF (2016) Microsatellite markers: what do they mean and why are they so useful. Genet Mol Biol 39:312–328. https://doi.org/10.1590/1678-4685-GMB-2016-0027
Vyas S, Gaur A, Tyagi AK, Purohit SD (2008) Assessment of genetic diversity in Feronia Limonia (L.) Swingle using inter simple sequence repeats. J Sustain for 27:328–342. https://doi.org/10.1080/10549810802256239
Wang Z, Fang B, Chen J et al (2010) De novo assembly and characterization of root transcriptome using Illumina paired-end sequencing and development of cSSR markers in sweet potato (Ipomoea batatas). BMC Genomics 11:726. https://doi.org/10.1186/1471-2164-11-726
Weising K, Nybom H, Wolff K, Kahl G (2005) DNA fingerprinting in plants: principles, methods, and applications, 2nd edn. CRC Press, Boca Raton, p 472
Woo J, Park YC, Lee JW, Yun SH, Kim M, Park S, Lee Y, Song KJ, Kim HB (2019) Evaluation of polyembryony for genetic resources and efficacy of simple sequence repeat markers for the identification of nucellar and zygotic embryo-derived individuals in Citrus. Appl Biol Chem 62:30. https://doi.org/10.1186/s13765-019-0437-1
Wu Q, Zang F, Xie X, Ma Y, Zheng Y, Zang D (2020) Full-length transcriptome sequencing analysis and development of EST-SSR markers for the endangered species Populus wulianensis. Sci Rep 10:16249. https://doi.org/10.1038/s41598-020-73289-5
Yang X, Li H, Liang M, Xu Q, Chai L, Deng X (2015) Genetic diversity and phylogenetic relationships among citron (Citrus medica L.) and its relatives in south-west China. Tree Genet Genome 11:129. https://doi.org/10.1007/s11295-015-0955-x
Zhu S, Wang F, Shen W, Jiang D, Hong Q, Zhao X (2015) Genetic diversity of Poncirus and phylogenetic relationships with its relatives revealed by SSR and SNP/InDel markers. Acta Physiol Pl 37:141. https://doi.org/10.1007/s11738-015-1890-2-z
Acknowledgements
The authors are thankful to the director of CSIR-National Botanical Research Institute (CSIR-NBRI), Lucknow for providing facilities and support to conduct this work. AL is grateful to the University Grants Commission for the award of a fellowship (National Fellowship Scheme for Scheduled Caste) to perform her research work. This manuscript has an institutional communication number: CSIR-NBRI_MS/2021/10/04.
Funding
This work was supported by a Research Fellowship awarded to the first author (AL) by the University Grants Commission of India (Grant No. F1-17.1/2013-14/RGNF-2013-14-SC-UTT-52795).
Author information
Authors and Affiliations
Contributions
Annu Lata performed the experiments and prepared the manuscript; Annu Lata and H.K. Yadav analysed the data; N. K. Nair and H.K. Yadav critically reviewed the manuscript; N. K. Nair and H.K. Yadav formulated the original research plans, supervised the research and finalized the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lata, A., Yadav, H.K. & Nair, N.K. Genetic diversity and population structure analysis in wood-apple (Limonia acidissima L.) using simple sequence repeat (SSR) markers. Genet Resour Crop Evol 71, 1355–1368 (2024). https://doi.org/10.1007/s10722-023-01699-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01699-1