Abstract
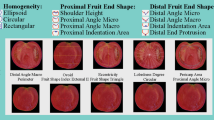
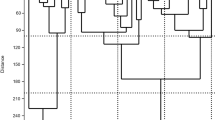
The recovery of ancient germplasm in tomato (Solanum lycopersicon L.) has become necessary to limit the wide genetic erosion caused by the employment of modern cultivars. Among germplasm collections, long shelf-life landraces could represent an important source of biodiversity. The present study provides a first set of molecular and phenotypic data on long shelf-life (so called “da serbo” in southern Italy) tomato collection, mainly originated from Sicily together with some landraces from Campania and Apulia. The analysis of fruit traits showed a low intra-varietal variation, while exhibiting a quite higher inter-varietal variability. Overall, the cultivars have been classified in six fruit shape classes of which flattened and slightly flattened included the 54.76 % of the collection. The principal component analysis (PCA) showed a large cluster in which almost all landraces from Sicily were included. The microsatellite (SSR) analysis confirmed a low intra-varietal variation, and the very low heterozygosity (Ho) revealed a high degree of homozygosity in these landraces. In accordance with limited morphological variability, the values of microsatellite polymorphism (PIC) showed a low genetic variability among these long shelf-life tomato cultivars. Cluster analysis based on 10 polymorphic SSR was not able to distinguish landraces for their different origin, while allowed to classify similar genotypes in four groups. Three groups showed a limited genetic distance while in a fourth largest and genetic variable cluster was included genotypes more selectable for traits of agronomic interest. Overall, the phenotypic and genetic variation allowed us to classify a collection of Sicilian long shelf-life tomato landraces.



Similar content being viewed by others
References
Alvarez AE, van de Wie CCM, Smulders MJM (2001) Use of microsatellites to evaluate genetic diversity and species relationships in the genus Lycopersicon. Theor Appl Genet 103:1283–1292
Andreakis N, Giordano I, Pentangelo A, Fogliano V, Graziani G, Monti LM, Rao R (2004) DNA fingerprinting and quality traits of corbarino cherry-like tomato landraces. J Agric Food Chem 52:3366–3371
Barbagallo RN, Chisari M, Branca F, Argento S, Spagna G (2006) Enzymatic degradative trait in long storage Sicilian tomato fruits (Lycopersicon esculentum Mill.). Italus Hortus 13:622-625
Barbagallo RN, Chisari M, Branca F, Spagna G (2008) Pectin methylesterase, polyphenol oxidase and physicochemical properties of typical long-storage cherry tomatoes cultivated under water stress regime. J Sci Food Agric 88:389–396
Benor S, Zhang M, Wang Z, Zhang H (2008) Assessment of genetic variation in tomato (Solanum lycopersicum) inbred lines using SSR molecular markers. J Genet Genom 35:373–379
Bredemeijer G, Cooke R, Ganal M, Peeters R, Isaac P, Noordijk Y, Rendell S, Jackson J, Röder MS, Wendehake K, Dijcks M, Amelaine M, Wickaert K, Bertrand L, Vosman B (2002) Construction and testing of a microsatellite database containing more than 500 tomato varieties. Theor Appl Genet 105:1019–1026
Breto MP, Asins MJ, Carbonell EA (1993) Genetic variability in Lycopersicon species and their genetic relationships. Theor Appl Genet 86:113–120
Bruni A (1855) Degli ortaggi e loro coltivazione presso la città di Napoli. Rivista agronomica, giornale di agricoltura, pastorizia, veterinaria e scienze affini. Annuario della Reale Scuola Superiore d’Agricoltura in Portici, vol I Stab Tip 1855-57-58 Napoli
Caramante M, Rao R, Monti LM, Corrado G (2009) Discrimination of ‘San Marzano’ accessions: a comparison of minisatellite, CAPS and SSR markers in relation to morphological traits. Sci Hortic 120:560–564
Caramante M, Corrado G, Monti LM, Rao R (2011) Simple sequence repeats are able to trace tomato cultivars in tomato food chains. Food Control 22:549–554
Casals J, Pascual L, Cañizares J, Cebolla-Cornejo J, Casañas F, Nuez F (2012) Genetic basis of long shelf life and variability into Penjar tomato. Genet Resour Crop Evol 59:219–229
Cebolla-Cornejo J, Roselló S, Nuez F (2013) Phenotypic and genetic diversity of Spanish tomato landraces. Sci Hortic 162:150–164
Chen J, Wang H, Shen H, Chai M, Li J, Qi M, Yang W (2009) Genetic variation in tomato populations from four breeding programs revealed by single nucleotide polymorphism and simple sequence repeat markers. Sci Hortic 122:6–16
Doyle JJ, Doyle LJ (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Figás MR, Prohens J, Raigón MD, Fernández-de-Córdova P, Fita A, Soler S (2014) Characterization of a collection of local varieties of tomato (Solanum lycopersicum L.) using conventional descriptors and the high-throughput phenomics tool tomato analyzer. Genet Resour Crop Evol. doi:10.1007/s10722-014-0142-1
Frary A, Xu Y, Liu J, Mitchell S, Tedeschi E, Tanksley S (2005) Development of a set of PCR based anchor markers encompassing the tomato genome and evaluation of their usefulness for genetics and breeding experiments. Theor Appl Genet 111:291–312
García-Martínez S, Andreani L, Garcia-Gusano M, Geuna F, Ruiz JJ (2006) Evaluation of amplified fragment length polymorphism and simple sequence repeats for tomato germplasm fingerprinting: utility for grouping closely related traditional cultivars. Genome 49:648–656
García-Martínez S, Corrado G, Ruiz JJ (2013) Diversity and structure of a sample of traditional Italian and Spanish tomato accessions. Genet Resour Crop Evol 60:789–798
Geethanjali S, Chen KY, Pastrana DV, Wang JF (2010) Development and characterization of tomato SSR markers from genomic sequences of anchored BAC clones on chromosome 6. Euphytica 173:85–97
He C, Poysa V, Yu K (2003) Development and characterization of simple sequence repeat (SSR) markers and their use in determining relationships among Lycopersicon esculentum cultivars. Theor Appl Genet 106:363–373
Huang X, Börner A, Röder M, Ganal M (2002) Assessing genetic diversity of wheat (Triticum aestivum L.) germplasm using microsatellite markers. Theor Appl Genet 105:699–707
Jones CJ, Edwards KJ, Castaglione S, Winfield MO, Sala F, van de Wiel C, Bredemeijer G, Vosman B, Matthes M, Daly A, Brettschneider R, Bettini P, Buiatti M, Maestri E, Malcevschi A, Marmiroli N, Aert R, Volckaert G, Rueda J, Linacero R, Vazquez A, Karp A (1997) Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Mol Breed 3:381–390
Liu K, Muse SV (2005) Powermarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Mazzucato A, Papa R, Bitocchi R, Pietro P, Nanni L, Negri V, Picarella ME, Siligato F, Soressi GP, Tiranti B, Veronesi F (2008) Genetic diversity, structure and marker-trait associations in a collection of Italian tomato (Solanum lycopersicum L.) landraces. Theor Appl Genet 116:657–669
Mazzucato A, Ficcadenti N, Caioni M, Mosconi P, Piccinini E, Sanampudi VRR, Sestili S, Ferrari V (2010) Genetic diversity and distinctiveness in tomato (Solanum lycopersicum L.) landraces: the Italian case study of ‘A pera Abruzzese’. Sci Hortic 125:55–62
Miller JC, Tanksley SD (1990) RFLP analysis of phylogenetic relationships and genetic variation in the genus Lycopersicon. Theor Appl Genet 80:437–448
Morgante M, Olivieri AM (1993) PCR-amplified microsatellites as markers in plant genetics. Plant J 3:175–182
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol 19:153–170
Nesbitt TC, Tanksley SD (2002) Comparative sequencing in the genus Lycopersicon: implications for the evolution of fruit size in the domestication of cultivated tomatoes. Genetics 162:365–379
Ohyama A, Asamizu E, Negoro S, Miyatake K, Yamaguchi H, Tabata S, Fukuoka H (2009) Characterization of tomato SSR markers developed using BAC-end and cDNA sequences from genome databases. Mol Breed 23:685–691
Page RDM (1996) Treeview: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Park YH, West MA, St. Clair DA (2004) Evaluation of AFLPs for germplasm fingerprinting and assessment of genetic diversity in cultivars of tomato (Lycopersicon esculentum L.). Genome 47:510–518
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Pressoir G, Berthaud J (2004) Population structure and strong divergent selection shape phenotypic diversification in maize landraces. Heredity 92:95–101
Primieri Carelli BP, Tseng L, Gerald S, Grazziotin FG, Echeverrigarays S (2006) Genetic diversity among Brazilian cultivars and landraces of tomato Lycopersicon esculentum Mill. revealed by RAPD markers. Genet Resour Crop Evol 53:395–400
Rick CM (1976) Tomato. Lycopersicon esculentum (Solanaceae). In: Simmonds N (ed) Evolution of crop plants. Longman, London, pp 268–273
Rick CM, Holle M (1990) Andean Lycopersicon esculentum var. cerasiforme: genetic variation and its evolutionary significance. Econ Bot 44:69–78
Rick CM, Laterrot H, Philouze J (1990) A revised key for the Lycopersicon species. Tomato Genet Coop Rep 40:31
Riggi E, Patanè C, Cosentino SL (2006) Contenuto di composti antiossidanti in ecotipi di pomodoro da serbo del Meridione d’Italia. Italus Hortus 13:779–782
Savo Sardaro ML, Marmiroli M, Maestri E, Marmiroli N (2013) Genetic characterization of Italian tomato varieties and their traceability in tomato food products. Food Sci Nutr 1:54–62
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality complete samples. Biometrika 52:591–611
Sharma KC, Verma S (2001) Analysis of genetic divergence in tomato. Ann Agric Res 22:71–73
Stachel M, Lelley T, Grausgruber H, Vollmann J (2000) Application of microsatellites in wheat (Triticum aestivum L.) for studying genetic differentiation caused by selection for adaptation and use. Theor Appl Genet 100:242–248
Stevens MR, Robbins MD (2007) Molecular markers in selection of tomato germplasm. In: Razdan MK, Mattoo AK (eds) Genetic improvement of solanaceous crops, vol 2, tomato. Science Publishers, Enfield, pp 239–260
Tam SM, Mhiri C, Vogelaar A, Kerkveld M, Pearce SR, Grandbastien MA (2005) Comparative analyses of genetic diversities within tomato and pepper collections detected by retrotransposon-based SSAP, AFLP and SSR. Theor Appl Genet 110:819–831
Tautz D (1989) Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res 17:6463–6471
Terzopoulos PJ, Bebeli PJ (2010) Phenotypic diversity in Greek tomato (Solanum lycopersicum L.) landraces. Sci Hortic 126:138–144
Villand J, Skroch PW, Lai T, Hanson P, Kuo CG, Nienhuis J (1998) Genetic variation among tomato accessions from primary and secondary centers of diversity. Crop Sci 38:1339–1347
Wang J, Tian L, Lee HS, Wei NE, Jiang H, Watson B, Madlung A, Osborn TC, Doerge RW, Comai L, Chen ZJ (2006) Genomewide nonadditive gene regulation in Arabidopsis allotetraploids. Genetics 172:507–517
Weber JL (1990) Informativeness of human (dC-dA)n(dG-dT)n polymorphisms. Genomics 7:524–530
Williams CE, St. Clair DA (1993) Phenetic relationships and levels of variability detected by restriction fragment length polymorphism and random amplified polymorphic DNA analysis of cultivated and wild accessions of Lycopersicon esculentum. Genome 36:619–630
Yi SS, Jato SA, Fujimura T, Yamanaka S, Watanabe J, Watanabe KN (2008) Potential loss of unique genetic diversity in tomato landraces by genetic colonization of modern cultivars at a non-center of origin. Plant Breed 127:189–196
Zeven AC (1998) Landraces: a review of definitions and classifications. Euphytica 104:127–139
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mercati, F., Longo, C., Poma, D. et al. Genetic variation of an Italian long shelf-life tomato (Solanum lycopersicon L.) collection by using SSR and morphological fruit traits. Genet Resour Crop Evol 62, 721–732 (2015). https://doi.org/10.1007/s10722-014-0191-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-014-0191-5