Abstract
Understanding how natural populations will respond to contemporary changes in climate is becoming increasingly urgent and of fundamental importance for the preservation of future biodiversity. Among vertebrates, amphibians and reptiles are more sensitive to environmental perturbations than endotherms and ectotherm diversity will likely be disproportionally impacted by climate change. Notwithstanding concerns surrounding the climate change resilience of ectotherm populations, accurately predicting future population trajectories based on contemporary ecological and physiological data alone remains challenging and much can be learnt by studying how populations have responded to climate change in the past. Genomic approaches can now assay the genetic diversity of contemporary population at an unprecedented scale but to date have been relatively underutilised when studying the demographic history of amphibians and reptiles. In this review, we first summarise how changing climatic conditions may influence the ectotherm phenotype and how this can translate to changes in fitness and population dynamics. We then discuss how the relative role of past climate in shaping ectotherm diversity has traditionally been approached in a phylogeographic context and how expanding genomic resources for ectotherm species can be leveraged to improve the study of past demography for many amphibian and reptilian groups. An integrative approach that links known proximate effects on phenotype due to climate change, with past changes in demographic trajectories will ultimately enable us to generate more accurate models of future population change and improve our ability to assess climate change resilience for many ectotherm groups.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Climate change has a broad impact on environmental conditions and as such can be a major driver of biodiversity turnover. The rate of environmental change and the diversity in taxa specific responses to changing habitats are important determinants of persistence and extinction (Hewitt 2004; Davis et al. 2005; Potter et al. 2018). For example, species can differ in their ability to mitigate the effects of climate change (resilience) with some taxa being more flexible in their ability to track suitable habitat or the capacity to adapt to novel circumstances in-situ, while others that lack such resilience facing local extinction (Davis et al. 2005; Sexton et al. 2017). Such idiosyncratic responses to climate change are not only observed between species from distinct taxonomic groups, but responses can sometimes even vary between closely related species that face common environmental changes (Deutsch et al. 2008; Moussalli et al. 2009; Hoffmann and Sgrò, 2018). With the global average annual temperature projected to rise between 1 and 4 °C by 2100 (Collins et al. 2013; IPCC 2014) and habitat destruction simultaneously accelerating (Habel et al. 2019), there is a growing need to understand how individual species and communities will respond to the ongoing changes in environmental conditions. Many studies address this topic by modelling prospective changes in species distribution and extinction risk (Araújo et al. 2006; Deutsch et al. 2008; Hof et al. 2011; Kafash et al. 2020). However, model accuracy strongly scales with the availability of (reliable) occurrence data and informative model parameters depend on our ability to quantify ecological and physiological requirements for a given species (Le Galliard et al. 2012; Waldvogel et al. 2020). For example, (environmental) specialists and generalists vary substantially in niche breadth and may therefore also differ in terms of resilience under changing conditions (Moussalli et al. 2009; Afonso Silva et al. 2017). Moreover, spatial heterogeneity in microclimatic conditions and complex interactions between taxa can also complicate the prediction of future effects of climate change on individual species (Araújo and Rahbek 2006; Araújo et al. 2006; Moritz et al. 2012; Taheri et al. 2021). Thus, it remains challenging to predict the degree of resilience for individual species by generalising across organismal groups and there is a pressing need to evaluate climate change resilience at a more detailed resolution across the Tree-of-Life.
Among vertebrates, ectotherms are particularly sensitive to changes in external conditions and have frequently been cited as a group where many species may be less able to cope with the predicted changes in climate (Deutsch et al. 2008; Rohr and Palmer 2013; Bestion et al. 2015; Winter et al. 2016). Ectotherms depend on ambient temperatures for thermoregulation and are sensitive to sizeable temperature changes in their surroundings (Winter et al. 2016). The worldwide decline in terrestrial ectotherms, reptiles and amphibians, has been partially attributed to their inability to track or adapt to changing climatic conditions and corresponding changes in prevalence of invasive species or foreign pathogens (Camargo et al. 2010; Hof et al. 2011; Oficialdegui et al. 2019). Many reptiles and amphibians have a limited dispersal capacity (Sinsch 1991; Araújo et al. 2006), are highly dependent on local environmental conditions for reproduction and development (Araújo et al. 2006; Blaustein et al. 2001; Jensen et al. 2018) and are generally considered to rely more heavily on external conditions than thermoregulating species (Deutsch et al. 2008; Bestion et al. 2015; Pie et al. 2017; Rolland et al. 2018). On the other hand, while these traits may be negative for local persistence during climate change, studying the demographic history of species with a narrow climatic niche in a geographic context can simultaneously provide important insights about the location and size of regions that have remained climatically stable over longer time scales (refugia; Hewitt 1996, 2004; Araújo et al. 2008). Thus, the study of climate change resilience in reptiles and amphibians is of particular interest due to their potential vulnerability under changing external conditions and terrestrial ectotherms are at the same time highly suited to evaluate how climatic stability may vary across landscapes.
The overarching aim of this Review is twofold. First, we will outline why climate change resilience is of particular concern in reptiles and amphibians by reviewing how changes in climate directly affect fitness via important biological dimensions, such as thermal regulation, reproduction and development. While more detailed Reviews on each of the aforementioned topics are in place (e.g. Blaustein et al. 2010; Diele-viegas et al. 2018; Butler 2019; Singh et al. 2020), providing a succinct overview on all topics combined will give a holistic view on the various ways that climate change can affect the ectotherm phenotype and how this may lead to population change. We will then continue by assessing how past changes in climate and extant patterns of ectotherm diversity have largely been studied in a phylogeographic rather than a demographic context. We discuss how genomic resources remain relatively absent for large groups of terrestrial ectotherms and how the lack of such resources has hindered the study of demographic history compared to other vertebrates. By providing an overview of different types of genomic data and corresponding developments in inference approaches, we highlight how such advances can be leveraged for the study of demographic history in amphibians and reptiles. This Review therefore outlines the myriad ways in which climate change has direct consequences for amphibian and reptilian fitness and diversity, while simultaneously discussing how we can improve our ability to quantify demographic history during past periods of climate change. These are both key requisites for identifying the proximate factors that have shaped population dynamics in the past and are thus essential for improving our ability to predict how contemporary populations will respond to future changes in environmental conditions. While reviewing (paleo-)environmental niche and predictive models themselves is outside the scope of the present Review (but see Sillero et al. 2021), here we emphasize the vulnerability of terrestrial ectotherms relative to endotherms and simultaneously highlight the outstanding challenges and opportunities for improving demographic inference in amphibians and reptiles. By doing so, we aim to bring herpetologists and population geneticists closer together and stimulate future efforts to study population change and the role of climate in one of the most evolutionary diverse groups of vertebrates.
The evolutionary consequences of climate change on terrestrial ectotherms
Identifying and quantifying the challenges that ectothermic vertebrates face under current climate change models is an essential but challenging task (Deutsch et al. 2008; Huey et al. 2009). Reptiles and amphibians are diverse groups that currently represent around 19,000 described species (Taylor et al. 2021). However, the true number of unique evolutionary lineages is putatively much higher due to the high incidence of cryptic diversity in both groups and extreme levels of species diversity in relatively understudied regions (e.g. the Tropics, Bickford et al. 2007). They are present on every continent except Antarctica, occupy a huge variety of habitats and cover a wide diversity in life-history traits (Pie et al. 2017; Diele-viegas et al. 2018; Taylor et al. 2021). They are strongly reliant on ambient temperature for thermoregulation and metabolic regulation and changes in temperature or corresponding environmental conditions such as the alteration of precipitation patterns, can therefore have many direct physiological and behavioural consequences (Huey 1982; Diele-viegas et al. 2018; Domínguez-Guerrero et al. 2019; Fig. 1). For example, reproductive output can be negatively affected when higher environmental temperatures bias the sex ratio of reptile species with temperature-dependent sex determination (Mitchell and Janzen 2010; Jensen et al. 2018). Similarly, changes in local precipitation and temperature can increase eutrophication in aquatic habitats, leading to e.g. elevated nutrient loading, low light levels, spread of pathogenic infections and hypoxic conditions, which in turn can affect growth and survival of amphibians (Johnson et al. 2007; Blaustein et al. 2010; Isaza et al. 2020; Rodgers 2021). Examples such as these, highlight how ectotherms may be more strongly affected and less likely to cope with rapidly changing environments than endotherms due to the direct interaction between external conditions and physiology (Pie et al. 2017; Rolland et al. 2018). In this section we will briefly summarise the diverse ways in which climate change has a direct impact on the ecology, behaviour and ultimately persistence of reptiles and amphibians.
Direct consequences of climate change for amphibians and reptiles. Reptilian and amphibian populations are strongly reliant on their environment and changes in e.g. temperature and precipitation regimes can have many direct physiological and behavioural consequences. These changes can be largely classified across three biological dimensions: (i) thermal regulation, (ii) reproduction and development and (iii) habitat dependency and dispersal, with the latter being considered a consequence of the first two dimensions. The figure illustrates under each category some examples found in the literature
Thermal regulation
Thermoregulation is the process with which organisms maintain their body temperature within an optimal thermal range in order to optimise biological processes, such as enzymatic activity or embryological development (Hutchison and Dupré 1992). In contrast to endotherms, ectotherms cannot thermoregulate internally and thus rely on external heat sources to adjust body temperature and optimise their performance (Huey 1982; Huey and Berrigan 2001; Rolland et al. 2018). Thermal regulation strategies, such as habitat selection or behavioural adjustments, allow ectotherms to counteract reductions in performance and shifts in thermal optima when environmental temperatures are suboptimal (i.e. due to daily and seasonal temperature fluctuations; Huey 1982; Huey and Berrigan 2001; Kearney et al. 2009; Blaustein et al. 2010; Le Galliard et al. 2012; Sunday et al. 2014; Muñoz et al. 2016; Taylor et al. 2021). The thermal range of a given species sets the utmost boundaries in which an individual can operate and maximise performance at an optimal temperature (Huey et al. 2012), and in the case of ectotherms, increases with latitude (Deutsch et al. 2008; Huey et al. 2009). With more variable environments and broader seasonal fluctuations in temperature, ectotherms in temperate regions generally have wider thermal optima and warming tolerances than tropical ectotherms (Snyder and Weathers 1975; Deutsch et al. 2008; Clusella-Trullas et al. 2011; Huey et al. 2012; Quintero and Wiens 2013), which are more sensitive to temperature changes and will reach their critical maximum faster (Deutsch et al. 2008; Moritz et al. 2012). Nonetheless, even among ectotherms in temperate regions, some species are adapted to cool and humid habitats (e.g. montane environments) and may have relatively narrow thermal ranges and low thermal optima (Garcia-Porta et al. 2019).
When variation in environmental conditions cannot be optimally accounted for via thermoregulation, climate change can lead to major differences in physiological performance. Changes in temperature and precipitation regimes, as well as the increased occurrence of extreme meteorological events, can induce significant changes in water and energy balances of ectotherms, such as increased evaporative water loss, as well as changes in their metabolic activity, greatly influencing their performance (Huey et al. 2012). Several ectotherm species depend on hibernation during harsh seasons and maintain their metabolic requirements at very low levels during periods of low ambient temperatures and decreased food availability (Reading 2007). For such species, changes in temperature patterns leading to bouts of higher environmental temperatures during winter might result in increased metabolic requirements caused by cycles of rewarming and metabolic depression (Reading 2007; Blaustein et al. 2010), resulting in lower fitness at time of arousal (Reading 2007). Altered metabolic activities and decreased performance resulting from shifts from thermal optima can negatively affect immune function, making ectotherms more susceptible to infection (Cohen et al. 2017). This may be particularly concerning in light of the widespread fungal infection outbreaks that have affected many amphibian groups (Blaustein et al. 2010, 2012; Cohen et al. 2019).
Reproduction and development
Changes in local environmental conditions can have important direct and indirect consequences for (the timing of) reproduction and development of reptiles and amphibians (Blaustein et al. 2010; Bestion et al. 2015). Breeding time onset is often affected by temperature (Dunn 2004) and an earlier start of the breeding season has been observed for many amphibian and reptilian species in regions where Spring commences earlier (Blaustein et al. 2010; Green 2017). While experimental studies have reported a positive correlation between breeding time onset, body size and reproductive capacity (Beebee 1995; Gibbs and Breisch 2001; Chadwick et al. 2006; Bestion et al. 2015), shifts in breeding time can have negative consequences for populations when competition for resources and habitat takes place between species that normally do not have overlapping breeding seasons (Blaustein et al. 2010). Breeding activity itself can also be negatively affected when climate change leads to variation in the strength and frequency of rainfall events (Blaustein et al. 2010; Akmentins et al. 2015), particularly for species with aquatic life stages and species that rely on ephemeral water bodies. Moreover, precipitation can be an important prerequisite for egg deposition and spawning activity of amphibians (Jensen et al. 2003). Alterations in reproductive behaviour to counteract the effects of climate change can include the adjustment of nest site selection towards cooler or more protected areas and altering nest site construction, in species where nesting depth is modulated (Doody et al. 2006).
Once eggs have been laid, incubation temperature has a direct impact on important life-history traits such as body size, growth rate, body shape, locomotor performance, thermoregulatory behaviour and sexual phenotype (Noble et al. 2018; Singh et al. 2020). For example, swimming performance and behaviour of hatchlings of the Mary River turtle, Elusor macrurus, can vary with incubation temperature. Individuals that are incubated at cooler temperatures have higher performance and swim more than turtles that are incubated at warmer temperatures (Micheli-Campbell et al. 2011). Most reptilian species lay eggs, with embryos that cannot thermoregulate (Cordero et al. 2018; Singh et al. 2020) and while warmer temperatures may accelerate embryonic growth and hatching time (Bestion et al. 2015; Singh et al. 2020), high temperatures during early stages of embryological development might negatively affect cell, tissue and organ differentiation (Singh et al. 2020). Increased temperatures during development might also accelerate larval growth rate in amphibians and lead to increased body size at the moment of metamorphosis (Peng et al. 2020). Incubation temperature does not only influence development itself, but can even determine offspring sex. There are many reptilian species, such as crocodiles and some turtles, where offspring sex is determined by incubation temperature. An increase in environmental temperature can therefore lead to a significant population bias in sex ratios (Hulin et al. 2009; Mitchell and Janzen 2010; Simoncini et al. 2014; Valenzuela et al. 2019). A skew in sex ratios can increase mate competition and negatively affect reproductive output (Doody et al. 2006; Blaustein et al. 2010; Singh et al. 2020). Rising sand temperature in breeding locations of green turtles, Chelonia mydas, for example, has led to an increased feminization of two populations on the Australian Great Barrier Reef during the last two decades; a sex ratio bias that could potentially lead to the complete feminization of these populations in the near future (Jensen et al. 2018). However, interestingly, some species with temperature-dependent sex determination seem to be able to shift their breeding onset as a behavioural reproductive strategy to compensate for biased sex ratios. These species can adjust the onset of reproduction towards temperatures that favour the production of the less frequent sex (Doody et al. 2006), but it remains unclear to what extent they can compensate.
Habitat dependency and dispersal capacity
The climatic niche, the range of environmental conditions in which a species can occur and persist (Pie et al. 2017; Wollenberg Valero et al. 2019), is generally more restricted for ectotherms than endotherms (Huey 1991; Pie et al. 2017; Rolland et al. 2018). Their dependence on external conditions for thermoregulation, reproduction and development frequently limits their physiological capacity to adopt a broad climatic niche and restricts their ability to disperse across non-suitable habitat and, while behavioural changes may partially alleviate, phenotypic plasticity can only mitigate such negative consequences to a certain extent (Muñoz et al. 2016; Domínguez-Guerrero et al. 2019). The low dispersal capacity of many amphibian and reptile species has likely contributed to an increase in ecological specialisation (Jocque et al. 2010) and while ectotherms globally occupy a wide diversity of ecological niches, many species are highly specialised to local conditions. Tropical regions in particular exhibit high degrees of species endemism and distinct species, which only marginally differ in ecological niche, can frequently be found in close proximity (Jocque et al. 2010; Oliver et al. 2019).
Ecological specialisation to highly localised conditions coupled with low-dispersal capacity increases reliance on habitat stability and rapid changes in climate may pose a considerable threat to ectotherms that are small range endemics. Additionally, the increased spread of invasive species facilitated by climate change, coupled with habitat suitability loss, might introduce further challenges, such as competition for resources or exposure to foreign pathogens (Falaschi et al. 2019; Pabijan et al. 2020). Whereas ecological generalists may be able to track suitable habitat during periods of change, ecological specialists are locally trapped and may go extinct with the disappearance of microhabitat (Moussalli et al. 2009; Garcia-Porta et al. 2019; Nguyen et al. 2019). Ectotherms may therefore be particularly dependent on geographic regions that have served as evolutionary refugia in the past and such refugia have likely shaped diversity patterns at a macroevolutionary scale for many reptiles and amphibians (Hewitt 1996, 2008; Schneider and Moritz 1999; Beheregaray 2008; Byrne et al. 2008; Carnaval et al. 2009; Moussalli et al. 2009; Noble et al. 2018; Leaché et al. 2019). Refugia where ectotherms have persisted during climate change can often be characterised as heterogeneous landscapes, in terms of geography, that provide both shelter and an opportunity for the persistence of microclimatic conditions that differ from the surrounding area. For example, local gorges where small water bodies remain can retain humidity and buffer temperatures, providing refuge for amphibian and reptile populations during past periods of aridification in the southern hemisphere (Byrne et al. 2008). Linking the study of past demography with habitat suitability over time can therefore yield important insights on the location and characteristics of regions where populations may potentially persist during contemporary changes in climate.
The role of climate in shaping demographic history
There are now ample examples that demonstrate how changes in climate may have considerable consequences for contemporary ectotherm populations, but a more detailed view on past population dynamics can help to understand how these proximate causes ultimately shape long-term demographic change. For example, it is important to study how climate-induced variation in traits such as thermal regulation, reproduction and development dovetail with demographic changes or stasis during Pleistocene periods of climatic instability. By looking at the past, we can evaluate the relative importance of these key traits for population persistence and decline in an empirical context and use these insights to improve models that predict future population responses to climate change (Brown et al. 2016; Waldvogel et al. 2020). However, even though terrestrial ectotherms are among the most well studied animal groups in terms of systematics and taxonomy, studies that have modelled demographic history remain relatively scarce for most amphibians and reptiles.
Herpetologists have to date, largely focused on the distribution and phylogenetic history of closely related species in a (bio)geographic context (Carnaval et al. 2009; Moussalli et al. 2009; Camargo et al. 2010; Pepper et al. 2011; Edwards et al. 2016, 2022). Phylogeographic studies have improved our understanding on the broad scale evolutionary history of many reptilian and amphibian species and have emphasised the importance of climate in shaping the long-term diversity of ectotherms (Dolman and Moritz 2006; Moussalli et al. 2009; Chapple et al. 2011; Moritz et al. 2012; Singhal and Moritz 2013; Melville et al. 2016; Hawlitschek et al. 2017; Potter et al. 2018; Ansari et al. 2019; Dinis et al. 2019; Dufresnes et al. 2020; Leaché et al. 2020; Ledo et al. 2020; Jaynes et al. 2021). For instance, the distribution of phylogenetic diversity across the landscape has been used to identify geographic regions of long-term evolutionary persistence (refugia) and their role in promoting speciation. Such methods are particularly powerful in a comparative framework, when phylogeographic patterns across many species are used to identify regions with a relatively high proportion of unique lineages (endemism; Dolman & Moritz 2006; Moussalli et al. 2009; Fujita et al. 2010; Rosauer et al. 2018).
Phylogeographic inference has been an important tool to study the long-term consequences of climatic variation on ectotherm diversification, but may be less suited to understand the interaction between recent shifts in climatic variables, phenotypic responses and associated changes in demography for specific populations and species. Phylogeographic methods largely depend on nucleotide substitutions, for example fixed differences between populations, and are therefore less accurate when studying more recent changes in population size (Edwards et al. 2016, 2022). However, major changes in climate have occurred since the Pleistocene and have certainly affected many ectotherm populations to varying degrees (Araújo et al. 2008). Understanding to what extent climate dependent fitness traits and changes in climatic variables directly drive demographic change, is an important prerequisite to accurately estimate the resilience of individual populations and species under current models of future change. We will therefore conclude this Review with a brief primer into demographic inference, give an overview of the type of data that is commonly used for terrestrial ectotherms to infer demography, discuss opportunities and challenges of these frequently used approaches and summarise how recent technological advances in sequencing methodology now set the stage for studying demographic history across a broader temporal continuum in many ectotherm groups.
Inferring population dynamics using genome-reduction data
To predict how contemporary changes in climate may affect extant populations, the study of long-term processes that shape phylogeographic diversity needs to be coupled with in-depth analyses that evaluate how more recent (e.g. < 1 MYA) variation in climate have led to demographic change. Traditionally, population parameters such as effective population size, population connectivity and divergence times, were estimated based on a small number of loci and quantified by comparing how a given summary statistic deviated from an equilibrium model (Tollis et al. 2012; Salmona et al. 2019). These estimates could vary substantially due to the widespread variation in coalescent rates between loci (Salmona et al. 2019). The accuracy of demographic parameter estimation has benefited substantially from the introduction of high-throughput sequencing (HTS) with recent models based on thousands of independently evolving loci (Beichman et al. 2018).
With large-scale genomic datasets emerging, many recent demographic studies in ectotherms have largely relied on changes in the site-frequency spectrum (SFS) to model population dynamics. The SFS is the distribution of allele frequencies at polymorphic sites and is calculated for a given population of interest (Salmona et al. 2019; Bourgeois and Warren 2021). Changes in effective population size will influence the expected distribution of allele frequency counts and therefore the overall shape of the SFS (Bourgeois and Warren 2021). For example, under a neutral evolution model, an expanding population has a higher number of low-frequency variants, compared to a population of constant size. In contrast, a population that has recently undergone a bottleneck will have a deficiency in low-frequency variants. Moreover, correlations between the SFS of individual populations can be levied to quantify the degree of shared history (i.e. 2D-SFS or joint-SFS; Salmona et al. 2019). Model based inference (Sousa and Hey 2013) can be employed to compare the observed SFS to an expected SFS (e.g. based on coalescent simulations) and identify a population model that best fits the data. Likelihood-based methods for model comparison, such as Fastsimcoal2 (Excoffier et al. 2013) and dadi (Gutenkunst et al. 2009), are popular but can be computationally taxing when the number of possible models is large and parameter estimation needed. Approximate Bayesian Computation (ABC, Beaumont et al. 2002) provides another framework for model comparison and is a versatile approach that can utilise various kinds of summary-statistics (e.g. Fst). More recent approaches, such as machine learning algorithms like random forest, include training simulations to predict the best model that fits the observed data. Random forest approaches can be used within ABC frameworks and significantly reduce computing effort (Pudlo et al. 2016; Schrider and Kern 2018).
To date, demographic inference in ectotherms has largely focused on methods that evaluate changes in the SFS or other summary-statistics, due to a general lack in genomic resources for most amphibians and reptiles. Genome reduction approaches, such as double-digest RAD-sequencing (ddRAD; Peterson et al. 2012) or sequence-capture (Faircloth et al. 2012; Lemmon et al. 2012; Jones and Good 2016), have provided an effective alternative to generate large-scale datasets of orthologous loci when reference genomes are lacking or when genome sizes are large (as is the case for some amphibian species). RAD-sequencing methods use cut-site specific restriction enzymes and orthologous fragments can be retrieved when the distribution of cut-sites across the genome is relatively similar between populations (Baird et al. 2008; Harvey et al. 2016). Sequence-capture methods target conserved loci of the genome (such as exons or ultra-conserved elements) across close or distantly related taxa (Harvey et al. 2016). Genome reduction approaches yield 100 s to 1000 s of unlinked orthologous loci which, in comparison to traditional sequencing approaches, increases the statistical power to differentiate between models and benefits the accuracy of parameter estimation. For instance, while a few hundred unlinked loci may suffice to differentiate between distinct demographic scenarios, such as population expansion or contraction (Prates et al. 2016; Afonso Silva et al. 2017; Potter et al. 2018), large numbers of loci may be needed for complex multi-population models or when simultaneously quantifying population parameters such as divergence time, population size or gene flow (Portik et al. 2017; Nguyen et al. 2019; Farleigh et al. 2021; Thörn et al. 2021; Wen and Fu 2021). By coupling genome-reduction with model-based inference, demographic studies have been able to test the directionality of colonisation (expansion) into different geographical regions, such as the expansion of populations of the common toad, Bufo bufo, in Sweden (Thörn et al. 2021). Moreover, these studies have been able to quantify the symmetry and amount of gene flow between populations and estimate general changes in population size, as in the study of Wen and Fu (2021), where populations of green odorous frogs, Odorrana margaretae, from western China have come into secondary contact at several points along their distribution after initial isolation. In addition, comparative methods that quantify the degree of synchrony in demographic dynamics between populations have been developed (Xue and Hickerson 2017). This is especially relevant for co-distributed populations that have diversified or persisted under similar climatic or ecological conditions, such as Anolis species found in the Amazonian and Atlantic Forest, for which idiosyncratic responses to common change in climate have been found (Prates et al. 2016). This also reiterates the importance of studying these dynamics at a species- or lineage-specific level. However, notwithstanding their important contribution in advancing our understanding of past population dynamics, it is worthwhile to recognize the limitations of SFS or summary-statistic based inference and discuss how other data types and inference approaches may mitigate some of these challenges.
A genome-wide view on demographic history
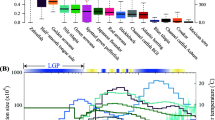
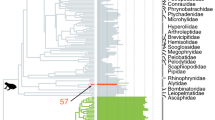
While model-based inference using SFS or summary-statistics is well suited to study very recent changes in population history, these approaches generally tend to be less accurate when inferring demographic dynamics over longer time scales (Fig. 2). The inference of older dynamics can be confounded when recent demographic history overrides previous changes in the SFS and thus leads to the saturation of sites in the SFS relative to more ancient processes (Liu and Fu 2015; Lapierre et al. 2017; Patton et al. 2019). To study demographic history over longer time scales, a distinct suite of methods that model the temporal distribution of coalescent histories may be more appropriate. For example, a recent empirical evaluation by Patton et al. (2019) demonstrated how sequentially Markovian coalescent (SMC) approaches are more accurate when inferring demographic patterns between 300 and one hundred thousand generations before present (gbp) whereas SFS based approaches are more reliable up to 30 gbp. In other words, coalescent based methods such as the SMC and allied approaches can complement methods that rely on the SFS alone and represent an important suite of tools that are more useful for studying demographic dynamics in ectotherms across the Pleistocene.
Inferring evolutionary history along the “phylogeographic-demographic” continuum. Data types and inference approaches vary in their suitability for inferring evolutionary history across time. Gradient lines under data types and methods roughly illustrate where each type of molecular marker or method excels. While an approximate timeline is provided, exact boundaries depend on generation time, effective population size, mutation rate, etc. Important to note is that methods based on coalescent based models (marked with * in the figure), such as Relate, MSMC, PSMC and SMC + + , require whole-genome data as input, and for most of these methods, high quality genomic data and the availability of a good reference genome is required. Finally, the Analyses section illustrates the most common inference outcomes for each of those models. For a more detailed overview of population genetics methods refer to Bourgeois and Warren (2021)
SMC approaches are based on coalescent theory which is a statistical framework that can relate changes in population size to the frequency of coalescent events over time (Kingman 1982; Mather et al. 2020). The rate of coalescence, when two lineages/haplotypes combine to become one ancestral lineage, is inversely correlated to population size under a neutral model since coalescent events are more likely to occur when populations are small. Since recombination breaks linkage between loci, each diploid genome contains many independently evolving tracts and the coalescent time of two alleles within each tract can vary. SMC based methods use a hidden-Markov model (HMM) to work their way across genomes, identify recombination events and record the distribution of coalescent times at a genome-wide scale. For diploid species with recombining genomes, a single genome can already contain a large number of individually evolving tracts and the corresponding coalescent variation between tracts can be used to infer demographic change over time. The Pairwise Sequentially Markovian Coalescent (PSMC; Li & Durbin 2011) approach uses single (unphased) genomes for estimating past demography and has now been widely used in a diverse array of organisms (Mather et al. 2020) for which whole-genome (resequencing) data is available. With only two alleles from a population sampled, the accuracy of inferring recent change is often limited however due to a lack of coalescent events closer to the present (Beichman et al. 2018; Mather et al. 2020). The Multiple Sequence Markovian Coalescent (MSMC/2) partially bypasses this by enabling the inclusion of multiple genomes but requires genomes to be phased (Schiffels and Durbin 2014). While the use of multiple genomes improves the accuracy of inferring size changes in the very recent past, accurate phasing can be challenging in non-model organisms and the method itself is computationally demanding which limits the number of genomes that can be simultaneously analysed. Since the introduction of PSMC and MSMC, mitigating these challenges has been a major area of development. For example, SMC + + (Terhorst et al. 2017) uses both linkage information (coalescent HMM) as well as the SFS, to accurately model demographic change across a longer time span. Moreover, SMC + + allows the use of many unphased genomes, limiting the effects of phasing errors, and can estimate recent effective population sizes with higher resolution than MSMC and PSMC (if sufficient samples available; Patton et al. 2019). Other promising approaches still require data to be phased, but can now scale up to thousands of samples (e.g. TSinfer, Kelleher et al. 2019; Relate, Speidel et al. 2019). All the aforementioned methods (SFS-based, ABC and SMC) assume that populations are neutrally evolving, however discerning between natural selection and demographic events can be challenging, as both processes can lead to similar patterns of genetic variation (Li and Durbin 2011; Schrider and Kern 2018; Beichman et al. 2018; Marchi et al. 2021). This may be particularly important for organisms such as amphibians and reptiles that have faced strong selective pressures in a scenario of climate change. Genome-wide datasets are beneficial in such instances because they allow for the exclusion of loci that may be under selection and focus on neutrally evolving loci to infer demographic history. For example, loci in low-recombination regions can be filtered out since they are often undergoing linked selection and exons may be less suited than intergenic regions (Beichman et al. 2018). Notwithstanding known challenges of accurately inferring demographic history due to confounding biological processes (e.g. population structure, selective sweeps, migration after divergence, etc. Schrider and Kern 2018; Beichman et al. 2018; Salmona et al. 2019; Sellinger et al. 2020, 2021; Spence et al. 2018), the use of SMC based methods has taken flight in recent years with the increasing availability of genome-wide datasets and complements our ability to accurately infer demographic history across a broader range of time scales.
For those species where whole-genome data is available, birds and mammals in particular, SMC and allied methods have been widely used to study the demographic consequences of environmental change. For example, SMC approaches have been used to estimate fluctuations in effective population size in relation to glacial cycles throughout the Quaternary (brown bears, Ursus arctos and polar bears, Ursus maritimus: Miller et al. 2012; birds: Nadachowska-Brzyska et al. 2015; Natesh et al. 2020; Eurasian lynx, Lynx lynx: Lucena-Perez et al. 2020; African golden wolves, Canis lupaster: Sarabia et al. 2020; bottlenose dolphins, Tursiops truncatus and Tursiops aduncus: Vijay et al. 2018; humans: Li and Durbin 2011). Contrasting population trajectories can reveal differences in the capacity of species to couple with local environmental change, even among closely related lineages, such as in the case of bottlenose dolphins (Vijay et al. 2018), African golden wolves (Sarabia et al. 2020) and walnut trees (Bai et al. 2018). These idiosyncratic responses can be further investigated to evaluate the role of ecological specialisation in mitigating climatic change, as Natesh et al. (2020) for example observed in their study on two Eurasian owlet species, Athene brama and Glaucidium radiatum, where the population size of species associated to open habitats followed the expansion and contraction of their habitat until the Mid-Holocene, whereas forest adapted species were not affected by climate-induced changes in vegetation. Similarly, geographic history and ecological specialisation have very likely played a role in the demographic history and diversification of Anolis lizards in Cuba, in which idiosyncratic trends in population size changes between ecomorphs and between thermal habitat specialists are likely due to habitat change caused by the varying timing of inundation of lowland areas during the mid-Pleistocene (Kanamori et al. 2022). Moreover, SMC methods have also proven to be highly valuable when studying the role of changing climate in historic population fluctuations of threatened and extinct taxa (Leitwein et al. 2020) where sample size is limited, such as the extinct woolly mammoth, Mammuthus primigenius (Palkopoulou et al. 2015). Studies modelling historical demographic responses to past climate based on genome-wide linkage and coalescent information have demonstrated their potential, but their uptake for quantifying demographic change in ectotherms remains limited to date due to a lack of high-quality reference genomes.
Reptiles and amphibians are among the most species rich groups of vertebrates but the relative number of species with a sequenced and annotated genome likely remains below 1% (Hotaling et al. 2021). While the availability of high-quality genomes is scarce across many vertebrate groups, the lack of genomic resources for reptiles and amphibians is particularly glaring given their extraordinary richness in species number and evolutionary diversity (Card et al. 2023). This scarcity in reference genomes can partially be attributed to the technical challenge of assembling reptilian and amphibian genomes. For example, amphibians tend to have larger genomes than most vertebrates and on average have a higher proportion of repetitive sequences which complicates assembly (Canapa et al. 2016). However, genome size actually varies a lot between amphibians (Liedtke et al. 2018) and, while the assembly of the 32 Gb. axolotl genome has been a major challenge (Smith et al. 2019), some amphibians have genome sizes that are relatively modest (e.g. the 1 Gb. genome of the ornate burrowing frog Platyplectrum ornatum; Lamichhaney et al. 2021). Moreover, the archetypal squamate genome is less than double in size (1.3–2.8 Gb; Pasquesi et al. 2018) of the average avian genome and smaller than most mammalian genomes. Nonetheless, to date, there are still only 59 amphibian and 130 reptilian genomes publicly available in NCBI (https://www.ncbi.nlm.nih.gov/genome last accessed on September 5th, 2023), in comparison to more than 760 birds and 720 mammals, and many remain highly fragmented and/or unannotated.
Powered by technological development and large-scale sequencing initiatives, such as the Earth BioGenome Project, the Genome 10 K consortium, the European Reference Genome Atlas and the Vertebrate Genomes Project, the number of high-quality amphibian and reptilian genomes will soon be a better reflection of the underlying species diversity among terrestrial vertebrates. Long-read sequencing (LRS) platforms, such as Oxford nanopore technologies (ONT) and Pacific biosciences (PacBio), now generate reads that can span some of the most complex genomic regions (Mao and Zhang 2022) and methods to infer chromatin-interaction maps (e.g. Hi-C, Lieberman-Aiden et al. 2009) can be used to construct chromosome models. These approaches offer a cost-efficient way to assemble larger, more heterogeneous and repetitive genomes and can now even be effectively accomplished by a single research group (rather than large genome consortia). For example, recently published chromosome-scale reference genomes for ectotherms include the brown anole, Anolis sagrei (Geneva et al. 2022), the desert horned lizard, Phrynosoma platyrhinos (Koochekian et al. 2022), the western spadefoot toad, Pelobates cultripes (Liedtke et al. 2022), two caecilian species: the Gaboon caecilian, Geotrypetes seraphini and Microcaecilia unicolor (Ovchinnikov et al. 2023) and two lizard species that are extinct-in-the-wild: the Christmas Island blue-tailed skink, Cryptoblepharus egeriae, and Lister’s gecko, Lepidodactylus listeri (Dodge et al. 2023). The increasing adoption of LRS accelerates the generation of reference genomes, which will enable the use of a wider suite of demographic analyses and eventually benefit the accuracy of demographic inference. Increasing read lengths will ease the ability to phase data and reconstruct both haplotypes of the same chromosome in diploid organisms. LRS has the potential to reduce the error associated with statistical phasing and will therefore improve demographic studies that depend on linkage disequilibrium (LD) and recombination estimation. Thus, the simultaneous development of inference methods that model coalescent variation at a genome wide scale, the integration of sequencing technologies that ease the generation of high-quality genomes and phased resequencing data, will allow us to study the role of climate in driving population dynamics in reptiles and amphibians across the Pleistocene in an unprecedented detail. These analyses will complement our understanding of how recent climate-induced changes in population dynamics dovetail with macroevolutionary (phylogeographic) patterns and will ultimately allow us to estimate how climate dependent fitness traits have shaped demographic change during recent changes in climate.
Conclusion and perspective
Climate change will have far reaching consequences for Earth’s biodiversity and the climate change resilience of amphibians and reptiles is of particular concern. In this review, we have synthesized the myriad of ways that environmental change can have a direct impact on terrestrial ectotherm populations and discuss their potential vulnerability in the face of future climate change. Studying how amphibians and reptiles have responded to changing conditions in the past, is a promising avenue to identify how these traits and climatic variation have ultimately shaped demographic history throughout the Pleistocene. However, while the evolutionary consequences of climate change have largely been studied in a phylogeographic context, the number of studies focusing on demographic change remains relatively limited. With an increasing ease to assay genetic variation at a genome-wide scale, we then provided a short primer on the most frequently used methods to generate and utilise genomic data for demographic inference and relate this to empirical work presented thus far. One major limitation that has prevented the uptake of such approaches is the relative lack of genomic resources for many amphibian and reptiles. Given their sensitivity to environmental perturbations, we therefore call for a more balanced focus among vertebrates and encourage the much-needed development of genomic resources for almost all ectotherm groups. This effort will greatly benefit from recent advances in long-read sequencing and even modest sized research groups can now produce high-quality assemblies for many amphibian and reptilian species. By both emphasizing the marked sensitivity of ectotherms to climate change, as well as the need for detailed studies quantifying demographic history, we have aimed to bridge the gap between herpetologists and population geneticists, and spur future studies into the demographic history of species groups that to date have been relatively neglected in comparison to other terrestrial vertebrates.
A high-resolution model of past population dynamics throughout the Pleistocene can then be coupled with recent estimates of climatic variation over time and space. Ecological Niche Modelling and paleoclimate data are now frequently combined to estimate the distribution of suitable habitat over time and are continuously improving with the integration of more sophisticated model prediction algorithms (e.g. Machine Learning). Traditionally, there has been a temporal mismatch between paleoclimate modelling and the time scale where phylogeographic studies excel. Contemporary demographic models are better suited to model population change during the same time span and increase our ability to identify biologically relevant correlations between environmental variables and changes in population dynamics. We argue that ectotherms are particularly suited for such investigations due to their direct physiological dependence on external conditions and sensitivity to environmental perturbations. Simultaneously, such empirically informed models are urgently needed to predict how contemporary populations of amphibians and reptiles will ultimately fare in the rapidly changing environment of the twenty-first century.
References
Afonso Silva AC, Bragg JG, Potter S, Fernandes C, Coelho MM, Moritz C (2017) Tropical specialist vs. climate generalist: diversification and demographic history of sister species of Carlia Skinks from Northwestern Australia. Mol Ecol 26(15):4045–4058. https://doi.org/10.1111/mec.14185
Akmentins MS, Pereyra LC, Sanabria EA, Vaira M (2015) Patterns of daily and seasonal calling activity of a direct-developing frog of the subtropical Andean forests of Argentina. Bioacoustics 24(2):89–99. https://doi.org/10.1080/09524622.2014.965217
Ansari MH, Cooper SJB, Schwarz MP, Ebrahimi M, Dolman G, Reinberger L, Saint KM, Donnellan SC, Bull CM, Gardner MG (2019) Plio-pleistocene diversification and biogeographic barriers in Southern Australia reflected in the phylogeography of a widespread and common lizard species. Mol Phylogenet Evol 133:107–119. https://doi.org/10.1016/j.ympev.2018.12.014
Araújo MB, Rahbek C (2006) How does climate change affect biodiversity? Science 313(5792):1396–1397. https://doi.org/10.1126/science.1131758
Araújo MB, Thuiller W, Pearson RG (2006) Climate warming and the decline of amphibians and reptiles in Europe. J Biogeogr 33(10):1712–1728. https://doi.org/10.1111/j.1365-2699.2006.01482.x
Araújo MB, Nogués-Bravo D, Diniz-Filho JAF, Haywood AM, Valdes PJ, Rahbek C (2008) Quaternary climate changes explain diversity among reptiles and amphibians. Ecography 31(1):8–15. https://doi.org/10.1111/j.2007.0906-7590.05318.x
Bai WN, Yan PC, Zhang BW, Woeste KE, Lin K, Zhang DY (2018) Demographically idiosyncratic responses to climate change and rapid pleistocene diversification of the walnut genus Juglans (Juglandaceae) revealed by whole-genome sequences. N Phytol 217(4):1726–1736. https://doi.org/10.1111/nph.14917
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA, Selker EU, Cresko WA, Johnson EA (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3(10):1–7. https://doi.org/10.1371/journal.pone.0003376
Beaumont MA, Zhang W, Balding DJ (2002) Approximate Bayesian computation in population genetics. Genetics 162(4):2025–2035. https://doi.org/10.1093/genetics/162.4.2025
Beebee TJC (1995) Amphibian breeding and climate. Nature 374(6519):219–220. https://doi.org/10.1038/374219a0
Beheregaray LB (2008) Twenty years of phylogeography: the state of the field and the challenges for the Southern Hemisphere. Mol Ecol 17(17):3754–3774. https://doi.org/10.1111/j.1365-294X.2008.03857.x
Beichman AC, Huerta-Sanchez E, Lohmueller KE (2018) Using genomic data to infer historic population dynamics of nonmodel organisms. Annu Rev Ecol Evol Syst 49:433–456. https://doi.org/10.1146/annurev-ecolsys-110617-062431
Bestion E, Teyssier A, Richard M, Clobert J, Cote J (2015) Live fast, die young: experimental evidence of population extinction risk due to climate change. PLoS Biol 13(10):1–19. https://doi.org/10.1371/journal.pbio.1002281
Bickford D, Lohman DJ, Sodhi NS, Ng PKL, Meier R, Winker K, Ingram KK, Das I (2007) Cryptic species as a window on diversity and conservation. Trends Ecol Evol 22(3):148–155. https://doi.org/10.1016/j.tree.2006.11.004
Blaustein AR, Belden LK, Olson DH, Green DM, Root TL, Kiesecker JM (2001) Amphibian breeding and climate change. Conserv Biol 15(6):1804–1809. https://doi.org/10.1046/j.1523-1739.2001.00307.x
Blaustein AR, Walls SC, Bancroft BA, Lawler JJ, Searle CL, Gervasi SS (2010) Direct and indirect effects of climate change on amphibian populations. Diversity 2(2):281–313. https://doi.org/10.3390/d2020281
Blaustein AR, Gervasi SS, Johnson PTJ, Hoverman JT, Belden LK, Bradley PW, Xie GY (2012) Ecophysiology meets conservation: understanding the role of disease in amphibian population declines. Philos Trans R Soc Lond B Biol Sci 367(1596):1688–1707. https://doi.org/10.1098/rstb.2012.0011
Bourgeois YXC, Warren BH (2021) An overview of current population genomics methods for the analysis of whole-genome resequencing data in eukaryotes. Mol Ecol 30(23):6036–6071. https://doi.org/10.1111/mec.15989
Brown JL, Weber JJ, Alvarado-Serrano DF, Hickerson MJ, Franks SJ, Carnaval AC (2016) Predicting the genetic consequences of future climate change: the power of coupling spatial demography, the coalescent, and historical landscape changes. Am J Bot 103(1):153–163. https://doi.org/10.3732/ajb.1500117
Butler CJ (2019) A review of the effects of climate change on chelonians. Diversity 11(8):138. https://doi.org/10.3390/d11080138
Byrne M, Yeates DK, Joseph L, Kearney M, Bowler J, Williams MAJ, Cooper S, Donnellan SC, Keogh JS, Leys R, Melville J, Murphy DJ, Porch N, Wyrwoll KH (2008) Birth of a biome: insights into the assembly and maintenance of the Australian arid zone biota. Mol Ecol 17(20):4398–4417. https://doi.org/10.1111/j.1365-294X.2008.03899.x
Camargo A, Sinervo B, Sites JW (2010) Lizards as model organisms for linking phylogeographic and speciation studies. Mol Ecol 19(16):3250–3270. https://doi.org/10.1111/j.1365-294X.2010.04722.x
Canapa A, Barucca M, Biscotti MA, Forconi M, Olmo E (2016) Transposons, genome size, and evolutionary insights in animals. Cytogenet Genome Res 147(4):217–239. https://doi.org/10.1159/000444429
Card DC, Jennings WB, Edwards SV (2023) Genome evolution and the future of phylogenomics of non-avian reptiles. Animals 13(3):471. https://doi.org/10.3390/ani13030471
Carnaval AC, Hickerson MJ, Haddad CFB, Rodrigues MT, Moritz C (2009) Stability predicts genetic diversity in the Brazilian Atlantic forest hotspot. Science 323(5915):785–789. https://doi.org/10.1126/science.1166955
Chadwick EA, Slater FM, Ormerod SJ (2006) Inter- and intraspecific differences in climatically mediated phenological change in coexisting Triturus species. Glob Chang Biol 12(6):1069–1078. https://doi.org/10.1111/j.1365-2486.2006.01156.x
Chapple DG, Hoskin CJ, Chapple SNJ, Thompson MB (2011) Phylogeographic divergence in the widespread delicate skink (Lampropholis Delicata) corresponds to dry habitat barriers in eastern Australia. BMC Evol Biol 11(1):191. https://doi.org/10.1186/1471-2148-11-191
Clusella-Trullas S, Blackburn TM, Chown SL (2011) Climatic predictors of temperature performance curve parameters in ectotherms imply complex responses to climate change. Am Nat 177(6):738–751. https://doi.org/10.1086/660021
Cohen JM, Venesky MD, Sauer EL, Civitello DJ, McMahon TA, Roznik EA, Rohr JR (2017) The thermal mismatch hypothesis explains host susceptibility to an emerging infectious disease. Ecol Lett 20(2):184–193. https://doi.org/10.1111/ele.12720
Cohen JM, Civitello DJ, Venesky MD, McMahon TA, Rohr JR (2019) An interaction between climate change and infectious disease drove widespread amphibian declines. Glob Chang Biol 25(3):927–937. https://doi.org/10.1111/gcb.14489
Collins M, Knutti R, Arblaster J, Dufresne JL, Fichefet T, Friedlingstein P, Gao X, Gutowski WJ, Johns T, Krinner G, Shongwe M, Tebaldi C, Weaver AJ, Wehner M (2013) Long-term climate change: projections, commitments and irreversibility. In: Stocker TF, Qin D, Plattner GK, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds) Climate change 2013: the physical science basis contribution of working group i to the fifth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge
Cordero GA, Telemeco RS, Gangloff EJ (2018) Reptile embryos are not capable of behavioral thermoregulation in the egg. Evol Dev 20(1):40–47. https://doi.org/10.1111/ede.12244
Davis MB, Shaw RG, Etterson JR (2005) Evolutionary responses to changing climate. Ecology 86(7):1704–1714. https://doi.org/10.1890/03-0788
Deutsch CA, Tewksbury JJ, Huey RB, Sheldon KS, Ghalambor CK, Haak DC, Martin PR (2008) Impacts of climate warming on terrestrial ectotherms across latitude. Proc Natl Acad Sci USA 105(18):6668–6672. https://doi.org/10.1073/pnas.0709472105
Diele-viegas LM, Frederico C, Rocha D (2018) Unraveling the in Fl Uences of climate change in Lepidosauria (Reptilia). J Therm Biol 78:401–414. https://doi.org/10.1016/j.jtherbio.2018.11.005
Dinis M, Merabet K, Martínez-Freiría F, Steinfartz S, Vences M, Burgon JD, Elmer KR, Donaire D, Hinckley A, Fahd S, Joger U, Fawzi A, Slimani T, Velo-Antón G (2019) Allopatric diversification and evolutionary melting pot in a North African palearctic relict: the biogeographic history of Salamandra algira. Mol Phylogenet Evol 130:81–91. https://doi.org/10.1016/j.ympev.2018.10.018
Dodge TO, Farquharson KA, Ford C, Cavanagh L, Schubert K, Schumer M, Belov K, Hogg CJ (2023) Genomes of two Extinct-in-the-Wild reptiles from Christmas Island reveal distinct evolutionary histories and conservation insights. Mol Ecol Res 00:1–17. https://doi.org/10.1111/1755-0998.13780
Dolman G, Moritz C (2006) A multilocus perspective on refugial isolation and divergence in rainforest skinks (Carlia). Evolution 60(3):573. https://doi.org/10.1111/j.0014-3820.2006.tb01138.x
Domínguez-Guerrero SF, Muñoz MM, de Pasten-Téllez D, Jesús AM, Diego M, Rodríguez-Miranda LA, Manríquez-Morán NL, Méndez-de la Cruz FR (2019) Interactions between thermoregulatory behavior and physiological acclimatization in a wild lizard population. J Therm Biol 79:135–143. https://doi.org/10.1016/j.jtherbio.2018.12.001
Doody JS, Guarino E, Georges A, Corey B, Murray G, Ewert M (2006) Nest site choice compensates for climate effects on sex ratios in a lizard with environmental sex determination. Evol Ecol 20(4):307–330. https://doi.org/10.1007/s10682-006-0003-2
Dufresnes C, Nicieza AG, Litvinchuk SN, Rodrigues N, Jeffries DL, Vences M, Perrin N, Martínez-Solano I (2020) Are glacial refugia hotspots of speciation and cytonuclear discordances? Answers from the genomic phylogeography of Spanish common frogs. Mol Ecol 29(5):986–1000. https://doi.org/10.1111/mec.15368
Dunn P (2004) Breeding dates and reproductive performance. Adv Ecol Res 35(4):69–87. https://doi.org/10.1016/S0065-2504(04)35004-X
Edwards SV, Potter S, Schmitt CJ, Bragg JG, Moritz C (2016) Reticulation, divergence, and the phylogeography-phylogenetics continuum. Proc Natl Acad Sci USA 113(29):8025–8032. https://doi.org/10.1073/pnas.1601066113
Edwards SV, Robin VV, Ferrand N, Moritz C (2022) The evolution of comparative phylogeography: putting the geography (and More) into comparative population genomics. Genome Biol Evol 14(1):1–16. https://doi.org/10.1093/gbe/evab176
Excoffier L, Dupanloup I, Huerta-Sánchez E, Sousa VC, Foll M (2013) Robust demographic inference from genomic and SNP data. PLoS Genet. https://doi.org/10.1371/journal.pgen.1003905
Faircloth BC, McCormack JE, Crawford NG, Harvey MG, Brumfield RT, Glenn TC (2012) Ultraconserved elements anchor thousands of genetic markers spanning multiple evolutionary timescales. Syst Biol 61(5):717–726. https://doi.org/10.1093/sysbio/sys004
Falaschi M, Manenti R, Thuiller W, Ficetola GF (2019) Continental-scale determinants of population trends in European amphibians and reptiles. Glob Chang Biol 25(10):3504–3515. https://doi.org/10.1111/gcb.14739
Farleigh K, Vladimirova SA, Blair C, Bracken JT, Koochekian N, Schield DR, Card DC, Finger N, Henault J, Leaché AD, Castoe TA, Jezkova T (2021) The effects of climate and demographic history in shaping genomic variation across populations of the desert horned lizard (Phrynosoma Platyrhinos). Mol Ecol 30(18):4481–4496. https://doi.org/10.1111/mec.16070
Fujita MK, McGuire JA, Donnellan SC, Moritz C (2010) Diversification and persistence at the arid-monsoonal interface: Australia-wide biogeography of the Bynoe’s Gecko (Heteronotia Binoei; Gekkonidae). Evolution 64(8):2293–2314. https://doi.org/10.1111/j.1558-5646.2010.00993.x
Garcia-Porta J, Irisarri I, Kirchner M, Rodríguez A, Kirchhof S, Brown JL, MacLeod A, Turner AP, Ahmadzadeh F, Albaladejo G, Crnobrnja-Isailovic J, De la Riva I, Fawzi A, Galán P, Göçmen B, Harris DJ, Jiménez-Robles O, Joger U, Jovanović Glavaš O, Karış M, Koziel G, Künzel S, Lyra M, Miles D, Nogales M, Oğuz MA, Pafilis P, Rancilhac L, Rodríguez N, Rodríguez Concepción B, Sanchez E, Salvi D, Slimani T, S’khifa A, Qashqaei AT, Žagar A, Lemmon A, Moriarty Lemmon E, Carretero MA, Carranza S, Philippe H, Sinervo B, Müller J, Vences M, Wollenberg Valero KC (2019) Environmental temperatures shape thermal physiology as well as diversification and genome-wide substitution rates in lizards. Nat Commun 10(1):4077. https://doi.org/10.1038/s41467-019-11943-x
Geneva AJ, Park S, Bock D, de Mello P, Sarigol F, Tollis M, Donihue C, Reynolds RG, Feiner N, Rasys AM, Lauderdale JD, Minchey SG, Alcala AJ, Infante CR, Kolbe JJ, Schluter D, Menke DB, Losos JB (2022) Chromosome-scale genome assembly of the brown anole (Anolis Sagrei), an emerging model species. Commun Biol 5(1):1–13. https://doi.org/10.1038/s42003-022-04074-5
Gibbs JP, Breisch AR (2001) Climate warming and calling phenology of frogs near Ithaca, New York, 1900–1999. Conserv Biol 15(4):1175–1178. https://doi.org/10.1046/j.1523-1739.2001.0150041175.x
Green DM (2017) Amphibian breeding phenology trends under climate change: predicting the past to forecast the future. Glob Chang Biol 23(2):646–656. https://doi.org/10.1111/gcb.13390
Gutenkunst RN, Hernandez RD, Williamson SH, Bustamante CD (2009) Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet. https://doi.org/10.1371/journal.pgen.1000695
Habel JC, Rasche L, Schneider UA, Engler JO, Schmid E, Rödder D, Meyer ST, Trapp N, Sos del Diego R, Eggermont H, Lens L, Stork NE (2019) Final countdown for biodiversity hotspots. Conserv Lett 12(6):1–9. https://doi.org/10.1111/conl.12668
Harvey MG, Smith BT, Glenn TC, Faircloth BC, Brumfield RT (2016) Sequence capture versus restriction site associated DNA sequencing for shallow systematics. Syst Biol 65(5):910–924. https://doi.org/10.1093/sysbio/syw036
Hawlitschek O, Toussaint EFA, Gehring PS, Ratsoavina FM, Cole N, Crottini A, Nopper J, Lam AW, Vences M, Glaw F (2017) Gecko phylogeography in the Western Indian ocean region: the oldest clade of Ebenavia Inunguis lives on the youngest island. J Biogeogr 44(2):409–420. https://doi.org/10.1111/jbi.12912
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58(3):247–276. https://doi.org/10.1111/j.1095-8312.1996.tb01434.x
Hewitt GM (2004) Genetic consequences of climatic oscillations in the quaternary. Philos Trans R Soc Lond B Biol Sci 359(1442):183–195. https://doi.org/10.1098/rstb.2003.1388
Hewitt GM (2008) Speciation, hybrid zones and phylogeography—or seeing genes in space and time. Mol Ecol 10(3):537–549. https://doi.org/10.1046/j.1365-294x.2001.01202.x
Hof C, Araújo MB, Jetz W, Rahbek C (2011) Additive threats from pathogens, climate and land-use change for global amphibian diversity. Nature 480(7378):516–519. https://doi.org/10.1038/nature10650
Hoffmann AA, Sgrò CM (2018) Comparative studies of critical physiological limits and vulnerability to environmental extremes in small ectotherms: how much environmental control is needed? Integr Zool 13(4):355–371. https://doi.org/10.1111/1749-4877.12297
Hotaling S, Kelley JL, Frandsen PB (2021) Toward a genome sequence for every animal: where are we now? Proc Natl Acad Sci USA 118(52):1–8. https://doi.org/10.1073/pnas.2109019118
Huey RB (1982) Temperature, physiology, and the ecology of reptiles. Biol Reptil 1982:25–74
Huey RB (1991) Physiological consequences of habitat selection. Am Nat 137:S91-115. https://doi.org/10.1086/285141
Huey RB, Berrigan D (2001) Temperature, demography, and ectotherm fitness. Am Nat 158(2):204–210. https://doi.org/10.1086/321314
Huey RB, Deutsch CA, Tewksbury JJ, Vitt LJ, Hertz PE, Álvarez Pérez HJ, Garland T (2009) Why tropical forest lizards are vulnerable to climate warming. Proc r Soc B: Biol Sci 276(1664):1939–1948. https://doi.org/10.1098/rspb.2008.1957
Huey RB, Kearney MR, Krockenberger A, Holtum JAM, Jess M, Williams SE (2012) Predicting organismal vulnerability to climate warming: roles of behaviour, physiology and adaptation. Philos Trans R Soc B Biol Sci 367:1665–1679. https://doi.org/10.1098/rstb.2012.0005
Hulin V, Delmas V, Girondot M, Godfrey MH, Guillon JM (2009) Temperature-dependent sex determination and global change: are some species at greater risk? Oecologia 160(3):493–506. https://doi.org/10.1007/s00442-009-1313-1
Hutchison VH, Dupré RK (1992) Thermoregulation. Environ. Physiol. Amphib. 1992:206–249
IPCC (2014) Climate change 2014: synthesis report. contribution of working groups I, II and III to the fifth assessment report of the intergovernmental panel on climate change. IPCC, Geneva, p 151
Isaza DFG, Cramp RL, Franklin CE (2020) Living in polluted waters: a meta-analysis of the effects of nitrate and interactions with other environmental stressors on freshwater taxa. Environ Pollut 261:114091. https://doi.org/10.1016/j.envpol.2020.114091
Jaynes KE, Myers EA, Gvoždík V, Blackburn DC, Portik DM, Greenbaum E, Jongsma GFM, Rödel MO, Badjedjea G, Bamba-Kaya A, Baptista NL, Akuboy JB, Ernst R, Kouete MT, Kusamba C, Masudi FM, McLaughlin PJ, Nneji LM, Onadeko AB, Penner J, Pinto PV, Stuart BL, Tobi E, Zassi-Boulou AG, Leaché AD, Fujita MK, Bell RC (2021) Giant tree frog diversification in west and Central Africa: isolation by physical barriers, climate, and reproductive traits. Mol Ecol 31(15):3979–3998. https://doi.org/10.1111/mec.16169
Jensen JB, Bailey MA, Blankenship EL, Camp CD (2003) The relationship between breeding by the gopher frog, Rana Capito (Amphibia: Ranidae) and rainfall. Am Midl Nat 150(1):185–190. https://doi.org/10.1674/0003-0031(2003)150[0185:TRBBBT]2.0.CO;2
Jensen MP, Allen CD, Eguchi T, Bell IP, LaCasella EL, Hilton WA, Hof CAM, Dutton PH (2018) Environmental warming and feminization of one of the largest sea turtle populations in the world. Curr Biol 28(1):154-159.e4. https://doi.org/10.1016/j.cub.2017.11.057
Jocque M, Field R, Brendonck L, De Meester L (2010) Climatic control of dispersal-ecological specialization trade-offs: a metacommunity process at the heart of the latitudinal diversity gradient? Glob Ecol Biogeogr 19(2):244–252. https://doi.org/10.1111/j.1466-8238.2009.00510.x
Johnson PT, Chase JM, Dosch KL, Hartson RB, Gross JA, Larson DJ, Sutherland DR, Carpenter SR (2007) Aquatic eutrophication promotes pathogenic infection in amphibians. Proc Natl Acad Sci USA 104(40):15781–15786. https://doi.org/10.1073/pnas.0707763104
Jones MR, Good JM (2016) Targeted capture in evolutionary and ecological genomics. Mol Ecol 25(1):185–202. https://doi.org/10.1111/mec.13304
Kafash A, Ashrafi S, Yousefi M, Rastegar-Pouyani E, Rajabizadeh M, Ahmadzadeh F, Grünig M, Pellissier L (2020) Reptile species richness associated to ecological and historical variables in Iran. Sci Rep 10(1):1–11. https://doi.org/10.1038/s41598-020-74867-3
Kanamori S, Díaz LM, Cádiz A, Yamaguchi K, Shigenobu S, Kawata M (2022) Draft genome of six Cuban Anolis lizards and insights into genetic changes during their diversification. BMC Ecol Evo 22:129. https://doi.org/10.1186/s12862-022-02086-7
Kearney M, Shine R, Porter WP (2009) The potential for behavioral thermoregulation to buffer “cold-blooded” animals against climate warming. Proc Natl Acad Sci USA 106(10):3835–3840. https://doi.org/10.1073/pnas.0808913106
Kelleher J, Wong Y, Wohns AW, Fadil C, Albers PK, McVean G (2019) Inferring whole-genome histories in large population datasets. Nat Genet 51(9):1330–1338. https://doi.org/10.1038/s41588-019-0483-y
Kingman JFC (1982) The coalescent. Stoch Process Appl 13(3):235–248. https://doi.org/10.1016/0304-4149(82)90011-4
Koochekian N, Ascanio A, Farleigh K, Card DC, Schield DR, Castoe TA, Jezkova T (2022) A chromosome-level genome assembly and annotation of the desert horned lizard, Phrynosoma platyrhinos, provides insight into chromosomal rearrangements among Reptiles. GigaScience 11:1–14. https://doi.org/10.1093/gigascience/giab098
Lamichhaney S, Catullo R, Keogh JS, Clulow S, Edwards SV, Ezaz T (2021) A bird-like genome from a frog: mechanisms of genome size reduction in the ornate burrowing frog, Platyplectrum ornatum. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.2011649118
Lapierre M, Lambert A, Achaz G (2017) Accuracy of demographic inferences from the site frequency spectrum: the case of the Yoruba population. Genetics 206(1):139–449. https://doi.org/10.1534/genetics.116.192708
Le Galliard JF, Massot M, Baron JP, Clobert J (2012) Ecological effects of climate change on European reptiles. Wildlife Conserv Chang Clim. https://doi.org/10.13140/RG.2.1.3523.0248
Leaché AD, Portik DM, Rivera D, Rödel MO, Penner J, Gvoždík V, Greenbaum E, Jongsma GFM, Ofori-Boateng C, Burger M, Eniang EA, Bell RC, Fujita MK (2019) Exploring rain forest diversification using demographic model testing in the African foam-nest treefrog Chiromantis rufescens. J Biogeogr 46(12):2706–2721. https://doi.org/10.1111/jbi.13716
Leaché AD, Oaks JR, Ofori-Boateng C, Fujita MK (2020) Comparative phylogeography of West African amphibians and reptiles. Evolution 74(4):716–724. https://doi.org/10.1111/evo.13941
Ledo RMD, Domingos FMCB, Giugliano LG, Sites JW, Werneck FP, Colli GR (2020) Pleistocene expansion and connectivity of mesic forests inside the south American dry diagonal supported by the Phylogeography of a small lizard. Evolution 74(9):1988–2004. https://doi.org/10.1111/evo.13978
Leitwein M, Duranton M, Rougemont Q, Gagnaire PA, Bernatchez L (2020) Using haplotype information for conservation genomics. Trends Ecol Evol 35(3):245–258. https://doi.org/10.1016/j.tree.2019.10.012
Lemmon AR, Emme SA, Lemmon EM (2012) Anchored hybrid enrichment for massively high-throughput phylogenomics. Syst Biol 61(5):727–744. https://doi.org/10.1093/sysbio/sys049
Li H, Durbin R (2011) Inference of human population history from individual whole-genome sequences. Nature 475(7357):493–496. https://doi.org/10.1038/nature10231
Lieberman-Aiden E, Van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, Amit I, Lajoie BR, Sabo PJ, Dorschner MO, Sandstrom R, Bernstein B, Bender MA, Groudine M, Gnirke A, Stamatoyannopoulos J, Mirny LA (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326(5950):289–293. https://doi.org/10.1126/science.1181369
Liedtke HC, Gower DJ, Wilkinson M, Gomez-Mestre I (2018) Macroevolutionary shift in the size of amphibian genomes and the role of life history and climate. Nat Ecol Evol 2(11):1792–1799. https://doi.org/10.1038/s41559-018-0674-4
Liedtke HC, Cruz F, Gómez-Garrido J, Fuentes Palacios D, Marcet-Houben M, Gut M, Alioto T, Gabaldón T, Gomez-Mestre I (2022) Chromosome-level assembly, annotation and phylome of Pelobates Cultripes, the Western spadefoot toad. DNA Res 29(3):1–11. https://doi.org/10.1093/dnares/dsac013
Liu X, Fu YX (2015) Exploring population size Changes using SNP frequency spectra. Nat Genet 47(5):555–559. https://doi.org/10.1038/ng.3254
Lucena-Perez M, Marmesat E, Kleinman-Ruiz D, Martínez-Cruz B, Węcek K, Saveljev AP, Seryodkin IV, Okhlopkov I, Dvornikov MG, Ozolins J, Galsandorj N, Paunovic M, Ratkiewicz M, Schmidt K, Godoy JA (2020) Genomic patterns in the widespread Eurasian lynx shaped by late quaternary climatic fluctuations and anthropogenic impacts. Mol Ecol 29(4):812–828. https://doi.org/10.1111/mec.15366
Mao Y, Zhang G (2022) A complete, telomere-to-telomere human genome sequence presents new opportunities for evolutionary genomics. Nat Methods 19(6):635–638. https://doi.org/10.1038/s41592-022-01512-4
Marchi N, Schlichta F, Excoffier L (2021) Demographic inference. Curr Biol 31(6):R276–R279. https://doi.org/10.1016/j.cub.2021.01.053
Mather N, Traves SM, Ho SYW (2020) A practical introduction to sequentially markovian coalescent methods for estimating demographic history from genomic data. Ecol Evol 10(1):579–589. https://doi.org/10.1002/ece3.5888
Melville J, Haines ML, Hale J, Chapple S, Ritchie EG (2016) Concordance in phylogeography and ecological niche modelling identify dispersal corridors for reptiles in arid Australia. J Biogeogr 43(9):1844–1855. https://doi.org/10.1111/jbi.12739
Micheli-Campbell MA, Campbell HA, Cramp RL, Booth DT, Franklin CE (2011) Staying cool, keeping strong: incubation temperature affects performance in a freshwater turtle. J Zool 285(4):266–273. https://doi.org/10.1111/j.1469-7998.2011.00840.x
Miller W, Schuster SC, Welch AJ, Ratan A, Bedoya-Reinaa OC, Zhao F, Kim HL, Burhans RC, Drautz DI, Wittekindt NE, Tomsho LP, Ibarra-Laclette E, Herrera-Estrella L, Peacock E, Farley S, Sage GK, Rode K, Obbard M, Montiel R, Bachmann L, Ingólfsson Ó, Aars J, Mailund T, Wiig Ø, Talbot SL, Lindqvist C (2012) Polar and brown bear genomes reveal ancient admixture and demographic footprints of past climate change. Proc Natl Acad Sci USA 109(36):10. https://doi.org/10.1073/pnas.1210506109
Mitchell NJ, Janzen FJ (2010) Temperature-dependent sex determination and contemporary climate change. Sex Dev 4(1–2):129–140. https://doi.org/10.1159/000282494
Moritz C, Langham G, Kearney M, Krockenberger A, vanDerWal J, Williams SE (2012) Integrating phylogeography and physiology reveals divergence of thermal traits between central and peripheral lineages of tropical rainforest lizards. Philos Trans R Soc B Biol Sci 367(1596):1680–1687. https://doi.org/10.1098/rstb.2012.0018
Moussalli A, Moritz C, Williams SE, Carnaval AC (2009) Variable responses of skinks to a common history of rainforest fluctuation: concordance between phylogeography and palaeo-distribution models. Mol Ecol 18(3):483–499. https://doi.org/10.1111/j.1365-294X.2008.04035.x
Muñoz MM, Langham GM, Brandley MC, Rosauer DF, Williams SE, Moritz C (2016) Basking behavior predicts the evolution of heat tolerance in Australian rainforest lizards. Evolution 70(11):2537–2549. https://doi.org/10.1111/evo.13064
Nadachowska-Brzyska K, Li C, Smeds L, Zhang G, Ellegren H (2015) Temporal dynamics of avian populations during pleistocene revealed by whole-genome sequences. Curr Biol 25(10):1375–1380. https://doi.org/10.1016/j.cub.2015.03.047
Natesh M, Vinay KL, Ghosh S, Jayapal R, Mukherjee S, Vijay N, Robin VV (2020) Contrasting trends of population size change for two Eurasian owlet species—Athene brama and Glaucidium radiatum from South Asia over the late quaternary. Front Ecol Evol 8(608339):1–7. https://doi.org/10.3389/fevo.2020.608339
Nguyen HN, Lu CW, Chu JH, Grismer LL, Hung CM, Lin SM (2019) Historical demography of four gecko species specializing in boulder cave habitat: implications in the evolutionary dead end hypothesis and conservation. Mol Ecol 28(4):772–784. https://doi.org/10.1111/mec.14985
Noble DW, Stenhouse V, Schwanz LE (2018) Developmental temperatures and phenotypic plasticity in reptiles: a systematic review and meta-analysis. Biol Rev 93(1):72–97. https://doi.org/10.1111/brv.12333
Oficialdegui FJ, Sánchez MI, Monsalve-Carcaño C, Boyero L, Bosch J (2019) The invasive red swamp crayfish (Procambarus clarkii) increases infection of the amphibian chytrid fungus (Batrachochytrium dendrobatidis). Biol Invasions 21:3221–3231. https://doi.org/10.1007/s10530-019-02041-6
Oliver PM, Ashman LG, Bank S, Laver RJ, Pratt RC, Tedeschi LG, Moritz C (2019) On and off the rocks: persistence and ecological diversification in a tropical Australian lizard radiation. BMC Evol Biol 19(1):1–15. https://doi.org/10.1186/s12862-019-1408-1
Ovchinnikov V, Uliano-Silva M, Wilkinson M, Wood J, Smith M, Oliver K, Sims Y, Torrance J, Suh A, McCarthy SA, Durbin R, O’Connell MJ (2023) Caecilian genomes reveal the molecular basis of adaptation and convergent evolution of limblessness in snakes and caecilians. Mol Biol Evol. https://doi.org/10.1093/molbev/msad102
Pabijan M, Palomar G, Antunes B, Antoł W, Zieliński P, Babik W (2020) Evolutionary principles guiding amphibian conservation. Evol Appl 13(5):857–878. https://doi.org/10.1111/eva.12940
Palkopoulou E, Mallick S, Skoglund P, Enk J, Rohland N, Li H, Omrak A, Vartanyan S, Poinar H, Götherström A, Reich D, Dalén L (2015) Complete genomes reveal signatures of demographic and genetic declines in the woolly mammoth. Curr, Biol 25(10):1395–1400. https://doi.org/10.1016/j.cub.2015.04.007
Pasquesi GI, Adams RH, Card DC, Schield DR, Corbin AB, Perry BW, Reyes-Velasco J, Ruggiero RP, Vandewege MW, Shortt JA, Castoe TA (2018) Squamate reptiles challenge paradigms of genomic repeat element evolution set by birds and mammals. Nat Commun 9(1):1–11. https://doi.org/10.1038/s41467-018-05279-1
Patton AH, Margres MJ, Stahlke AR, Hendricks S, Lewallen K, Hamede RK, Ruiz-Aravena M, Ryder O, McCallum HI, Jones ME, Hohenlohe PA, Storfer A (2019) Contemporary demographic reconstruction methods are robust to genome assembly quality: a case study in Tasmanian devils. Mol Biol Evol 36(12):2906–2921. https://doi.org/10.1093/molbev/msz191
Peng LQ, Tang M, Liao JH, Liang SY, Gan LT, Hua KJ, Chen Y, Li H, Chen W, Merilä J (2020) Effects of temperature on growth and development of amphibian larvae across an altitudinal gradient in the Tibetan Plateau. Anim Biol 70(3):239–250. https://doi.org/10.1163/15707563-20201196
Pepper M, Ho SYW, Fujita MK, Keogh JS (2011) The genetic legacy of aridification: climate cycling fostered lizard diversification in Australian montane refugia and left low-lying deserts genetically depauperate. Mol Phylogenet Evol 61(3):750–759. https://doi.org/10.1016/j.ympev.2011.08.009
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo snp discovery and genotyping in model and non-model species. PLoS ONE 7(5):e37135. https://doi.org/10.1371/journal.pone.0037135
Pie MR, Campos LLF, Meyer ALS, Duran A (2017) The evolution of climatic niches in squamate reptiles. Proc Royal Soc B Biol Sci 284(1858):20170268. https://doi.org/10.1098/rspb.2017.0268
Portik DM, Leaché AD, Rivera D, Barej MF, Burger M, Hirschfeld M, Rödel MO, Blackburn DC, Fujita MK (2017) Evaluating mechanisms of diversification in a Guineo-Congolian tropical forest frog using demographic model selection. Mol Ecol 26(19):5245–5263. https://doi.org/10.1111/mec.14266
Potter S, Xue AT, Bragg JG, Rosauer DF, Roycroft EJ, Moritz C (2018) Pleistocene climatic changes drive diversification across a tropical savanna. Mol Ecol 27(2):520–532. https://doi.org/10.1111/mec.14441
Prates I, Xue AT, Brown JL, Alvarado-Serrano DF, Rodrigues MT, Hickerson MJ, Carnaval AC (2016) Inferring responses to climate dynamics from historical demography in neotropical forest lizards. Proc Natl Acad Sci USA 113(29):7978–7985. https://doi.org/10.1073/pnas.1601063113
Pudlo P, Marin JM, Estoup A, Cornuet JM, Gautier M, Robert CP (2016) Reliable ABC model choice via random forests. Bioinformatics 32(6):859–866. https://doi.org/10.1093/bioinformatics/btv684
Quintero I, Wiens JJ (2013) What determines the climatic niche width of species? The role of spatial and temporal climatic variation in three vertebrate clades. Glob Ecol Biogeogr 22(4):422–432. https://doi.org/10.1111/geb.12001
Reading CJ (2007) Linking global warming to amphibian declines through its effects on female body condition and survivorship. Oecologia 151(1):125–131. https://doi.org/10.1007/s00442-006-0558-1
Rodgers EM (2021) Adding climate change to the mix: Responses of aquatic ectotherms to the combined effects of eutrophication and warming. Biol Lett 17(10):20210442. https://doi.org/10.1098/rsbl.2021.0442
Rohr JR, Palmer BD (2013) Climate change, multiple stressors, and the decline of ectotherms. Conserv Biol 27(4):741–751. https://doi.org/10.1111/cobi.12086
Rolland J, Silvestro D, Schluter D, Guisan A, Broennimann O, Salamin N (2018) The impact of endothermy on the climatic niche evolution and the distribution of vertebrate diversity. Nat Ecol Evol 2(3):459–464. https://doi.org/10.1038/s41559-017-0451-9
Rosauer DF, Byrne M, Blom MPK, Coates DJ, Donnellan S, Doughty P, Keogh JS, Kinloch J, Laver RJ, Myers C, Oliver PM, Potter S, Rabosky DL, Afonso Silva AC, Smith J, Moritz C (2018) Real-world conservation planning for evolutionary diversity in the Kimberley, Australia sidesteps uncertain taxonomy. Conserv Lett 11(4):1–10. https://doi.org/10.1111/conl.12438
Salmona J, Heller R, Lascoux M, Shafer A (2019) Inferring demographic history using genomic data. In: Rajora OP (ed) Population genomics: concepts, approaches and applications. Springer International Publishing, Cham, pp 511–537
Sarabia C, vonHoldt B, Larrasoaña JC, Uríos V, Leonard JA (2020) Pleistocene climate fluctuations drove demographic history of African golden wolves (Canis lupaster). Mol Ecol 30(23):6101–6120. https://doi.org/10.1111/mec.15784
Schiffels S, Durbin R (2014) Inferring human population size and separation history from multiple genome sequences. Nat Genet 46(8):919–925. https://doi.org/10.1038/ng.3015
Schneider C, Moritz C (1999) Rainforest refugia and evolution in Australia’s wet tropics. Proc Royal Soc B Biol Sci 266(1415):191–196. https://doi.org/10.1098/rspb.1999.0621
Schrider DR, Kern AD (2018) Supervised machine learning for population genetics: a new paradigm. Trends Genet 34(4):301–312. https://doi.org/10.1016/j.tig.2017.12.005
Sellinger TPP, Abu-Awad D, Moest M, Tellier A (2020) Inference of past demography, dormancy and self-fertilization rates from whole genome sequence data. PLoS Genet 16(4):1–28. https://doi.org/10.1371/journal.pgen.1008698
Sellinger TPP, Abu-Awad D, Tellier A (2021) Limits and convergence properties of the sequentially markovian coalescent. Mol Ecol Resour 21(7):2231–2248. https://doi.org/10.1111/1755-0998.13416
Sexton JP, Montiel J, Shay JE, Stephens MR, Slatyer RA (2017) Evolution of ecological niche breadth. Annu Rev Ecol Evol Syst 48:183–206. https://doi.org/10.1146/annurev-ecolsys-110316-023003
Sillero N, Arenas-Castro S, Enriquez-Urzelai U, Vale CG, Sousa-Guedes D, Martínez-Freiría F, Real R, Barbosa AM (2021) Want to model a species niche? A step-by-step guideline on correlative ecological niche modelling. Ecol Modell 456:109671. https://doi.org/10.1016/j.ecolmodel.2021.109671
Simoncini MS, Cruz FB, Larriera A, Piña CI (2014) Effects of climatic conditions on sex ratios in nests of broad-snouted caiman. J Zool 293(4):243–251. https://doi.org/10.1111/jzo.12140
Singh SK, Das D, Rhen T (2020) Embryonic temperature programs phenotype in reptiles. Front Physiol. https://doi.org/10.3389/fphys.2020.00035
Singhal S, Moritz C (2013) Reproductive isolation between phylogeographic lineages scales with divergence. Proc Royal Soc B Biol Sci. https://doi.org/10.1098/rspb.2013.2246
Sinsch U (1991) Mini-review: The orientation behaviour of amphibians. Herpetol J 1(54):1–544
Smith JJ, Timoshevskaya N, Timoshevskiy VA, Keinath MC, Hardy D, Voss SR (2019) A chromosome-scale assembly of the axolotl genome. Genome Res 29(2):317–324. https://doi.org/10.1101/gr.241901.118
Snyder GK, Weathers WW (1975) Temperature adaptations in amphibians. Am Nat 109(965):93–101. https://doi.org/10.1086/282976
Sousa V, Hey J (2013) Understanding the origin of species with genome-scale data: modelling gene flow. Nat Rev Genet 14(6):404–414. https://doi.org/10.1038/nrg3446
Speidel L, Forest M, Shi S, Myers SR (2019) A method for genome-wide genealogy estimation for thousands of samples. Nat Genet 51(9):1321–1329
Spence JP, Steinrücken M, Terhorst J, Song YS (2018) Inference of population history using coalescent HMMs: review and outlook. Curr Opin Genet Dev 53:70–76. https://doi.org/10.1016/j.gde.2018.07.002
Sunday JM, Bates AE, Kearney MR, Colwell RK, Dulvy NK, Longino JT, Huey RB (2014) Thermal-safety margins and the necessity of thermoregulatory behavior across latitude and elevation. Proc Natl Acad Sci USA 111(15):5610–5615. https://doi.org/10.1073/pnas.1316145111
Taheri S, Naimi B, Rahbek C, Araújo MB (2021) Improvements in reports of species redistribution under climate change are required. Sci Adv 7(15):1–12. https://doi.org/10.1126/sciadv.abe1110
Taylor EN, Diele-Viegas LM, Gangloff EJ, Hall JM, Halpern B, Massey MD, Rödder D, Rollinson N, Spears S, Sun BJ, Telemeco RS (2021) The thermal ecology and physiology of reptiles and amphibians: a user’s guide. J Exp Zool A Ecol Integr Physiol 335(1):13–44. https://doi.org/10.1002/jez.2396
Terhorst J, Kamm JA, Song YS (2017) Robust and scalable inference of population history from hundreds of unphased whole genomes. Nat Genet 49(2):303–309. https://doi.org/10.1038/ng.3748
Thörn F, Rödin-Mörch P, Cortazar-Chinarro M, Richter-Boix A, Laurila A, Höglund J (2021) The Effects of drift and selection on latitudinal genetic variation in scandinavian common toads (Bufo Bufo) following postglacial recolonisation. Heredity 126(4):656–667. https://doi.org/10.1038/s41437-020-00400-x
Tollis M, Ausubel G, Ghimire D, Boissinot S (2012) Multi-locus phylogeographic and population genetic analysis of Anolis carolinensis: historical demography of a genomic model species. PLoS ONE 7(6):1–14. https://doi.org/10.1371/journal.pone.0038474
Valenzuela N, Literman R, Neuwald JL, Mizoguchi B, Iverson JB, Riley JL, Litzgus JD (2019) Extreme thermal fluctuations from climate change unexpectedly accelerate demographic collapse of vertebrates with temperature-dependent sex determination. Sci Rep 9(1):1–11. https://doi.org/10.1038/s41598-019-40597-4
Vijay N, Park C, Oh J, Jin S, Kern E, Kim HW, Zhang J, Park JK (2018) Population genomic analysis reveals contrasting demographic changes of two closely related dolphin species in the last glacial. Mol Biol Evol 35(8):2026–2033. https://doi.org/10.1093/molbev/msy108
Waldvogel AM, Feldmeyer B, Rolshausen G, Exposito-Alonso M, Rellstab C, Kofler R, Mock T, Schmid K, Schmitt I, Bataillon T, Savolainen O, Bergland A, Flatt T, Guillaume F, Pfenninger M (2020) Evolutionary genomics can improve prediction of species’ responses to climate change. Evol Lett 4(1):4–18. https://doi.org/10.1002/evl3.154
Wen G, Fu J (2021) Isolation and reconnection: demographic history and multiple contact zones of the green odorous frog (Odorrana Margaretae) around the Sichuan Basin. Mol Ecol 30(16):4103–4117. https://doi.org/10.1111/mec.16021
Winter M, Fiedler W, Hochachka WM, Koehncke A, Meiri S, De La Riva I (2016) Patterns and biases in climate change research on amphibians and reptiles: a systematic review. R Soc Open Sci 3(9):160158. https://doi.org/10.1098/rsos.160158
Wollenberg Valero KC, Marshall JC, Bastiaans E, Caccone A, Camargo A, Morando M, Niemiller ML, Pabijan M, Russello MA, Sinervo B, Werneck FP, Sites JW Jr, Wiens JJ, Steinfartz S (2019) Patterns, mechanisms and genetics of speciation in reptiles and amphibians. Genes 10(646):47. https://doi.org/10.3390/genes10090646
Xue AT, Hickerson MJ (2017) Multi-dice: R package for comparative population genomic inference under hierarchical co-demographic models of independent single-population size changes. Mol Ecol Resour 17(6):e212–e224. https://doi.org/10.1111/1755-0998.12686
Acknowledgements
The authors would like to thank Sally Potter, Joshua Penalba, Filip Thörn, Mario Ernst and Darija Josić for their helpful comments on the earlier versions of the manuscript, as well as three anonymous reviewers for their insightful comments during submission. S.I.H.B. was funded by the Elsa-Neumann scholarship from the State of Berlin, Germany.
Funding
Open Access funding enabled and organized by Projekt DEAL. Author S.I.H.B was supported by the Elsa-Neumann scholarship from the State of Berlin, Germany.
Author information
Authors and Affiliations
Contributions
All authors contributed to the review conception and design. The first draft of the manuscript was written by S.I.H.B and M.P.K.B and both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Pedro Aragón.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hayden Bofill, S.I., Blom, M.P.K. Climate change from an ectotherm perspective: evolutionary consequences and demographic change in amphibian and reptilian populations. Biodivers Conserv 33, 905–927 (2024). https://doi.org/10.1007/s10531-023-02772-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-023-02772-y