Abstract
Evolution can occur over contemporary timescales, which may be crucial for the invasive success of non-native plant species. Many studies have shown rapid evolution by comparing native and non-native populations in common gardens. However, our understanding of the mechanisms underpinning rapid evolution is still incomplete. Here, we identify the progress, applications, and limitations of studies on rapid evolution of non-native plants with respect to sampling, experimental design and experimental methods. To encompass broad variation within and between the ranges, we recommend sampling across large-scale environmental gradients. We also suggest careful consideration of pitfalls related to the choice of seed families and of the biotic interaction under focus. The latter should be chosen with a view on both the experimental treatment and the corresponding field data to estimate population history. Furthermore, we suggest exploiting multiple omics approaches to address the complexity of biotic interactions, and to account for non-adaptive evolution with molecular data on demographic history of populations. We also reviewed papers that studied rapid evolution in non-native plants and quantified how many of these met our criteria. We anticipate that disentangling adaptive and non-adaptive drivers of among-population variation can increase the accuracy of research on rapid evolution, and that integrating phenotypic, metabolomic and population genomic data can bring opportunities for studying complex biotic interactions. We also illustrate the importance of large collaborative networks and present our scientific network iCONNECT (integrative CONyza NEtwork for Contemporary Trait evolution), with the goal of motivating similar studies on the mechanistic understanding of rapid evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Natural selection can act on very short ecological scales, shifting the genomes, metabolomes and phenotypes of populations over a few generations, a process which is called “rapid evolution” (e.g., Thompson 1998) or in some other contexts also “contemporary evolution” (e.g., Hendry and Kinnison 1999). We here follow the definition of Thompson (1998) using the term rapid evolution as an adaptive process that has immediate ecological consequences on very short evolutionary scales. Rapid evolution can result from standing genetic variation (e.g., altering genotype frequencies through lineage sorting) and from the emergence of novel genotypes, either through recombination within populations or through admixture of previously isolated gene pools (Turcotte et al. 2011; Cameron et al. 2013; Dlugosch et al. 2016). The ability to rapidly adapt to environmental change is of major importance for plant population survival under global change (Hoffmann and Sgrò 2011; Franks et al. 2014; Rosche et al. 2018a; Rauschkolb et al. 2022) and for successful range expansion by non-native plant species (Ochocki and Miller 2017; Szűcs et al. 2017; Woods and Sultan 2022).
Here we focus on rapid evolution in the course of plant invasions. This may happen when non-native plant populations rapidly adapt to novel environmental conditions in their non-native ranges. Many plant invasions are characterized by a “lag-phase”, i.e., delay between the introduction of a non-native species and its successful colonization in a new area (Osunkoya et al. 2021). This process likely coincides with rapid adaptation to the novel environment and is often followed by the colonization of the invader (Aikio et al. 2010; Clements and DiTommaso 2011). Indeed, many studies have shown genomic, metabolomic and phenotypic differences between native and non-native populations under common garden conditions, indicating rapid evolution as a common occurrence in biological invasions reviewed by Bossdorf et al. (2005); Colautti and Lau (2016); van Kleunen et al. (2018); Clements and Jones (2021). In many cases, rapid evolution was associated with higher performance or greater competitive ability of non-native compared to native populations in common gardens, especially for successful invaders (e.g., Zheng et al. 2015; Montesinos et al. 2019).
There are two reasons why rapid evolution is frequent in biological invasions. First, non-native populations undergo dramatic demographic changes. In particular, population genetic studies often observe colonization bottlenecks or founder effects associated with genetic drift on the one hand and multiple introductions that intensify gene pool admixture on the other hand. Together, these processes can lead to fission and fusion of native gene pools in the non-native ranges (Rius and Darling 2014; Estoup et al. 2016; Rosche et al. 2016). Alexander et al. (2009), for example, showed that population genetic structure was much weaker in the introduced area, and populations were not isolated by distance. Second, such rearrangements of gene pools occur when non-native populations experience fundamentally altered selection pressures as compared to populations in the native range (van Kleunen et al. 2018). Shifts in biotic interactions are considered as the most important selection pressures of rapid evolution in non-native ranges (Moran and Alexander 2014; Dlugosch et al. 2016; Moles et al. 2022; Sheng et al. 2022). The most prominent example in this context may be the release from specialized herbivores and pathogens present in their native ranges (enemy release hypothesis, Keane and Crawley 2002).
In recent years, several hypotheses have been proposed regarding the underlying mechanisms of rapid evolution reviewed by Jeschke and Heger (2018). These hypotheses have been tested in numerous case studies and have significantly improved our understanding of the importance of biotic interactions in biological invasions (Dlugosch et al. 2016). However, studies of rapid evolution are not always carefully set up with rigorous methodologies and there are many contradictory findings when testing theories based on rapid evolution (Colautti and Lau 2016), including, for example, the “evolution of increased competitive ability-hypothesis” (EICA, Blossey and Notzold 1995; Rotter and Holeski 2018; Callaway et al. 2022).
Because comprehensive frameworks are essential to elucidate the intricacies of adaptive processes over short evolutionary scales, many recent research calls for more sophisticated eco-evolutionary frameworks to study rapid evolution (e.g., Colautti and Lau 2016; Schrieber and Lachmuth 2017; Schrieber et al. 2017; Rotter and Holeski 2018; Rosche et al. 2018b, 2019; Sheng et al. 2022). They identified manifold issues that may arise when studying rapid evolution in biotic interaction traits. First, the samplings in the native and the non-native ranges can be unbalanced, thereby not representing the full breadth of conditions in either range (Colautti and Lau 2016). Second, the experimental set-ups are diverse and each set up comes with its own specific limitations (de Villemereuil et al. 2016; Schrieber and Lachmuth 2017). Third, biotic interactions are complex by nature and therefore difficult to describe and quantify (e.g., competitive ability), making their assessment as selective forces a challenging task for evolutionary ecologists (Aschehoug et al. 2016).
To address apparent issues in studying rapid evolution, we review progress, applications and limitations of current research. Different empirical methods are applied to study rapid evolution from the genetic to the phenotypic levels. Some experiments observe phenotypic shifts under different treatments over a few generations (Williams et al. 2016; Szűcs et al. 2017) whereas others use resequencing approaches to identify allele frequency shifts (Turner et al. 2011; Schlötterer et al. 2015). Also, herbarium specimens can identify trait shifts through invasion histories (Wu and Colautti 2022) and resurrection approaches are used to compare revived ancestors from stored propagules with their progeny (Sultan et al. 2013; Franks et al. 2018; Rauschkolb et al. 2022). However, we focus here on classical population ecological studies which represent the vast majority of studies dealing with rapid evolution, i.e., experimental setups that compare native and non-native populations under common garden conditions. In this context, we focus on three topics where we believe progress is needed to advance our mechanistic understanding of rapid evolution (Fig. 1):
-
(1)
Sampling Consider comparable spatial and environmental variation in both the native and non-native ranges to avoid misleading conclusions drawn from unrepresentative samplings in either range.
-
(2)
Design of experiments Conceptualize approaches for testing biotic interactions of interest and recognize common experimental pitfalls to design an appropriate experiment.
-
(3)
Experimental methods Utilize interdisciplinary approaches to examine complex biotic interaction traits with applicable omics tools.
Conceptual outline how to overcome current limitations in (1) sampling, (2) experimental design, and (3) experimental methods to study rapid evolution between native and non-native ranges. We suggest (1a) sampling broad and comparable variation in environmental gradients within and between native and non-native ranges, (1b) sampling comparable spatial distribution in either range, (2a) evaluating the focal biotic interaction for both the design of experimental treatments and the collection of field data on population history. Experiments should also consider (2b) pitfalls related to seed family selection. In addition, we propose to use multiple omics approaches to (3a) capture the complexity of biotic interactions with currently available tools and to (3b) account for non-adaptive evolution with molecular data on population demographic history
We propose considerations that researchers should make when designing studies that test differences between native and non-native populations. We also reviewed papers from the journal Biological Invasions that studied rapid evolution in non-native plants using common garden approaches (Table 1) and quantified how many of these met our criteria (related to the topics 1 to 3 above). Finally, we illustrate the importance of large collaborative networks and present our scientific network iCONNECT (integrative CONyza NEtwork for Contemporary Trait evolution) in which we are studying rapid evolution in competitive ability of native and non-native populations of Conyza canadensis (syn. Erigeron canadensis), as an example of how this could be achieved.
Sampling comparable spatial and environmental variation in both ranges
Common garden experiments have frequently found differences between native and non-native populations (Kulmatiski 2019; Montesinos 2022; Zhang et al. 2022). Such between-range differences may suggest evidence for rapid evolution assuming that they are based on altered selection pressures between the ranges. However, between-range variation is just a part of the global spectrum of among-population variation (APV, i.e., intraspecific trait variation that is due to selection pressures associated with population history). In this regard, it is important to consider that populations from both native and non-native ranges can show strong within-range variation in their population history (Oduor et al. 2022; Sheng et al. 2022). This means that the choice of sampled populations from either range can obviously affect the results of native vs. non-native comparisons in the common garden (Rosche et al. 2018b, 2019).
Colautti and Lau (2016) reviewed 31 studies that postulated rapid evolution in non-native populations. They used data simulations and found that 24 of these studies did not present sufficient data to support their conclusions. Most of them compared only few and rather haphazardly chosen populations from large areas and therefore underestimated APV within each range. Many studies also sampled along different environmental gradients in the native vs. non-native ranges. We found that only 23.5% of our reviewed studies sampled native and non-native populations across comparable environmental gradients in either range. In those cases, significant differences between native and non-native populations might result from within-range APV, which was not measured in both ranges to a comparable extent (Rosche et al. 2019). To ensure appropriate between-range comparisons testing for rapid evolution, studies should have sufficient and representative environmental variation in both the native and non-native ranges (Fig. 1-1a). This can be achieved by sampling as many populations as possible while maximizing environmental variation in both ranges. Another solution was recently demonstrated by Sheng et al. (2022) who paired four matching bioclimatic regions in the native and non-native ranges to have comparable climatic backgrounds for their study populations from either range.
Besides incorporating environmental gradients in the sampling design, studies should also sample comparable spatial distributions in native vs. non-native ranges (Fig. 1-1b). We found that none of our reviewed studies sampled native and non-native populations along comparable spatial distances in either range. This is important because variables measured from locations close to each other often exhibit more similar values than those from further apart (i.e., spatial autocorrelation, Legendre 1993). A regional clustering of samplings can help to account for spatial auto-correlation if number and spatial distances of populations within geographical regions are comparable among the regions and between native and non-native ranges (Sheng et al. 2022). Such sampling design may allow setting population nested within region as a random effect which was recently shown to appropriately account for spatial auto-correlation in native vs. non-native range comparisons (Rosche et al. 2019). There are many other statistical approaches that account for spatial auto-correlation in ecology but all of which benefit from a judicial representation of spatial distributions (see Kuehn and Dormann 2012).
Conceptual approaches for experimental designs
To investigate the degree to which differences between native and non-native populations are attributable to rapid evolution, rather than other components of APV, it is important to identify biotic interactions (Fig. 1-2a) that differ between the ranges with the potential to alter selection pressures (Jones and Gomulkiewicz 2012; van Kleunen et al. 2018). There are many types of biotic interactions that determine the success of invasive species, such as pollination (Harmon‐Threatt et al. 2009; Mackin et al. 2021), herbivory (Hu and Dong 2019; Yin et al. 2023), pathogen infestation (Goss et al. 2020), soil biota mutualisms (Callaway and Lucero 2020; Sheng et al. 2022) and competitive interactions (Shah et al. 2014). For the sake of clarity, we here focus exemplarily on competitive interactions, but our descriptions arguments below can be applied in a similar way to other biotic interactions under focus.
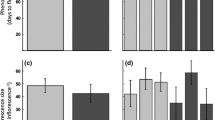
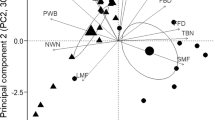
Conceptual scheme of the current research in the iCONNECT project that aims to disentangle drivers of rapid evolution in the competitive ability of Conyza canadensis. The project has four connected and interdisciplinary work packages (WPs) that analyze data from one greenhouse experiment (olive boxes). The experiment investigates among-population variation in the response of 112 native and 177 non-native populations to experimental competition × drought treatments. Available field data will be used as a proxy for the population history of the local drought regime, the competitive regime, mutualist-pathogen ratio in the field rhizosphere (gray boxes). The population history data and data on neutral genetic relatedness of the populations are anticipated to help disentangling how adaptive and non-adaptive evolution determinants drive among population variation with respect to the response to experimental competition × drought treatments, both within and between native and non-native ranges
Competitive interactions are among the most important biotic interactions that determine the success of invasions (Callaway et al. 2011) because native and non-native species often interact differently with co-existing species (Felker-Quinn et al. 2013; Shah et al. 2014; Pearse et al. 2019; Pal et al. 2020; Nagy et al. 2022). Studies on rapid evolution often select treatments that simulate the particular biotic interaction assumed to drive rapid evolution (e.g., Turcotte et al. 2011; Schrieber et al. 2017; Macel et al. 2017; Rosche et al. 2018b; Javed et al. 2020). For example, if different competitive interactions between the native and non-native ranges are observed in the field, studies can apply a competition treatment under common conditions (e.g., neighbor vs. no neighbor; e.g., Shah et al. 2014). Such a design allows recording whether the response to the experimental treatment differs between the ranges (i.e., range x treatment interaction).
To integrate the effects of APV within and between ranges, field data that deal with the biotic interaction of interest can be included in the models as a proxy for population history. Such proxies can explicitly test whether the strength and direction of natural selection in recent population history differ between native and non-native ranges (i.e., range x population history x treatment interaction; e.g., Schrieber et al. 2017; Rosche et al. 2018b; Villellas et al. 2021). Moreover, such assessments allow testing for differential effects of plasticity vs. local adaptation in both native and non-native ranges (Villellas et al. 2021). In studies of competition as the main driver of rapid evolution, field data on the competitive regime can be used to test whether competition in the population history has the same effects on competitive ability in the greenhouse in native vs. non-native populations (Lachmuth et al. 2011; Heger et al. 2014). As an example different from competitive interactions, Schrieber et al. (2017) investigated how the herbivory that native and non-native populations experienced in the field affected experimental herbivory under common garden conditions. However, only 11.8% of the reviewed studies investigated how biotic interactions experienced in the field might affect the outcome of corresponding experimental treatments as an indication of rapid evolution.
Another important point for designing experiments is the consideration of seed family variation to minimize unexplained variance in common garden experiments (Fig. 1-2b). Within populations, genotypes may differ in phenotype with some genotypes showing a high performance across multiple environments (e.g., general purpose genotype, Barrett 2016), which can be a problem for experimental designs. For example, in competition experiments, an overrepresentation of general purpose genotypes in the “competition treatment” as compared to the “no competition treatment” may lead to overestimation of the population´s competitive ability. One way to avoid this pitfall is to control for equal distribution of seed families across the experimental treatments in the greenhouse (Lachmuth et al. 2011; Schrieber et al. 2017). 29.4% of the papers that we reviewed used different seed families in their experiments to distribute them equally across their treatments. This means that the effect of individual genomes has been underestimated in the majority of the studies.
Other considerations in experimental design regarding the seed families include maternal effects and breeding background (Pico et al. 2003). Offspring may show responses to conditions experienced by the mother (maternal effects, Herman and Sultan 2011; Heger 2016). Moreover, field conditions influence the resource availability of the mother plant and thus indirectly affect the offspring in the experiment (Veselá et al. 2021). Greenhouse estimates of population fitness thus depend on the environmental conditions that the mothers faced in the sampling year (de Villemereuil et al. 2016; de Kort et al. 2020). Differences in mother fitness on offspring performance may be accounted for by including seed mass as a covariate in the models (Dyer et al. 2010). Also, the breeding background of an individual (e.g., selfing, biparental inbreeding, outcrossing) can affect its performance (reviewed by Angeloni et al. 2011). This may result in unexplained variance in experiments on rapid evolution (Rosche et al. 2017; Schrieber et al. 2017; Gustafsson et al. 2022). The use of F1-offspring generated under standardized environmental and breeding conditions can control for both maternal effects and breeding background. However, such approach would reduce epigenetic variation which—depending on the research question—can be preferable or not.
Experimental methods: utilizing various omics approaches
Traditionally recorded phenotypic data can only provide inference of rapid evolution and ideally should be accompanied by molecular data (de Villemereuil et al. 2016, 2022). However, most native vs. non-native range comparisons have focused on phenotypic data reviewed by Bossdorf et al. (2005). Others have used population genomic reviewed by Flucher et al. (2021), or metabolomic data (e.g., Macel et al. 2014; Wu et al. 2020; Yu et al. 2022; Yin et al. 2023), but these did not cover APV in both ranges. In fact, studies that cover both molecular and phenotypic data across broad spatio-environmental gradients are rare and almost exclusively focus on Arabidopsis thaliana (e.g., Mönchgesang et al. 2016; Exposito-Alonso et al. 2019). In our review, none of the studies used multi-omic approaches. This is an important knowledge gap as complex biological questions cannot be investigated using only a few genotypes or populations (Des Roches et al. 2018; Milcu et al. 2018). With insights into drivers of rapid evolution in biotic interactions, applying multiple omics approaches may be particularly promising (Fig. 1-3a) because biotic interactions are often characterized by trait complexes which are difficult to understand, quantify, or even to define. For example, competitive ability is a complex of traits comprising the tolerance of an individual to the suppressive effects of nearby plants, and traits that usurp resources from a finite pool and traits that directly inhibit growth of neighboring plants (Aschehoug et al. 2016).
Molecular interaction ecology
Eco-metabolomics is a powerful tool for understanding the underlying mechanisms of ecological processes (Peters and Worrich 2018), may help to understand hitherto unexplained variation in plant interactions (Walker et al. 2022; 2023) and can unravel strategies of invasive plants (Macel et al. 2014). Metabolites are viewed as one of the intermediary layers between the genome and expressed phenotypes, and thus represent a key component to understanding how species interact (Yang et al. 2021; Walker et al 2022; Auwerx et al. 2023). We here focus exemplarily on studying root exudates; yet, metabolomics can be applied for studying many other biotic interactions (e.g., studying secondary metabolites in leaves in herbivory studies, Wu et al. 2022).
Root exudates can have direct effects on the competitive ability of plants via the toxic suppression of nearby plant growth, known as allelopathy (Semchenko et al. 2014). The novel-weapons hypothesis posits that the overwhelming success of some plants in their non-native ranges is due to allelopathic suppression of native species that are not adapted to the novel chemical traits (Hierro and Callaway 2021). At the same time, an increased allelopathic susceptibility of local competitors in non-native ranges may stimulate rapid evolution for the production of allelopathic substances between native and non-native ranges, although this has rarely been investigated (but see Irimia et al. 2019 for leaf leachates). Moreover, root exudates may shape the soil biota community in the rhizosphere by attracting mutualists or repelling pathogens (Weidenhamer and Callaway 2010; van Dam and Bouwmeester 2016; Yu et al. 2022). This is important because shifts in soil biota communities affect resource availability and can have strong indirect effects on competitive interactions among plants, while these relationships may differ strongly in native vs. nonnative ranges (Lekberg et al. 2018).
Many studies support enemy release from soil biota in non-native ranges, particularly for specialized pathogens (Kulmatiski et al. 2008; Flory et al. 2011). The reduced pathogen pressure in non-native populations may also facilitate beneficial interactions with mutualists (Reinhart and Callaway 2006; Sheng et al. 2022). This aligns with competition experiments where non-native plants benefit more than native plants from the presence of soil fungi (e.g., Callaway and Lucero 2020). However, while the evolutionary consequences of biotic interactions have been studied extensively for aboveground interactions, we know very little on how altered soil biota communities may trigger rapid evolution in non-native plants (Lekberg et al. 2018; Sheng et al. 2022). Current meta-genomic tools such as next generation amplicon sequencing or the recent progress in analyzing root exudates increase our abilities to address such questions related to the “belowground black box” (van Kleunen et al. 2018).
Population genomics and rapid evolution
High throughput sequencing approaches are revolutionizing the field of eco-evolutionary genomics by providing large amounts of genome-wide single nucleotide polymorphisms (SNPs, Andrews et al. 2016). The availability of thousands of SNPs facilitates our ability to unravel how gene flow is controlled by the interplay of adaptive vs. non-adaptive drivers of the dispersal probability at the global scale (Orsini et al. 2013; Levy and Boone 2019). Quantifying gene flow among populations is crucial for a comprehensive understanding of the demographic history of populations (Fig. 1-3b, Al‐Gharaibeh et al. 2017; Nagy et al. 2018; Vendrami et al. 2019; Walsh et al. 2021), and important for predicting native source populations and reconstructing the invasion history of non-native populations (Fitzpatrick et al. 2012; Hou and Li 2020; Bieker et al. 2022; Encinas‐Viso et al. 2022; McCulloch et al 2023).
Moreover, knowledge on the demographic history of populations allows accounting for non-adaptive evolution when analysing phenotypic or metabolomic APV (Raeymaekers et al. 2017; de Villemereuil et al. 2022). APV in general, and between-range variation more specifically, is not only caused by natural selection but can also result from purely demographic processes such as genetic drift and migration history (i.e., population co-ancestry, Keller and Taylor 2008). For example, Rosche et al. (2019) used pedigree mixed-effects models that account for population co-ancestry and demonstrated for C. canadensis that non-adaptive evolution may false-positively indicate rapid evolution or obscure adaptive effects due to unexplained residual variance.This variance is particularly useful for studying rapid evolution because both fission and fusion of native gene pools can shift allele frequencies towards sweep scenarios, where adaptive allele frequencies vary greatly across non-native populations without actually corresponding to any selection pressures (Ravinet et al. 2017; Rosche et al. 2019; Irimia et al. 2023). However, our review indicated that none of the studies have taken population genomic information into account to analyses when studying rapid evolution in non-native plants.
Furthermore, mapping high throughput SNP data on assembled genomes can identify genomic regions that are associated with environmental gradients in population histories (McKown et al. 2014). Similarly, these data can detect genomic regions that are associated with phenotypic and metabolomic APV in common garden experiments. Combined, these data provide insights into whether selection patterns in the population history correspond with APV in experiments (Exposito-Alonso et al. 2019). Mapping of candidate SNP loci can also identify functional genes that are potentially related to APV. This may allow, for example, identifying gene size changes, novel genomic rearrangements and local adaptations underpinning rapid evolutionary change (Andrews et al. 2016; Hendry 2016; Rudman et al. 2018), and may also be used to design probes for sequence capture approaches in further studies (Fahrenkrog et al. 2017; Capblancq et al. 2020). Together, these approaches can assess the causality of adaptive correlations observed in eco-evolutionary studies.
Large collaborative networks to address adaptive evolution
Large collaborative networks are crucial for advancing our understanding of ecology and evolution (Goring et al. 2014; Papale et al. 2020). Existing networks that address adaptive evolution on global scales include, for example, the Global Urban Evolution Project (GLUE, Santangelo et al. 2022), the PLANTPOPNET (Smith et al. 2020), and the Genomics of Rapid Evolution in Novel Environments-net (Czech et al. 2022). These networks bring together large amounts of data across broad spatio-environmental scales and involve experts from diverse backgrounds, enabling the integration of various perspectives and methodologies. Together these characteristics address three important issues when studying complex evolutionary questions. First, studying evolutionary ecology requires extensive datasets across many populations and time periods (Vermeulen et al. 2013). Second, biogeographic regions can fundamentally differ in how environmental changes affect adaptation (Exposito-Alonso et al. 2019) and how this affects biotic interactions (Lee et al. 2022). And third, evolutionary biology is inherently interdisciplinary, drawing from genetics, metabolomics, ecology, and other fields (Craven et al. 2019).
Here, we introduce iCONNECT, which is a new large collaborative and interdisciplinary network investigating mechanisms driving rapid evolution in C. canadensis (https://conyzaiconnect.wixsite.com/iconnect). This framework is an open collaboration of researchers who contribute to the sampling of populations from the Northern Hemisphere, and researchers who investigate APV on sampled populations in their particular field. We use C. canadensis as a model because it is a successful invader, and an economically significant agricultural weed (Okada et al. 2015). The species is native to large parts of North America and non-native to large parts of the rest of the temperate and subtropical world. This cosmopolitan distribution allows studying APV in biotic interactions across large climatic gradients. Conyza canadensis has a high capability for rapid evolution given by that it was the first eudicot weed that evolved glyphosate resistance, independently at multiple locations (Okada et al. 2015).
Our current research investigates the drivers of rapid evolution in the competitive ability of C. canadensis under mesic vs. drought-stressed conditions. To do so, we investigate how population history determines APV under common garden conditions for competitive ability, drought responses, root exudate profiles, allelopathic activity, resource acquisition patterns and, interactions with mutualistic and pathogenic fungi (Fig. 2). In addition, correlating data across the interdisciplinary work packages can help to unravel how belowground mechanisms determine competitive ability. This includes genome-wide association studies to identify genomic regions that drive rapid evolution in competitive ability.
Conclusion
The understanding of rapid evolution is still incomplete despite tremendous efforts that have been made with diverse approaches in common garden studies. There are only few studies on rapid evolution that explicitly disentangle how population history drives APV within and between native and non-native ranges. There is also a lack of interdisciplinary frameworks that could identify the underlying mechanisms of rapid evolution, particularly studies involving multi-omics approaches. In this conceptual paper, we reiterate the call for more sophisticated eco-evolutionary frameworks to study rapid evolution (e.g., Mráz et al. 2014; Schrieber et al. 2017; Rosche et al. 2018b, 2019) by focusing on three topics: (1) sampling, (2) design of experiments, and (3) experimental methods.
We argue that integrating data-intensive research on phenotypes, metabolomics, belowground amplicon sequencing and population genomics offers promising opportunities for studying complex biotic interactions. Considering these aspects, we outline how to study rapid evolution in the context of competitive ability under changing environmental conditions and introduce our scientific network iCONNECT. Our conceptual approach, however, is applicable to other invasive species and other types of biotic interactions and we emphasize the value of large collaborative networks to address such research. We hope that our considerations can be helpful for researchers that design studies on rapid evolution to test for differences between native vs. non-native populations.
References
Aikio S, Duncan RP, Hulme PE (2010) Lag-phases in alien plant invasions: separating the facts from the artefacts. Oikos 119(2):370–378. https://doi.org/10.1111/j.1600-0706.2009.17963.x
Alexander JM, Poll M, Dietz H, Edwards PJ (2009) Contrasting patterns of genetic variation and structure in plant invasions of mountains. Divers Distrib 15(3):502–512. https://doi.org/10.1111/j.1472-4642.2008.00555.x
Al-Gharaibeh MM, Hamasha HR, Rosche C, Lachmuth S, Wesche K, Hensen I (2017) Environmental gradients shape the genetic structure of two medicinal Salvia species in Jordan. Plant Biol 19(2):227–238. https://doi.org/10.1111/plb.12512
Andrews KR, Good JM, Miller MR, Luikart G, Hohenlohe PA (2016) Harnessing the power of RADseq for ecological and evolutionary genomics. Nat Rev Genet 17(2):81–92. https://doi.org/10.1038/nrg.2015.28
Angeloni F, Ouborg NJ, Leimu R (2011) Meta-analysis on the association of population size and life history with inbreeding depression in plants. Biol Cons 144(1):35–43. https://doi.org/10.1016/j.biocon.2010.08.016
Aschehoug ET, Brooker R, Atwater DZ, Maron JL, Callaway RM (2016) The mechanisms and consequences of interspecific competition among plants. Annu Rev Ecol Evol Syst 47:263–281. https://doi.org/10.1146/annurev-ecolsys-121415-032123
Auwerx C, Sadler MC, Woh T, Reymond A, Kutalik Z, Porcu E (2023) Exploiting the mediating role of the metabolome to unravel transcript-to-phenotype associations. Elife 12:e81097. https://doi.org/10.7554/eLife.81097
Barrett SC (2016) Foundations of invasion genetics: the Baker and Stebbins legacy. Invasion Genet Baker Stebbins Leg. https://doi.org/10.1002/9781119072799.ch1
Bieker VC, Battlay P, Petersen B, Sun X, Wilson J, Brealey JC et al (2022) Uncovering the genomic basis of an extraordinary plant invasion. Sci Adv 8(34):eabo5115. https://doi.org/10.1126/sciadv.abo5115
Blossey B, Notzold R (1995) Evolution of increased competitive ability in invasive nonindigenous plants: a hypothesis. J Ecol 83(5):887–889. https://doi.org/10.2307/2261425
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11. https://doi.org/10.1007/s00442-005-0070-z
Callaway RM, Lucero JE (2020) Soil biota and non-native plant invasions. Plant invasions: the role of biotic interactions. CABI, Wallingford, pp 45–66
Callaway RM, Waller LP, Diaconu A, Pal R, Collins AR, Mueller-Schaerer H, Maron JL (2011) Escape from competition: neighbors reduce Centaurea stoebe performance at home but not away. Ecology 92(12):2208–2213. https://doi.org/10.1890/11-0518.1
Callaway RM, Lucero JE, Hierro JL, Lortie CJ (2022) The EICA is dead? Long live the EICA! Ecol Lett 25(10):2289–2302. https://doi.org/10.1111/ele.14088
Cameron TC, O’Sullivan D, Reynolds A, Piertney SB, Benton TG (2013) Eco-evolutionary dynamics in response to selection on life-history. Ecol Lett 16:754–763. https://doi.org/10.1111/ele.12107
Capblancq T, Butnor JR, Deyoung S, Thibault E, Munson H, Nelson DM et al (2020) Whole-exome sequencing reveals a long-term decline in effective population size of red spruce (Picea rubens). Evol Appl 13(9):2190–2205. https://doi.org/10.1111/eva.12985
Clements D, DiTommaso A (2011) Climate change and weed adaptation: can evolution of invasive plants lead to greater range expansion than forecasted? Weed Res 51:227–240. https://doi.org/10.1111/J.1365-3180.2011.00850.X
Clements DR, Jones VL (2021) Rapid evolution of invasive weeds under climate change: present evidence and future research needs. Front Agron 3:664034. https://doi.org/10.3389/fagro.2021.664034
Colautti RI, Lau JA (2016) Contemporary evolution during invasion: evidence for differentiation, natural selection, and local adaptation. Invasion Genet Baker Stebbins Leg. https://doi.org/10.1111/mec.13162
Craven D, Winter M, Hotzel K, Gaikwad J, Eisenhauer N, Hohmuth M et al (2019) Evolution of interdisciplinarity in biodiversity science. Ecol Evol 9(12):6744–6755. https://doi.org/10.1002/ece3.5244
Czech L, Peng Y, Spence JP, Lang PL, Bellagio T, Hildebrandt J et al (2022) Monitoring rapid evolution of plant populations at scale with pool-sequencing. BioRxiv. https://doi.org/10.1101/2022.02.02.477408
De Kort H, Panis B, Helsen K, Douzet R, Janssens SB, Honnay O (2020) Pre-adaptation to climate change through topography-driven phenotypic plasticity. J Ecol 108(4):1465–1474. https://doi.org/10.1111/1365-2745.13365
de Villemereuil P, Gaggiotti OE, Mouterde M, Till-Bottraud I (2016) Common garden experiments in the genomic era : new perspectives and opportunities. Heredity 116(3):249–254. https://doi.org/10.1038/hdy.2015.93
de Villemereuil P, Gaggiotti OE, Goudet J (2022) Common garden experiments to study local adaptation need to account for population structure. J Ecol 110(5):1005–1009. https://doi.org/10.1111/1365-2745.13528
Des Roches S, Post DM, Turley NE, Bailey JK, Hendry AP, Kinnison MT et al (2018) The ecological importance of intraspecific variation. Nat Ecol Evol 2(1):57–64. https://doi.org/10.1038/s41559-017-0402-5
Dlugosch KM, Anderson SR, Braasch J, Cang FA, Gillette HD (2016) The devil is in the details: genetic variation in introduced populations and its contributions to invasion. Invasion Genet Baker Stebbins Leg. https://doi.org/10.1111/mec.13183
Dyer AR, Brown CS, Espeland EK, McKay JK, Meimberg H, Rice KJ (2010) The role of adaptive trans-generational plasticity in biological invasions of plants. Evol Appl 3:179–192. https://doi.org/10.1111/j.1752-4571.2010.00118.x
Encinas-Viso F, Morin L, Raghu S, Knerr N, Roux C et al (2022) Population genomics reveal multiple introductions and admixture of Sonchus oleraceus in Australia. Divers Distrib 28(9):1951–1965. https://doi.org/10.1111/ddi.13597
Erfmeier A, Bruelheide H (2010) Invasibility or invasiveness? Effects of habitat, genotype, and their interaction on invasive Rhododendron ponticum populations. Biol Invasions 12:657–676. https://doi.org/10.1007/s10530-009-9472-x
Eriksen RL, Desronvil T, Hierro J, Kesseli R (2012) Morphological differentiation in a common garden experiment among native and non-native specimens of the invasive weed yellow starthistle (Centaurea solstitialis). Biol Invasions 14:1459–1467. https://doi.org/10.1007/s10530-012-0172-6
Estoup A, Ravigné V, Hufbauer R, Vitalis R, Gautier M, Facon B (2016) Is there a genetic paradox of biological invasion? Annu Rev Ecol Evol Syst 47:51–72. https://doi.org/10.1146/annurev-ecolsys-121415-032116
Exposito-Alonso M, Burbano HA, Bossdorf O, Nielsen R, Weigel D (2019) Natural selection on the Arabidopsis thaliana genome in present and future climates. Nature 573(7772):126–129. https://doi.org/10.1038/s41586-019-1520-9
Fahrenkrog AM, Neves LG, Resende MF Jr, Vazquez AI, de Los CG, Dervinis C et al (2017) Genome-wide association study reveals putative regulators of bioenergy traits in Populus deltoides. New Phytol 213(2):799–811. https://doi.org/10.1111/nph.14154
Felker-Quinn E, Schweitzer JA, Bailey JK (2013) Meta-analysis reveals evolution in invasive plant species but little support for evolution of increased competitive ability (EICA). Ecol Evol 3(3):739–751. https://doi.org/10.1002/ece3.488
Ferrero V, Navarro L, Castro S, Loureiro J, Sánchez JM, Carvallo GO, Barrett SC (2020) Global patterns of reproductive and cytotype diversity in an invasive clonal plant. Biol Invasions 22(5):1691–1703. https://doi.org/10.1007/s10530-020-02213-9
Fitzpatrick B, Fordyce JA, Niemiller ML, Reynolds RG (2012) What can DNA tell us about biological invasions? Biol Invasions 14:245–253. https://doi.org/10.1007/s10530-011-0064-1
Flory S, Kleczewski N, Clay K (2011) Ecological consequences of pathogen accumulation on an invasive grass. Ecosphere 2:1–12. https://doi.org/10.1890/ES11-00191.1
Flucher SM, Krapf P, Arthofer W, Suarez AV, Crozie RH, Steiner FM, Schlick-Steiner BC (2021) Effect of social structure and introduction history on genetic diversity and differentiation. Mol Ecol 30(11):2511–2527. https://doi.org/10.1111/mec.15911
Franks SJ, Weber JJ, Aitken SN (2014) Evolutionary and plastic responses to climate change in terrestrial plant populations. Evol Appl 7(1):123–139. https://doi.org/10.1111/eva.12112
Franks SJ, Hamann E, Weis AE (2018) Using the resurrection approach to understand contemporary evolution in changing environments. Evol Appl 11:17–28. https://doi.org/10.1111/eva.12528
Goring SJ, Weathers KC, Dodds WK, Soranno PA, Sweet LC, Cheruvelil KS et al (2014) Improving the culture of interdisciplinary collaboration in ecology by expanding measures of success. Front Ecol Environ 12(1):39–47. https://doi.org/10.1890/120370
Goss EM, Kendig AE, Adhikari A, Lane B, Kortessis N, Holt RD et al (2020) Disease in invasive plant populations. Annu Rev Phytopathol 58:97–117. https://doi.org/10.1146/annurev-phyto-010820-012757
Guo WY, Lambertini C, Guo X, Li XZ, Eller F, Brix H (2016) Phenotypic traits of the Mediterranean Phragmites australis M1 lineage: differences between the native and introduced ranges. Biol Invasions 18:2551–2561. https://doi.org/10.1007/s10530-016-1236-9
Gustafsson ALS, Gussarova G, Borgen L, Ikeda H, Antonell A, Marie-Orleach L et al (2022) Rapid evolution of post-zygotic reproductive isolation is widespread in Arctic plant lineages. Ann Bot 129(2):171–184. https://doi.org/10.1093/aob/mcab128
Harmon-Threatt AN, Burns JH, Shemyakina LA, Knigh TM (2009) Breeding system and pollination ecology of introduced plants compared to their native relatives. Am J Bot 96(8):1544–1550. https://doi.org/10.3732/ajb.0800369
He WM, Thelen GC, Ridenour WM, Callaway RM (2010) Is there a risk to living large? Large size correlates with reduced growth when stressed for knapweed populations. Biol Invasions 12:3591–3598. https://doi.org/10.1007/s10530-010-9753-4
Heger T (2016) Light availability experienced in the field affects ability of following generations to respond to shading in an annual grassland plant. J Ecol 104(5):1432–1440. https://doi.org/10.1111/1365-2745.12607
Heger T, Jacobs BS, Latimer AM, Kollmann J, Rice KJ (2014) Does experience with competition matter? Effects of source competitive environment on mean and plastic trait expression in Erodium cicutarium. Perspect Plant Ecol, Evol Syst 16(5):236–246. https://doi.org/10.1016/j.ppees.2014.06.002
Hendry AP (2016) Eco-evolutionary dynamics. Eco-evolutionary dynamics. Princeton University Press, Princeton. https://doi.org/10.1515/9781400883080
Hendry AP, Kinnison MT (1999) Perspective: the pace of modern life: measuring rates of contemporary microevolution. Evolution 53(6):1637–1653. https://doi.org/10.1111/j.1558-5646.1999.tb04550.x
Herman JJ, Sultan SE (2011) Adaptive transgenerational plasticity in plants: case studies, mechanisms, and implications for natural populations. Front Plant Sci 2:102. https://doi.org/10.3389/fpls.2011.00102
Hernández F, Poverene M, Garayalde A, Presotto A (2019) Re-establishment of latitudinal clines and local adaptation within the invaded area suggest rapid evolution of seed traits in Argentinean sunflower (Helianthus annuus L.). Biol Invasions 21:2599–2612. https://doi.org/10.1007/s10530-019-01998-8
Hierro JL, Callaway RM (2021) The ecological importance of allelopathy. Annu Rev Ecol Evol Syst 52:25–45. https://doi.org/10.1146/annurev-ecolsys-051120-030619
Hoffbeck C, TerHorst CP (2022) Trait differences between and within ranges of an invasive legume species. Biol Invasions 24(9):2873–2883. https://doi.org/10.21203/rs.3.rs-1197357/v1
Hoffmann AA, Sgrò CM (2011) Climate change and evolutionary adaptation. Nature 470:479–485. https://doi.org/10.1038/nature09670
Hou Z, Li A (2020) Population genomics reveals demographic history and genomic differentiation of Populus davidiana and Populus tremula. Front Plant Sci 11:553736. https://doi.org/10.3389/fpls.2020.01103
Hu XT, Dong BC (2019) Herbivory and nitrogen availability affect performance of an invader Alternanthera philoxeroides and its native congener A. sessilis. Flora 257:151412. https://doi.org/10.1016/j.flora.2019.05.011
Irimia RE, Lopes SM, Sotes G, Cavieres LA, Eren Ö, Lortie CJ et al (2019) Biogeographic differences in the allelopathy of leaf surface extracts of an invasive weed. Biol Invasions 21:3151–3168. https://doi.org/10.1007/s10530-019-02038-1
Irimia RE, Montesinos D, Chaturvedi A, Sanders I, Hierro JL, Sotes G et al (2023) Trait evolution during a rapid global weed invasion despite little genetic differentiation. Evol Appl. https://doi.org/10.1111/eva.13548
Javed Q, Sun J, Azeem A, Jabran K, Du D (2020) Competitive ability and plasticity of Wedelia trilobata (L.) under wetland hydrological variations. Sci Rep 10(1):1–11. https://doi.org/10.1038/s41598-020-66385-z
Jeschke JM, Heger T (eds) (2018) Invasion biology: hypotheses and evidence, vol 9. CABI, Wallingford
Jones EI, Gomulkiewicz R (2012) Biotic interactions, rapid evolution, and the establishment of introduced species. Am Nat 179(2):E28–E36. https://doi.org/10.1086/663678
Keane RM, Crawley MJ (2002) Exotic plant invasions and the enemy release hypothesis. Trends Ecol Evol 17(4):164–170. https://doi.org/10.1016/S0169-5347(02)02499-0
Keller SR, Taylor DR (2008) History, chance and adaptation during biological invasion: separating stochastic phenotypic evolution from response to selection. Ecol Lett 11(8):852–866. https://doi.org/10.1111/j.1461-0248.2008.01188.x
Kuehn I, Dormann CF (2012) Less than eight (and a half) misconceptions of spatial analysis. J Biogeogr 39(5):995–998. https://doi.org/10.1111/j.1365-2699.2012.02707.x
Kulmatiski A (2019) Plant-soil feedbacks predict native but not non-native plant community composition: a 7-year common-garden experiment. Front Ecol Evol 7:326. https://doi.org/10.3389/fevo.2019.00326
Kulmatiski A, Beard KH, Stevens JR, Cobbold SM (2008) Plant–soil feedbacks: a meta-analytical review. Ecol Lett 11(9):980–992. https://doi.org/10.1111/j.1461-0248.2008.01209.x
Lachmuth S, Durka W, Schurr FM (2011) Differentiation of reproductive and competitive ability in the invaded range of Senecio inaequidens: the role of genetic Allee effects, adaptive and nonadaptive evolution. New Phytol 192(2):529–541. https://doi.org/10.1111/j.1469-8137.2011.03808.x
Lee BR, Miller TK, Rosche C, Yang Y, Heberling JM, Kuebbing SE, Primack RB (2022) Wildflower phenological escape differs by continent and spring temperature. Nat Commun 13(1):7157. https://doi.org/10.1038/s41467-022-34936-9
Legendre P (1993) Spatial autocorrelation: trouble or new paradigm? Ecology 74(6):1659–1673. https://doi.org/10.2307/1939924
Lekberg Y, Bever JD, Bunn RA, Callaway RM, Hart MM, Kivlin SN et al (2018) Relative importance of competition and plant–soil feedback, their synergy, context dependency and implications for coexistence. Ecol Lett 21(8):1268–1281. https://doi.org/10.1111/ele.13093
Levy SE, Boone BE (2019) Next-generation sequencing strategies. Cold Spring Harbor Perspect Med. https://doi.org/10.1101/cshperspect.a025791
Luo X, Xu X, Zheng Y, Guo H, Hu S (2019) The role of phenotypic plasticity and rapid adaptation in determining invasion success of Plantago virginica. Biol Invasions 21:2679–2692. https://doi.org/10.1007/s10530-019-02004-x
Macel M, de Vos RCH, Jansen JJ, van der Putten WH, van Dam NM (2014) Novel chemistry of invasive plants: exotic species have more unique metabolomic profiles than native congeners. Ecol Evol 4:2777–2786. https://doi.org/10.1002/ece3.1132
Macel M, Dostálek T, Esch S, Bucharová A, van Dam NM, Tielbörger K, Verhoeven KJF, Münzbergová Z (2017) Evolutionary responses to climate change in a range expanding plant. Oecologia 184:543–554. https://doi.org/10.1007/s00442-017-3864-x
Mackin CR, Peña JF, Blanco MA, Balfou NJ, Castellanos MC (2021) Rapid evolution of a floral trait following acquisition of novel pollinators. J Ecol 109(5):2234–2246. https://doi.org/10.1111/1365-2745.13636
McCulloch G, Gurdasani K, Hereward J, Morin L, Walter G, Raghu S (2023) Invasion history of Lycium ferocissimum in Australia: the impact of admixture on genetic diversity and differentiation. Divers Distrib. https://doi.org/10.1111/ddi.13702
McKown AD, Klápště J, Geraldes A, Friedmann M, Cronk QC et al (2014) Geographical and environmental gradients shape phenotypic trait variation and genetic structure in Populus trichocarpa. New Phytol 201(4):1263–1276. https://doi.org/10.1111/nph.12601
Milcu A, Puga-Freitas R, Ellison AM, Blouin M, Scheu S, Freschet GT et al (2018) Genotypic variability enhances the reproducibility of an ecological study. Nat Ecol Evol 2(2):279–287. https://doi.org/10.1038/s41559-017-0434-x
Mimura M, Ono K, Goka K, Hara T (2013) Standing variation boosted by multiple sources of introduction contributes to the success of the introduced species, Lotus corniculatus. Biol Invasions 15:2743–2754. https://doi.org/10.1007/s10530-013-0488-x
Moles AT, Dalrymple RL, Raghu S, Bonser SP, Ollerton J (2022) Advancing the missed mutualist hypothesis, the under-appreciated twin of the enemy release hypothesis. Biol Lett 18(10):20220220
Mönchgesang S, Strehmel N, Schmidt S, Westphal L, Taruttis F, Müller E et al (2016) Natural variation of root exudates in Arabidopsis thaliana-linking metabolomic and genomic data. Sci Rep 6(1):29033. https://doi.org/10.1038/srep29033
Montesinos D (2022) Fast invasives fastly become faster: Invasive plants align largely with the fast side of the plant economics spectrum. J Ecol 110(5):1010–1014. https://doi.org/10.1111/1365-2745.13616
Montesinos D, Graebner RC, Callaway RM (2019) Evidence for evolution of increased competitive ability for invasive Centaurea solstitialis, but not for naturalized C. calcitrapa. Biol Invasions 21:99–110. https://doi.org/10.1007/s10530-018-1807-z
Moran EV, Alexander JM (2014) Evolutionary responses to global change: lessons from invasive species. Ecol Lett 17(5):637–649. https://doi.org/10.1111/ele.12262
Mráz P, Tarbush E, Müller-Schärer H (2014) Drought tolerance and plasticity in the invasive knapweed Centaurea stoebe s.l. (Asteraceae): effect of populations stronger than those of cytotype and range. Ann Bot 114(2):289–299. https://doi.org/10.1093/aob/mcu105
Nagy DU, Stranczinger S, Godi A, Weisz A, Rosche C, Suda J et al (2018) Does higher ploidy level increase the risk of invasion? A case study with two geo-cytotypes of Solidago gigantea Aiton (Asteraceae). J Plant Ecol 11(2):317–327. https://doi.org/10.1093/jpe/rtx005
Nagy DU, Rauschert ESJ, Callaway RM, Henn T, Filep R, Pal RW (2022) Intense mowing management suppresses invader, but shifts competitive resistance by a native to facilitation. Restor Ecol 30(1):e13483. https://doi.org/10.1111/rec.13483
Ochocki BM, Miller TE (2017) Rapid evolution of dispersal ability makes biological invasions faster and more variable. Nat Commun 8(1):1–8. https://doi.org/10.1038/ncomms14315
Oduor AM, Adomako MO, Yuan Y, Li JM (2022) Older populations of the invader Solidago canadensis exhibit stronger positive plant-soil feedbacks and competitive ability in China. Am J Bot 109(8):1230–1241. https://doi.org/10.1002/ajb2.16034
Okada M, Hanson BD, Hembree KJ, Peng Y, Shrestha A, Stewart CN Jr et al (2015) Evolution and spread of glyphosate resistance in Conyza bonariensis in California and a comparison with closely related Conyza canadensis. Weed Res 55(2):173–184. https://doi.org/10.1111/wre.12131
Orsini L, Vanoverbeke J, Swillen I, Mergeay J, De Meester L (2013) Drivers of population genetic differentiation in the wild: isolation by dispersal limitation, isolation by adaptation and isolation by colonization. Mol Ecol 22(24):5983–5999. https://doi.org/10.1111/mec.12561
Osunkoya O, Lock CB, Dhileepan K, Buru JC (2021) Lag times and invasion dynamics of established and emerging weeds: Insights from herbarium records of Queensland, Australia. Biol Invasions 23:3383–3408. https://doi.org/10.1007/s10530-021-02581-w
Pal RW, Maron J, Nagy DU, Waller L, Tosto A, Liao H, Callaway RM (2020) What happens in Europe stays in Europe: apparent evolution by an invader does not help at home. Ecology 101(8):e03072. https://doi.org/10.1002/ecy.3072
Papale F, Saget J, Bapteste É (2020) Networks consolidate the core concepts of evolution by natural selection. Trends Microbiol 28(4):254–265. https://doi.org/10.1016/j.tim.2019.11.006
Pearse IS, Sofaer HR, Zaya DN, Spyreas G (2019) Non-native plants have greater impacts because of differing per-capita effects and nonlinear abundance–impact curves. Ecol Lett 22(8):1214–1220. https://doi.org/10.1111/ele.13284
Peng Y, Yang JX, Zhou XH, Peng PH, Li JJ, Zhang SM, He WM (2019) An invasive population of Solidago canadensis is less sensitive to warming and nitrogen-addition than its native population in an invaded range. Biol Invasions 21:151–162. https://doi.org/10.1007/s10530-018-1812-2
Peters K, Worrich A (2018) Current challenges in plant eco-metabolomics. Int J Mol Sci 19(5):1385. https://doi.org/10.3390/ijms19051385
Pico FX, Ouborg NJ, Van Groenendael JM (2003) Fitness traits and dispersal ability in the herb Tragopogon pratensis (Asteraceae): decoupling the role of inbreeding depression and maternal effects. Plant Biol 5(5):522–530. https://doi.org/10.1055/s-2003-44788
Qing H, Cai Y, Xiao Y, Yao Y, An S (2012) Leaf nitrogen partition between photosynthesis and structural defense in invasive and native tall form Spartina alterniflora populations: effects of nitrogen treatments. Biol Invasions 14:2039–2048. https://doi.org/10.1007/s10530-012-0210-4
Raeymaekers JA, Chaturvedi A, Hablützel PI, Verdonck I, Hellemans B, Maes GE et al (2017) Adaptive and non-adaptive divergence in a common landscape. Nat Commun 8(1):267. https://doi.org/10.1038/s41467-017-00256-6
Rauschkolb R, Li Z, Godefroid S, Dixon L, Durka W, Májeková M et al (2022) Evolution of plant drought strategies and herbivore tolerance after two decades of climate change. New Phytol 235(2):773–785. https://doi.org/10.1111/nph.18125
Ravinet M, Faria R, Butlin RK, Galindo J, Bierne N, Rafajlović M et al (2017) Interpreting the genomic landscape of speciation: a road map for finding barriers to gene flow. J Evol Biol 30(8):1450–1477. https://doi.org/10.1111/jeb.13047
Reinhart KO, Callaway RM (2006) Soil biota and invasive plants. New Phytol 170(3):445–457. https://doi.org/10.1111/j.1469-8137.2006.01715.x
Rius M, Darling JA (2014) How important is intraspecific genetic admixture to the success of colonising populations? Trends Ecol Evol 29(4):233–242. https://doi.org/10.1016/j.tree.2014.02.003
Rosche C, Durka W, Hensen I, Mráz P, Hartmann M, Müller-Schärer H, Lachmuth S (2016) The population genetics of the fundamental cytotype-shift in invasive Centaurea stoebe sl: genetic diversity, genetic differentiation and small-scale genetic structure differ between cytotypes but not between ranges. Biol Invasions 18:1895–1910
Rosche C, Hensen I, Mraz P, Durka W, Hartmann M, Lachmuth S (2017) Invasion success in polyploids: the role of inbreeding in the contrasting colonization abilities of diploid versus tetraploid populations of Centaurea stoebe sl. J Ecol 105(2):425–435. https://doi.org/10.1111/1365-2745.12670
Rosche C, Heinicke S, Hensen I, Silantyeva MM, Stolz J, Gröning S, Wesche K (2018a) Spatio-environmental determinants of the genetic structure of three steppe species in a highly fragmented landscape. Basic Appl Ecol 28:48–59. https://doi.org/10.1016/j.baae.2018.02.001
Rosche C, Hensen I, Lachmuth S (2018b) Local pre-adaptation to disturbance and inbreeding–environment interactions affect colonization abilities of diploid and tetraploid Centaurea stoebe. Plant Biol 20(1):75–84. https://doi.org/10.1111/plb.12628
Rosche C, Hensen I, Schaar A, Zehra U, Jasieniuk M, Callaway RM et al (2019) Climate outweighs native vs. nonnative range-effects for genetics and common garden performance of a cosmopolitan weed. Ecol Monogr 89(4):e01386. https://doi.org/10.1002/ecm.1386
Rotter MC, Holeski LM (2018) A meta-analysis of the evolution of increased competitive ability hypothesis: genetic-based trait variation and herbivory resistance trade-offs. Biol Invasions 20:2647–2660. https://doi.org/10.1007/s10530-018-1724-1
Rudman SM, Barbour MA, Csilléry K, Gienapp P, Guillaume F, Hairston NG Jr et al (2018) What genomic data can reveal about eco-evolutionary dynamics. Nat Ecol Evol 2(1):9–15. https://doi.org/10.1038/s41559-017-0385-2
Santangelo JS, Ness RW, Cohan B, Fitzpatrick CR, Innes SG, Koch S et al (2022) Global urban environmental change drives adaptation in white clover. Science 375(6586):1275–1281. https://doi.org/10.1126/science.abk0989
Schlötterer C, Kofler R, Versace E, Tobler R, Franssen SU (2015) Combining experimental evolution with next-generation sequencing: a powerful tool to study adaptation from standing genetic variation. Heredity 114(5):431–440. https://doi.org/10.1038/hdy.2014.86
Schrieber K, Lachmuth S (2017) The genetic paradox of invasions revisited: the potential role of inbreeding × environment interactions in invasion success. Biol Rev 92(2):939–952. https://doi.org/10.1111/brv.12263
Schrieber K, Wolf S, Wypior C, Höhlig D, Hensen I, Lachmuth S (2017) Adaptive and non-adaptive evolution of trait means and genetic trait correlations for herbivory resistance and performance in an invasive plant. Oikos 126(4):572–582. https://doi.org/10.1111/oik.03781
Semchenko M, Saar S, Lepik A (2014) Plant root exudates mediate neighbour recognition and trigger complex behavioural changes. New Phytol 204(3):631–637. https://doi.org/10.1111/nph.12930
Shah MA, Callaway RM, Shah T, Houseman GR, Pal RW, Xiao S et al (2014) Conyza canadensis suppresses plant diversity in its nonnative ranges but not at home: a transcontinental comparison. New Phytol 202(4):1286–1296. https://doi.org/10.1111/nph.12733
Sheng M, Rosche C, Al-Gharaibeh M, Bullington LS, Callaway RM, Clark T et al (2022) Acquisition and evolution of enhanced mutualism—an underappreciated mechanism for invasive success? ISME J 16(11):2467–2478. https://doi.org/10.1038/s41396-022-01293-w
Shi J, Stahl M, de Vos RC, Tielbörger K, Verhoeven KJ, Macel M (2023) Metabolomic profiling reveals shifts in defenses of an invasive plant. Biol Invasions. https://doi.org/10.1007/s10530-023-03109-0
Smith AL, Hodkinson TR, Villellas J, Catford JA, Csergő AM, Blomberg SP et al (2020) Global gene flow releases invasive plants from environmental constraints on genetic diversity. Proc Natl Acad Sci 117(8):4218–4227. https://doi.org/10.1073/pnas.1915848117
Sultan SE, Horgan-Kobelski T, Nichols LM, Riggs CE, Waples RK (2013) A resurrection study reveals rapid adaptive evolution within populations of an invasive plant. Evol Appl 6(2):266–278. https://doi.org/10.1111/j.1752-4571.2012.00287.x
Szűcs M, Vahsen ML, Melbourne BA, Hoover C, Weiss-Lehman C, Hufbauer RA (2017) Rapid adaptive evolution in novel environments acts as an architect of population range expansion. Proc Natl Acad Sci 114(51):13501–13506. https://doi.org/10.1073/pnas.171293411
Tavares D, Loureiro J, Martins A, Castro M, Roiloa S, Castro S (2019) Genetically based phenotypic differentiation between native and introduced tetraploids of Oxalis pes-caprae. Biol Invasions 21(1):229–243. https://doi.org/10.1007/s10530-018-1820-2
Thompson JN (1998) Rapid evolution as an ecological process. Trends Ecol Evol 13(8):329–332. https://doi.org/10.1016/S0169-5347(98)01378-0
Turcotte MM, Reznick DN, Hare JD (2011) The impact of rapid evolution on population dynamics in the wild: experimental test of eco-evolutionary dynamics. Ecol Lett 14(11):1084–1092. https://doi.org/10.1111/j.1461-0248.2011.01676.x
Turner TL, Stewart AD, Fields AT, Rice WR, Tarone AM (2011) Population-based resequencing of experimentally evolved populations reveals the genetic basis of body size variation in Drosophila melanogaster. PLoS Genet 7:e1001336. https://doi.org/10.1371/journal.pgen.1001336
Turner KG, Nurkowski KA, Rieseberg LH (2017) Gene expression and drought response in an invasive thistle. Biol Invasions 19:875–893. https://doi.org/10.1007/s10530-016-1308-x
van Dam NM, Bouwmeester HJ (2016) Metabolomics in the rhizosphere: tapping into belowground chemical communication. Trends Plant Sci 21(3):256–265. https://doi.org/10.1016/j.tplants.2016.01.008
van Kleunen M, Bossdorf O, Dawson W (2018) The ecology and evolution of alien plants. Annu Rev Ecol Evol Syst 49:25–47. https://doi.org/10.1146/annurev-ecolsys-110617-062654
Vandegrift R, Blaser W, Campos-Cerda F, Heneghan AF, Carroll GC, Roy BA (2015) Mixed fitness effects of grass endophytes modulate impact of enemy release and rapid evolution in an invasive grass. Biol Invasions 17:1239–1251. https://doi.org/10.1007/s10530-014-0791-1
Vendrami DL, De Noia M, Telesca L, Handal W, Charrier G, Boudry P et al (2019) RAD sequencing sheds new light on the genetic structure and local adaptation of European scallops and resolves their demographic histories. Sci Rep 9(1):7455. https://doi.org/10.1038/s41598-019-43939-4
Vermeulen N, Parker JN, Penders B (2013) Understanding life together: a brief history of collaboration in biology. Endeavour 37(3):162–171. https://doi.org/10.1016/j.endeavour.2013.03.001
Veselá A, Hadincová V, Vandvik V, Münzbergová Z (2021) Maternal effects strengthen interactions of temperature and precipitation, determining seed germination of dominant alpine grass species. Am J Bot 108(5):798–810. https://doi.org/10.1002/ajb2.1657
Villellas J, EhrlénJ CEE, Csergő AM, Garcia MB, Laine AL et al (2021) Phenotypic plasticity masks range-wide genetic differentiation for vegetative but not reproductive traits in a short-lived plant. Ecol Lett 24(11):2378–2393. https://doi.org/10.1111/ele.13858
Walker TWN, Alexander JM, Allard P-M, Baines O, Baldy V, Bardgett RD et al (2022) Functional traits 2.0: the power of the metabolome for ecology. J Ecol 110:4–20. https://doi.org/10.1111/1365-2745.13826
Walker TWN, Schrodt F, Allard P-M, Defossez E, Jassey VEJ, Schuman MC et al (2023) Leaf metabolic traits reveal hidden dimensions of plant form and function. Sci Adv 9(35):eadi02. https://doi.org/10.1126/sciadv.adi4029
Walsh J, Kovach AI, Benham PM, Clucas GV, Winder VL, Lovette IJ (2021) Genomic data reveal the biogeographical and demographic history of Ammospiza sparrows in northeast tidal marshes. J Biogeogr 48(9):2360–2374. https://doi.org/10.1111/jbi.14208
Weidenhamer JD, Callaway RM (2010) Direct and indirect effects of invasive plants on soil chemistry and ecosystem function. J Chem Ecol 36(1):59–69. https://doi.org/10.1007/s10886-009-9735-0
Williams JL, Kendall BE, Levine JM (2016) Rapid evolution accelerates plant population spread in fragmented experimental landscapes. Science 353(6298):482–485. https://doi.org/10.1126/science.aaf6268
Woods EC, Sultan SE (2022) Post-introduction evolution of a rapid life-history strategy in a newly invasive plant. Ecology 103(11):e3803. https://doi.org/10.1002/ecy.3803
Wu Y, Colautti RI (2022) Evidence for continent-wide convergent evolution and stasis throughout 150 y of a biological invasion. Proc Natl Acad Sci 119(18):e2107584119. https://doi.org/10.1073/pnas.2107584119
Wu M, Ge Y, Xu C, Wang J (2020) Metabolome and transcriptome analysis of hexaploid Solidago canadensis roots reveals its invasive capacity related to polyploidy. Genes 11(2):187. https://doi.org/10.3390/genes11020187
Wu S, Chen L, Zhou Y, Xiao F, Liu D, Wang Y (2022) Invasive plants have higher resistance to native generalist herbivores than exotic noninvasive congeners. Environ Entomol 52:81–87. https://doi.org/10.1093/ee/nvac108
Yang X, Cheng J, Yao B, Lu H, Zhang Y, Xu J et al (2021) Polyploidy-promoted phenolic metabolism confers the increased competitive ability of Solidago canadensis. Oikos 130(6):1014–1025. https://doi.org/10.1111/oik.08280
Yin W, Zhou L, Yang K, Fang J, Biere A, Callaway RM et al (2023) Rapid evolutionary trade-offs between resistance to herbivory and tolerance to abiotic stress in an invasive plant. Ecol Lett. https://doi.org/10.1111/ele.14221
Yu H, He Y, Zhang W, Chen L, Zhang J, Zhang X et al (2022) Greater chemical signaling in root exudates enhances soil mutualistic associations in invasive plants compared to natives. New Phytol 236:1140–1153. https://doi.org/10.1111/nph.18289
Zhang Z, Liu Y, Hardrath A, Jin H, van Kleunen M (2022) Increases in multiple resources promote competitive ability of naturalized non-native plants. Commun Biol 5(1):1150. https://doi.org/10.1038/s42003-022-04113-1
Zheng YL, Feng YL, Zhang LK, Callaway RM, Valiente-Banuet A, Luo DQ et al (2015) Integrating novel chemical weapons and evolutionarily increased competitive ability in success of a tropical invader. New Phytol 205(3):1350–1359. https://doi.org/10.1111/nph.13135
Acknowledgements
We thank Heidi Hirsch for her help in developing the illustrations. We also gratefully acknowledge the financial support of the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) and the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig within the framework of Flexpool. The associate editor and two anonymous reviewers provided very helpful criticisms of earlier versions of the manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation; Project project number: RO 6418/1-1) and iDiv Flexpool funding (grant number W47038118). The German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig is funded by the German Research Foundation (DFG–FZT 118, 202548816).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. LMS wrote the manuscript with support from CR and IH, and substantial input from all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lucas, M.S., Hensen, I., Barratt, C.D. et al. Re-focusing sampling, design and experimental methods to assess rapid evolution by non-native plant species. Biol Invasions 26, 1327–1343 (2024). https://doi.org/10.1007/s10530-024-03249-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-024-03249-x