Abstract
Objective
To improve the activity of a water-forming NADH oxidase from Lactobacillus rhamnosus under neutral or alkaline pH for coupling NAD+-dependent dehydrogenases with an alkaline optimal pH.
Results
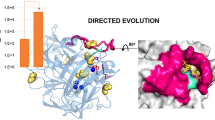
The water-forming NADH oxidase from Lactobacillus rhamnosus was engineered by replacing the aspartic acid or glutamic acid with arginine on the surface. The mutant D251R improved the activity with a 112%, 111%, and 244% relative activity to the wild-type at pH 6.5, pH 7.0, and pH 7.5, respectively. Docking substrate into the D251R mutant reveals that the NADH is access to the substrate-binding site with a larger substrate loop due to the enhanced electrostatic repulsion between ARG-251 and ARG-243. In the D251R-NADH complex, the carboxyl of NADH additionally forms two hydrogen bonds (2.6 and 2.9 Å) with G154 due to the changed interaction of substrate and the residues in the catalytic sites, and the hydrogen bond with the oxygen of carbonyl in P295 is shortened from 2.9 to 2.0 Å, which could account for the enhanced specific activity.
Conclusions
The D251R mutant displayed higher catalytic activity than the wild-type in the pH range 6.5–7.5, and further insight into those shorter and newly formed hydrogen bonds in substrate docking analysis could account for the higher bind affinity and catalytic efficiency of D251R mutant.





Similar content being viewed by others

References
Barin R, Biria D, Rashid-Nadimi S, Asadollahi MA (2018) Enzymatic CO2 reduction to formate by formate dehydrogenase from Candida boidinii coupling with direct electrochemical regeneration of NADH. J CO2 Util 28:117–125. https://doi.org/10.1016/j.jcou.2018.09.020
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye Binding. Anal Biochem 72:248–254
Case CL, Rodriguez JR, Mukhopadhyay B (2009) Characterization of an NADH oxidase of the flavin-dependent disulfide reductase family from Methanocaldococcus jannaschii. Microbiol-SGM 155:69–79. https://doi.org/10.1099/mic.0.024265-0
Cui D, Zhang L, Jiang S et al (2015) A computational strategy for altering an enzyme in its cofactor preference to NAD(H) and/or NADP(H). Febs J 282:2339–2351. https://doi.org/10.1111/febs.13282
Gao H, Tiwari MK, Kang YC, Lee J-K (2012) Characterization of H2O-forming NADH oxidase from Streptococcus pyogenes and its application in L-rare sugar production. Bioorg Med Chem Lett 22:1931–1935. https://doi.org/10.1016/j.bmcl.2012.01.049
Goren Z, Lapidot N, Willner I (1988) Photocatalysed regeneration of NAD(P)H by CdS and TiO2 semiconductors: applications in enzymatic synthesis. J Mol Catal. https://doi.org/10.1016/0304-5102(88)85069-7
Hummel W, Riebel B (2003) Isolation and biochemical characterization of a new NADH oxidase from Lactobacillus brevis. Biotechnol Lett 25:51–54. https://doi.org/10.1023/A:1021730131633
Kengen SWM, van der Oost J, de Vos WM (2003) Molecular characterization of H2O2-forming NADH oxidases from Archaeoglobus fulgidus. Eur J Biochem 270:2885–2894. https://doi.org/10.1046/j.1432-1033.2003.03668.x
Li F, Li Y-X, Cao Y-X et al (2018) Modular engineering to increase intracellular NAD (H/(+)) promotes rate of extracellular electron transfer of Shewanella oneidensis. Nat Commun 9:3637. https://doi.org/10.1038/s41467-018-05995-8
Li F-L, Zhou Q, Wei W et al (2019a) Switching the substrate specificity from NADH to NADPH by a single mutation of NADH oxidase from Lactobacillus rhamnosus. Int J Biol Macromol 135:328–336. https://doi.org/10.1016/j.ijbiomac.2019.05.146
Li F-L, Zhuang M-Y, Shen J-J et al (2019b) Specific Immobilization of Escherichia coli expressing recombinant glycerol dehydrogenase on mannose-functionalized magnetic nanoparticles. Catalysts 9:585. https://doi.org/10.3390/catal9070585
Li Q, Jiang T, Liu R et al (2019c) Tuning the pH profile of β-glucuronidase by rational site-directed mutagenesis for efficient transformation of glycyrrhizin. Appl Microbiol Biotechnol 103:4813–4823. https://doi.org/10.1007/s00253-019-09790-3
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Russell AJ, Fersht AR (1987) Rational modification of enzyme catalysis by engineering surface charge. Nature 328:496–500. https://doi.org/10.1038/328496a0
Saba T, Burnett JWH, Li J et al (2020) Assessing the environmental performance of NADH regeneration methods: a cleaner process using recyclable Pt/Fe3O4 and hydrogen. Catal Today 339:281–288. https://doi.org/10.1016/j.cattod.2019.01.049
Toomey D, Mayhew SG (1998) Purification and characterisation of NADH oxidase from Thermus aquaticus YT-1 and evidence that it functions in a peroxide-reduction system. Eur J Biochem 251:935–945. https://doi.org/10.1046/j.1432-1327.1998.2510935.x
Trott O, Olson AJ (2010) Software news and update AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
Turunen O, Vuorio M, Fenel F, Leisola M (2002) Engineering of multiple arginines into the Ser/Thr surface of Trichoderma reesei endo-1,4-β-xylanase II increases the thermotolerance and shifts the pH optimum towards alkaline pH. Protein Eng Des Sel 15:141–145. https://doi.org/10.1093/protein/15.2.141
Usher KC, Remington SJ, Martin DP, Drueckhammer DG (1994) A very short hydrogen bond provides only moderate stabilization of an enzyme-inhibitor complex of citrate synthase. Biochemistry 33:7753–7759. https://doi.org/10.1021/bi00191a002
Vidal LS, Kelly CL, Mordaka PM, Heap JT (2018) Review of NAD(P)H-dependent oxidoreductases: properties, engineering and application. Biochim Biophys Acta-Proteins Proteom 1866:327–347. https://doi.org/10.1016/j.bbapap.2017.11.005
Wang T, Qiu A, Meng F, Zhou H (2009) Changing the metal binding specificity of superoxide dismutase from Thermus thermophilus HB-27 by a single mutation. Mol Biotechnol 42:146–153. https://doi.org/10.1007/s12033-009-9149-9
Wang T, Ma F, Ma X, Wang P (2015) Spatially programmed assembling of oxidoreductases with single-stranded DNA for cofactor-required reactions. Appl Microbiol Biotechnol 99:3469–3477. https://doi.org/10.1007/s00253-014-6172-y
Xu M-Q, Wang S-S, Li L-N et al (2018) Combined cross-linked enzyme aggregates as biocatalysts. Catalysts 8:460. https://doi.org/10.3390/catal8100460
Xu M-Q, Li F-L, Yu W-Q et al (2020) Combined cross-linked enzyme aggregates of glycerol dehydrogenase and NADH oxidase for high efficiency in situ NAD+ regeneration. Int J Biol Macromol 144:1013–1021. https://doi.org/10.1016/j.ijbiomac.2019.09.178
Yokota K, Satou K, Ohki S (2006) Comparative analysis of protein thermo stability: differences in amino acid content and substitution at the surfaces and in the core regions of thermophilic and mesophilic proteins. Sci Technol Adv Mater 7:255–262. https://doi.org/10.1016/j.stam.2006.03.003
Yuan M, Kummer MJ, Milton RD et al (2019) Efficient NADH regeneration by a redox polymer-immobilized enzymatic system. ACS Catal 9:5486–5495. https://doi.org/10.1021/acscatal.9b00513
Zhang Y-W, Tiwari MK, Gao H et al (2012) Cloning and characterization of a thermostable H2O-forming NADH oxidase from Lactobacillus rhamnosus. Enzyme Microb Technol 50:255–262. https://doi.org/10.1016/j.enzmictec.2012.01.009
Acknowledgement
The authors appreciated the financial support from Natural Science Foundation of Guangxi province (2019GXNSFAA185059).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhou, Q., Gao, J. & Zhang, YW. Optimal pH shift of the NADH oxidase from Lactobacillus rhamnosus with a single mutation. Biotechnol Lett 43, 1413–1420 (2021). https://doi.org/10.1007/s10529-021-03129-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03129-7