Abstract
Objectives
To improve rhamnolipid production and its potential application in removal of crude oil, the recombinant Pseudomonas aeruginosa strain DAB was constructed to enhance yield of rhamnolipids.
Results
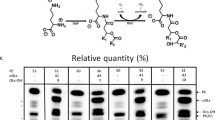
Strain DAB had a higher yield of 17.3 g rhamnolipids l−1 in the removal process with crude oil as the sole carbon source than 10 g rhamnolipids l−1 of wild-type strain DN1, where 1% crude oil was degraded more than 95% after 14 days cultivation. These rhamnolipids reduced the surface tension of water from 72.92 to 26.15 mN m−1 with CMC of 90 mg l−1. The predominant rhamnolipid congeners were Rha–C10–C10 and Rha–Rha–C10–C10 detected by MALDI-TOF MS analysis with approx. 70% relative abundance, although a total of 21 rhamnolipid congeners were accumulated.
Conclusion
Increasing the copy number of rhlAB genes efficiently enhanced the production of rhamnolipids by the recombinant P. aeruginosa DAB and thus presents a promising application for the bioremediation process.





Similar content being viewed by others
References
Abdelmawgoud AM, Lépine F, Déziel E (2010) Rhamnolipids: diversity of structures, microbial origins and roles. Appl Microbiol Biotechnol 86:1323–1336
Abidi N, Hequet E, Cabrales L (2010) Changes in sugar composition and cellulose content during the secondary cell wall biogenesis in cotton fibers. Cellulose 17:153–160
Amani H, Müller MM, Syldatk C, Hausmann R (2013) Production of microbial rhamnolipid by Pseudomonas Aeruginosa MM1011 for ex situ enhanced oil recovery. Appl Biochem Biotechnol 170:1080–1093
Bordoloi NK, Konwar BK (2008) Microbial surfactant-enhanced mineral oil recovery under laboratory conditions. Colloid Surf B Biointerfaces 63:73–82
Cayias JL, Schechter RS, Wade WH (1975) The measurement of low interfacial tension via the spinning drop technique. ACS Symp 8:234–247
Dobler L, Vilela LF, Almeida RV, Neves BC (2016) Rhamnolipids in perspective: gene regulatory pathways, metabolic engineering, production and technological forecasting. Nat Biotechnol 33:123–135
Dong W, He CQ, Li YP, Huang C, Chen FL, Ma YL (2017) Complete genome sequence of a versatile hydrocarbon degrader, Pseudomonas aeruginosa DN1 isolated from petroleum-contaminated soil. Gene Rep 7:123–126
Jansons I, Touchie G, Sharp R, Almquist K, Farinha MA, Lam JS, Kropinski AM (1994) Deletion and transposon mutagenesis and sequence analysis of the pRO1600 OriR region found in the broad-host-range plasmids of the pQF series. Plasmid 31:265–274
Kaczorek E, Olszanowski A (2011) Uptake of hydrocarbon by Pseudomonas fluorescens (P1) and Pseudomonas putida (K1) strains in the presence of surfactants: a cell surface modification. Water Air Soil Pollut 214:451–459
Ma KY, Sun MY, Dong W, He CQ, Chen FL, Ma YL (2016) Effects of nutrition optimization strategy on rhamnolipid production in a Pseudomonas aeruginosa strain DN1 for bioremediation of crude oil. Biocat Agric Biotechnol 6:144–151
Müller MM, Hörmann B, Kugel M, Syldatk C, Hausmann R (2011) Evaluation of rhamnolipid production capacity of Pseudomonas aeruginosa PAO1 in comparison to the rhamnolipid over-producer strains DSM 7108 and DSM 2874. Appl Microbiol Biotechnol 89:585–592
Nie M, Yin X, Ren C, Wang Y, Xu F, Shen Q (2010) Novel rhamnolipid biosurfactants produced by a polycyclic aromatic hydrocarbon-degrading bacterium Pseudomonas aeruginosa strain NY3. Biotechnol Adv 28:635–643
Nikolopoulou M, Pasadakis N, Norf H, Kalogerakis N (2013) Enhanced ex situ bioremediation of crude oil contaminated beach sand by supplementation with nutrients and rhamnolipids. Mar Pollut Bull 77:37–44
Nitschke M, Costa SG, Contiero J (2010) Structure and applications of a rhamnolipid surfactant produced in soybean oil waste. Appl Biochem Biotechnol 160:2066–2074
Sajna KV, Sukumaran RK, Gottumukkala LD, Pandey A (2015) Crude oil biodegradation aided by biosurfactants from Pseudozyma sp. NII 08165 or its culture broth. Bioresour Technol 191:133–139
Sarachat T, Pornsunthorntawee O, Chavadej S, Rujiravanit R (2010) Purification and concentration of a rhamnolipid biosurfactant produced by Pseudomonas aeruginosa SP4 using foam fractionation. Bioresour Technol 101:324–330
Zhang W, Li J, Huang G, Song W, Huang Y (2011) An experimental study on the bio-surfactant-assisted remediation of crude oil and salt contaminated soils. J Environ Sci Health A Tox Hazard Subst Environ Eng 46:306–313
Zhao F, Cui Q, Han S, Dong H, Zhang J, Ma F, Zhang Y (2015) Enhanced rhamnolipid production of Pseudomonas aeruginosa SG by increasing copy number of rhlAB genes with modified promoter. RS Adv 5:70546–70552
Acknowledgements
This research was supported by the National Science Foundation for Young Scientists of China (Grant No. 31000069) and the research project of Shaanxi Provincial Key Laboratory of Biotechnology (16JS108).
Supplementary information
Supplementary Figure 1—Schematic diagram of the construction of recombinant plasmid.
Supplementary Figure 2—Time course of pollutant degradation by the engineered strain DAB and DN1 after inoculation in the optimized medium consisting of BPLM with different PAHs as the sole carbon source (A) naphathalene; (B) phenanthrene; (C) pyrene; (D) fluoranthene.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, C., Dong, W., Li, J. et al. Characterization of rhamnolipid biosurfactants produced by recombinant Pseudomonas aeruginosa strain DAB with removal of crude oil. Biotechnol Lett 39, 1381–1388 (2017). https://doi.org/10.1007/s10529-017-2370-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-017-2370-x