Abstract
Lignocellulosic biomass is an attractive sustainable platform for fuel ethanol production. Xylose is a second after glucose most abounded sugar in lignocellulosic hydrolysates. Effective conversion of xylose to ethanol is one of key prerequisite for the development of an efficient conversion of biomass to ethanol. Engineered Saccharomyces cerevisiae strains are able to xylose fermentation. However, the yield and productivities of xylose fermentation remains lower in comparison with glucose fermentation. In this work, we studied impact of transcription factors Znf1, Sip4, Adr1, Tup1, and Hap4 on xylose catabolism. We have isolated znf1Δ, adr1Δ, tup1Δ and hap4Δ mutants, and strains overexpressing SIP4, ADR1 and HAP4 genes on the background of xylose-fermenting strain of S. cerevisiae aiming to explore involvement of these transcription factors in regulation of xylose growth and fermentation. It was shown that hap4Δ reveal 1.8-fold increase of ethanol production from xylose as compared to that of parental strain. The hap4Δ mutant accumulates 10.38 g l−1 of ethanol with an overall ethanol yield reaching 0.41 g g−1 of consumed xylose. While the other constructed strains revealed a decrease in ethanol production from this pentose.
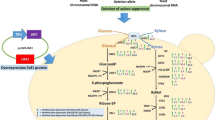
Graphical Abstract






Similar content being viewed by others
Data availability
All data generated during this study are included in this published article and its supplementary files.
References
Cunha JT, Costa CE, Ferraz L, Romaní A, Johansson B, Sá-Correia I, Domingues L (2018) HAA1 PRS3 overexpression boosts yeast tolerance towards acetic acid improving xylose or glucose consumption: unravelling the underlying mechanisms. Appl Microbiol Biotechnol 102:4589–4600. https://doi.org/10.1007/s00253-018-8955-z
Dikicioglu D, Pir P, Onsan ZI, Ulgen KO, Kirdar B, Oliver SG (2008) Integration of metabolic modeling phenotypic data in evaluation improvement of ethanol production using respiration-deficient mutants of Saccharomyces cerevisiae. Appl Environ Microbiol 74:5809–5816. https://doi.org/10.1128/AEM.00009-08
Dudley AM, Janse DM, Tanay A, Shamir R, Church GM (2005) A global view of pleiotropy phenotypically derived gene function in yeast. Mol Syst Biol 1(2005):0001. https://doi.org/10.1038/msb4100004
Dzanaeva L, Kruk B, Ruchala J, Nielsen J, Sibirny A, Dmytruk K (2020a) The role of peroxisomes in xylose alcoholic fermentation in the engineered Saccharomyces cerevisiae. Cell Biol Int 44:1606–1615. https://doi.org/10.1002/cbin.11353
Dzanaeva L, Ruchala J, Sibirny A, Dmytruk K (2020b) The impact of transcriptional factors Znf1 Sip4 on xylose alcoholic fermentation in recombinant strains of yeast Saccharomyces cerevisiae. Cytol Genet 54:386–392. https://doi.org/10.3103/S0095452720050035
Forsburg SL, Guarente L (1989) Identification characterization of HAP4: a third component of the CCAAT-bound HAP2/HAP3 heteromer. Genes Dev 3:1166–1178. https://doi.org/10.1101/gad.3.8.1166
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96. https://doi.org/10.1016/s0076-6879(02)50957-5
Harbison CT, Gordon DB, Lee TI, Rinaldi NG, Macisaac KD, Danford TD, Young AR (2004) Transcriptional regulatory code of a eukaryotic genome. Nature 431:99–104. https://doi.org/10.1038/nature02800
Jin YS, Laplaza JM, Jeffries TW (2004) Saccharomyces cerevisiae engineered for xylose metabolism exhibits a respiratory response. Appl Environ Microbiol 70:6816–6825. https://doi.org/10.1128/AEM.70.11.6816-6825.2004
Kwak S, Jin YS (2017) Production of fuels chemicals from xylose by engineered Saccharomyces cerevisiae: a review perspective. Microb Cell Fact 16:82. https://doi.org/10.1186/s12934-017-0694-9
Lee KS, Hong ME, Jung SC, Ha SJ, Yu BJ, Koo HM, Jin YS (2011) Improved galactose fermentation of Saccharomyces cerevisiae through inverse metabolic engineering. Biotechnol Bioeng 108:621–631. https://doi.org/10.1002/bit.22988
Lin X, Zhang CY, Bai XW, Song HY, Xiao DG (2014a) Effects of MIG1, TUP1 SSN6 deletion on maltose metabolism leavening ability of baker’s yeast in lean dough. Microb Cell Fact 13:93. https://doi.org/10.1186/s12934-014-0093-4
Lin Y, Chomvong K, Acosta-Sampson L, Estrela R, Galazka JM, Kim SR, Cate JH (2014b) Leveraging transcription factors to speed cellobiose fermentation by Saccharomyces cerevisiae. Biotechnol Biofuels 7:126. https://doi.org/10.1186/s13068-014-0126-6
Liu H, Liu K, Yan M, Xu L, Ouyang P (2011) gTME for improved adaptation of Saccharomyces cerevisiae to corn cob acid hydrolysate. Applied Biochem Biotechnol 164:1150–1159. https://doi.org/10.1007/s12010-011-9201-7
MacPherson S, Larochelle M, Turcotte B (2006) A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiol Mol Biol Rev 70:583–604. https://doi.org/10.1128/MMBR.00015-06
Malavé TM, Dent SY (2006) Transcriptional repression by Tup1-Ssn6. Biochem Cell Biol 84:437–443. https://doi.org/10.1139/o06-073
Matsushika A, Hoshino T (2015) Increased ethanol production by deletion of HAP4 in recombinant xylose-assimilating Saccharomyces cerevisiae. J Ind Microbiol Biotechnol 42:1623–1631. https://doi.org/10.1007/s10295-015-1695-5
Matsushika A, Goshima T, Hoshino T (2014) Transcription analysis of recombinant industrial laboratory Saccharomyces cerevisiae strains reveals the molecular basis for fermentation of glucose xylose. Microb Cell Fact 13:16. https://doi.org/10.1186/1475-2859-13-16
Miyagi H, Kawai S, Murata K (2009) Two sources of mitochondrial NADPH in the yeast Saccharomyces cerevisiae. J Biol Chem 284:7553–7560. https://doi.org/10.1074/jbc.M804100200
Moysés DN, Reis VCB, de Almeida JRM, de Moraes LMP, Torres FAG (2016) Xylose fermentation by Saccharomyces cerevisiae: challenges prospects. Int J Mol Sci 17:207. https://doi.org/10.3390/ijms17030207
Nijland JG, Shin HY, Boender LGM, de Waal PP, Klaassen P, Driessen AJM (2017) Improved xylose metabolism by a CYC8 mutant of Saccharomyces cerevisiae. Appl Environ Microbiol 83:e00095-e117. https://doi.org/10.1128/AEM.00095-17
Peng B, Shen Y, Li X, Chen X, Hou J, Bao X (2012) Improvement of xylose fermentation in respiratory-deficient xylose-fermenting Saccharomyces cerevisiae. Metab Eng 14:9–18. https://doi.org/10.1016/j.ymben.2011.12.001
Qi K, Zhong JJ, Xia XX (2014) Triggering respirofermentative metabolism in the crabtree-negative yeast Pichia guilliermondii by disrupting the CAT8 gene. Appl Environ Microbiol 80:3879–3887. https://doi.org/10.1128/AEM.00854-14
Raghevendran V, Patil KR, Olsson L, Nielsen J (2006) Hap4 is not essential for activation of respiration at low specific growth rates in Saccharomyces cerevisiae. J Biol Chem 281:12308–12314. https://doi.org/10.1074/jbc.M512972200
Ruchala J, Kurylenko OO, Soontorngun N, Dmytruk KV, Sibirny AA (2017) Transcriptional activator Cat8 is involved in regulation of xylose alcoholic fermentation in the thermotolerant yeast Ogataea (Hansenula) polymorpha. Microb Cell Fact 16:36. https://doi.org/10.1186/s12934-017-0652-6
Sakihama Y, Hasunuma T, Kondo A (2015) Improved ethanol production from xylose in the presence of acetic acid by the overexpression of the HAA1 gene in Saccharomyces cerevisiae. J Biosci Bioeng 119:297–302. https://doi.org/10.1016/j.jbiosc.2014.09.004
Sambrook J, Fritsh EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor. Cold Spring Harbor Laboratory Press, NY
Scalcinati G, Otero JM, Van Vleet JR, Jeffries TW, Olsson L, Nielsen J (2012) Evolutionary engineering of Saccharomyces cerevisiae for efficient aerobic xylose consumption. FEMS Yeast Res 12:582–597. https://doi.org/10.1111/j.1567-1364.2012.00808.x
Shi NQ, Davis B, Sherman F, Cruz J, Jeffries TW (1999) Disruption of the cytochrome c gene in xylose-utilizing yeast Pichia stipitis leads to higher ethanol production. FEMS Yeast Res 15(11):1021–1030. https://doi.org/10.1002/(SICI)1097-0061(199908)15:11%3c1021::AID-YEA429%3e3.0.CO;2-V
Songdech P, Ruchala J, Semkiv MV, Jensen LT, Sibirny A, Ratanakhanokchai K, Soontorngun N (2020) Overexpression of transcription factor ZNF1 of glycolysis improves bioethanol productivity under high glucose concentration enhances acetic acid tolerance of Saccharomyces cerevisiae. Biotechnology 15(7):e1900492. https://doi.org/10.1002/biot.201900492
Soontorngun N (2017) Reprogramming of nonfermentative metabolism by stress-responsive transcription factors in the yeast Saccharomyces cerevisiae. Curr Genet 63:1–7. https://doi.org/10.1007/s00294-016-0609-z
Souto-Maior AM, Runquist D, Hahn-Hägerdal B (2009) Crabtree-negative characteristics of recombinant xylose-utilizing Saccharomyces cerevisiae. Biotechnology 143:119–123. https://doi.org/10.1016/j.jbiotec.2009.06.022
Sunwoo IY, Sukwong P, Park YR, Jeong DY, Kim SR, Jeong GT, Kim SK (2020) Enhancement of galactose uptake from Kappaphycus alvarezii using Saccharomyces cerevisiae through deletion of negative regulators of GAL genes. Appl Biochem Biotechnol. https://doi.org/10.1007/s12010-020-03434-3
Tangsombatvichit P, Semkiv MV, Sibirny AA, Jensen LT, Ratanakhanokchai K, Soontorngun N (2015) Zinc cluster protein Znf1, a novel transcription factor of non-fermentative metabolism in Saccharomyces cerevisiae. FEMS Yeast Res 15:002. https://doi.org/10.1093/femsyr/fou002
van Maris AJ, Bakker BM, Brandt M, Boorsma A, de Mattos MJT, Grivell LA, Blom J (2001) Modulating the distribution of fluxes among respiration fermentation by overexpression of HAP4 in Saccharomyces cerevisiae. FEMS Yeast Res 1:139–149. https://doi.org/10.1111/j.1567-1364.2001.tb00025.x
Wang M, Wu M, Huo H (2007) Life-cycle energy greenhouse gas emission impacts of different corn ethanol plant types. Environ Res Lett 2:24001–24013. https://doi.org/10.1088/1748-9326/2/2/024001
Watanabe D, Hashimoto N, Mizuno M, Zhou Y, Akao T, Shimoi H (2013) Accelerated alcoholic fermentation caused by defective gene expression related to glucose derepression in Saccharomyces cerevisiae. Biosci Biotechnol Biochem 77:2255–2262. https://doi.org/10.1271/bbb.130519
Wenning L, Yu T, David F, Nielsen J, Siewers V (2017) Establishing very long-chain fatty alcohol wax ester biosynthesis in Saccharomyces cerevisiae. Biotechnol Bioeng 114:1025–1035. https://doi.org/10.1002/bit.26220
Acknowledgements
This study was supported by Polish National Science Center, grant Opus UMO-2016/21/B/NZ1/00280, DEC-2020/37/B/NZ1/02232 and by National Academy of Sciences of Ukraine (Grant 2-21 and 17-21).
Funding
This study was supported by Polish National Science Center, grant Opus UMO-2016/21/B/NZ1/00280, DEC-2020/37/B/NZ1/02232 and by National Academy of Sciences of Ukraine (Grant 2–21 and 17–21).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. LD, BK and JR performed the experiments. LD, BK, JR, AS and KD analysed the data. LD, AS and KD drafted the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare not to have any conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dzanaeva, L., Kruk, B., Ruchala, J. et al. The impact of transcription factors Znf1, Sip4, Adr1, Tup1, and Hap4 on xylose alcoholic fermentation in the engineered yeast Saccharomyces cerevisiae. Antonie van Leeuwenhoek 114, 1373–1385 (2021). https://doi.org/10.1007/s10482-021-01607-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01607-6