Abstract
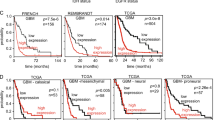
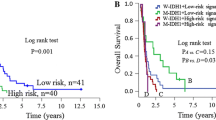
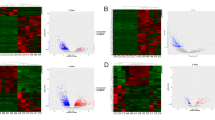
To investigate the m6a-related long non-coding RNAs (lncRNAs) that may be exploited as potential biomarkers in primary glioblastoma (pGBM), a cohort of 268 glioma samples from GSE16011 dataset was included for discovery. The Chinese Glioma Genome Atlas (CGGA) microarray and RNA sequencing databases were used for validation. Bioinformatic analyses were performed using the R software. The m6a-lncRNA co-expression networks were constructed, and four m6a-related lncRNAs (MIR9-3HG, LINC00900, MIR155HG, and LINC00515) were identified in pGBM patients on the univariate Cox regression analysis. Patients in the low-risk group had longer overall survival (OS) and progression-free survival (PFS) than those in the high-risk group (P = 0.0025, P = 0.0070). Moreover, the high-risk group displayed older age, isocitrate dehydrogenase (IDH) wild-type, classical and mesenchymal TCGA subtype, and G3 CGGA subtype. Distinct m6a status was identified according to histologic grade and two groups (low-risk and high-risk). Functional annotation showed that differentially expressed genes between the two groups were enriched in immune response, apoptosis, cell adhesion, negative regulation of transcription, negative regulation of RNA metabolic process, and regulation of RNA metabolic process. We profiled the m6a status in glioma and identified four m6a-related prognostic lncRNAs for pGBMs.





Similar content being viewed by others
References
Yang P, Wang Y, Peng X, You G, Zhang W, Yan W, Bao Z, Wang Y, Qiu X, Jiang T (2013) Management and survival rates in patients with glioma in China (2004-2010): a retrospective study from a single-institution. J Neuro-Oncol 113:259–266
Yang Y, Hsu PJ, Chen YS, Yang YG (2018) Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res 28:616–624
Weng H, Huang H, Wu H et al (2018) METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell 22:191–205 e199
Bansal H, Yihua Q, Iyer SP, Ganapathy S, Proia DA, Penalva LO, Uren PJ, Suresh U, Carew JS, Karnad AB, Weitman S, Tomlinson GE, Rao MK, Kornblau SM, Bansal S (2014) WTAP is a novel oncogenic protein in acute myeloid leukemia. Leukemia 28:1171–1174
Cui Q, Shi H, Ye P et al (2017) m(6)A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep 18:2622–2634
Liu J, Ren D, Du Z, Wang H, Zhang H, Jin Y (2018) m(6)A demethylase FTO facilitates tumor progression in lung squamous cell carcinoma by regulating MZF1 expression. Biochem Biophys Res Commun 502:456–464
Zhang S, Zhao BS, Zhou A et al (2017) m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31:591–606 e596
Patil DP, Chen CK, Pickering BF et al (2016) m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537:369–373
Zhao X, Chen Y, Mao Q, Jiang X, Jiang W, Chen J, Xu W, Zhong L, Sun X (2018) Overexpression of YTHDF1 is associated with poor prognosis in patients with hepatocellular carcinoma. Cancer Biomark 21:859–868
Bian EB, Li J, Xie YS, Zong G, Li J, Zhao B (2015) lncRNAs: new players in gliomas, with special emphasis on the interaction of lncRNAs with EZH2. J Cell Physiol 230:496–503
Sun Y, Wang Z, Zhou D (2013) Long non-coding RNAs as potential biomarkers and therapeutic targets for gliomas. Med Hypotheses 81:319–321
Zhang XQ, Leung GK (2014) Long non-coding RNAs in glioma: functional roles and clinical perspectives. Neurochem Int 77:78–85
Wang P, Ren Z, Sun P (2012) Overexpression of the long non-coding RNA MEG3 impairs in vitro glioma cell proliferation. J Cell Biochem 113:1868–1874
Wang Q, Zhang J, Liu Y, Zhang W, Zhou J, Duan R, Pu P, Kang C, Han L (2016) A novel cell cycle-associated lncRNA, HOXA11-AS, is transcribed from the 5-prime end of the HOXA transcript and is a biomarker of progression in glioma. Cancer Lett 373:251–259
Han Y, Wu Z, Wu T, Huang Y, Cheng Z, Li X, Sun T, Xie X, Zhou Y, du Z (2016) Tumor-suppressive function of long noncoding RNA MALAT1 in glioma cells by downregulation of MMP2 and inactivation of ERK/MAPK signaling. Cell Death Dis 7:e2123
Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, Liu J, Che L, Li J (2019) Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)A reader YTHDF3. Mol Cancer 18:143
Wu Y, Yang X, Chen Z et al (2019) m(6)A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol Cancer 18:87
He Y, Hu H, Wang Y, Yuan H, Lu Z, Wu P, Liu D, Tian L, Yin J, Jiang K, Miao Y (2018) ALKBH5 inhibits pancreatic cancer motility by decreasing long non-coding RNA KCNK15-AS1 methylation. Cell Physiol Biochem 48:838–846
Zhang X, Sun S, Pu JK, Tsang AC, Lee D, Man VO, Lui WM, Wong ST, Leung GK (2012) Long non-coding RNA expression profiles predict clinical phenotypes in glioma. Neurobiol Dis 48:1–8
Wang Y, Li Y, Yue M et al (2018) Publisher correction: N(6)-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nat Neurosci
Michelhaugh SK, Lipovich L, Blythe J, Jia H, Kapatos G, Bannon MJ (2011) Mining Affymetrix microarray data for long non-coding RNAs: altered expression in the nucleus accumbens of heroin abusers. J Neurochem 116:459–466
Johnson R (2012) Long non-coding RNAs in Huntington’s disease neurodegeneration. Neurobiol Dis 46:245–254
Cai J, Zhang W, Yang P, Wang Y, Li M, Zhang C, Wang Z, Hu H, Liu Y, Li Q, Wen J, Sun B, Wang X, Jiang T, Jiang C (2015) Identification of a 6-cytokine prognostic signature in patients with primary glioblastoma harboring M2 microglia/macrophage phenotype relevance. PLoS One 10:e0126022
Bao ZS, Li MY, Wang JY, Zhang CB, Wang HJ, Yan W, Liu YW, Zhang W, Chen L, Jiang T (2014) Prognostic value of a nine-gene signature in glioma patients based on mRNA expression profiling. CNS Neurosci Ther 20:112–118
Zhang CB, Zhu P, Yang P, Cai JQ, Wang ZL, Li QB, Bao ZS, Zhang W, Jiang T (2015) Identification of high risk anaplastic gliomas by a diagnostic and prognostic signature derived from mRNA expression profiling. Oncotarget 6:36643–36651
Lossos IS, Czerwinski DK, Alizadeh AA, Wechser MA, Tibshirani R, Botstein D, Levy R (2004) Prediction of survival in diffuse large-B-cell lymphoma based on the expression of six genes. N Engl J Med 350:1828–1837
Zhang W, Zhang J, Yan W, You G, Bao Z, Li S, Kang C, Jiang C, You Y, Zhang Y, Chen CC, Song SW, Jiang T (2013) Whole-genome microRNA expression profiling identifies a 5-microRNA signature as a prognostic biomarker in Chinese patients with primary glioblastoma multiforme. Cancer 119:814–824
Wang W, Yang F, Zhang L, Chen J, Zhao Z, Wang H, Wu F, Liang T, Yan X, Li J, Lan Q, Wang J, Zhao J (2016) LncRNA profile study reveals four-lncRNA signature associated with the prognosis of patients with anaplastic gliomas. Oncotarget 7:77225–77236
Wang W, Zhang L, Liu W, Zhu Q, Lan Q, Zhao J (2016) Antiplatelet agents for the secondary prevention of ischemic stroke or transient ischemic attack: a network meta-analysis. J Stroke Cerebrovasc Dis 25:1081–1089
Dai D, Wang H, Zhu L, Jin H, Wang X (2018) N6-methyladenosine links RNA metabolism to cancer progression. Cell Death Dis 9:124
Nikkanen J, Landoni JC, Balboa D, Haugas M, Partanen J, Paetau A, Isohanni P, Brilhante V, Suomalainen A (2018) A complex genomic locus drives mtDNA replicase POLG expression to its disease-related nervous system regions. EMBO Mol Med 10:13–21
Jia L, Wu D, Wang Y, You W, Wang Z, Xiao L, Cai G, Xu Z, Zou C, Wang F, Teoh JY, Ng CF, Yu S, Chan FL (2018) Orphan nuclear receptor TLX contributes to androgen insensitivity in castration-resistant prostate cancer via its repression of androgen receptor transcription. Oncogene 37:3340–3355
Thiele JA, Hosek P, Kralovcova E et al (2018) lncRNAs in non-malignant tissue have prognostic value in colorectal cancer. Int J Mol Sci 19
Song J, Peng J, Zhu C, Bai G, Liu Y, Zhu J, Liu J (2018) Identification and validation of two novel prognostic lncRNAs in kidney renal clear cell carcinoma. Cell Physiol Biochem 48:2549–2562
Wu X, Wang Y, Yu T, Nie E, Hu Q, Wu W, Zhi T, Jiang K, Wang X, Lu X, Li H, Liu N, Zhang J, You Y (2017) Blocking MIR155HG/miR-155 axis inhibits mesenchymal transition in glioma. Neuro-Oncology 19:1195–1205
Balasubramaniyan V, Bhat KP (2017) Targeting MIR155HG in glioma: a novel approach. Neuro-Oncology 19:1152–1153
Lu D, Yang C, Zhang Z, Cong Y, Xiao M (2018) Knockdown of Linc00515 inhibits multiple myeloma autophagy and chemoresistance by upregulating miR-140-5p and downregulating ATG14. Cell Physiol Biochem 48:2517–2527
Zheng ZG, Xu H, Suo SS et al (2016) The essential role of H19 contributing to cisplatin resistance by regulating glutathione metabolism in high-grade serous ovarian cancer. Sci Rep 6:26093
Funding
This work was supported by grants from the Beijing Municipal Administration of Hospitals’ Mission Plan (SML20150501), the “13th Five-Year Plan” National Science and Technology supporting plan (2015BAI09B04), and the Foundation of Beijing Tiantan Hospital (2018-YQN-6).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was approved by the Beijing Tiantan Hospital institutional review board (IRB).
Informed consent
Informed consent was obtained from each patient involved in our research.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Wang, W., Li, J., Lin, F. et al. Identification of N6-methyladenosine-related lncRNAs for patients with primary glioblastoma. Neurosurg Rev 44, 463–470 (2021). https://doi.org/10.1007/s10143-020-01238-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10143-020-01238-x