Abstract
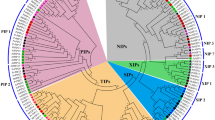
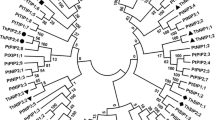
Tonoplast intrinsic proteins (TIPs) play a vital role in water transport across membranes. In the present study, we performed a comparative analysis of TIP genes in ten plant species including both monocots and dicots. A total of 100 TIP aquaporin genes were identified, and their relationships among the plant species were analyzed. Phylogenetic analysis was performed to evaluate the relationship of these genes within the plant species. Based on the phylogenetic analysis results, TIPs were classified into five distinct arbitrary groups (group I to group V), which represented TIP2, TIP5, TIP4, TIP1, and TIP3, respectively. Group I represented the largest arbitrary group, followed by group IV, in the phylogenetic tree. The result clearly indicates that TIP2 and TIP1 are abundant aquaporins and highly related among the species. In the present review, a comparative study of gene structure analysis between dicots and monocots has been performed to analyze their structural variation. Most of the predicted motifs are conserved among the species, signifying an evolutionary relationship. The gene expression analysis indicated that the expression of TIP genes varies during different developmental stages and also during stressed conditions. The results indicated a great degree of evolutionary relationship and variation in the expression levels of TIPs in plants.








Similar content being viewed by others
Abbreviations
- APQ:
-
Aquaporin
- TIPs:
-
Tonoplast intrinsic proteins
- CDS:
-
Coding DNA sequence
- NPA:
-
Asparagine-Proline-Alanine
References
Alexandersson E, Fraysse L, Sjovall-Larsen S, Gustavsson S, Fellert M, Karlsson M, Johanson U, Kjellbom P (2005) Whole gene family expression and drought stress regulation of aquaporins. Plant Mol Biol 59:469–484
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proceedings of the second international conference on 132 intelligent systems for molecular biology. AAAI Press, Menlo Park, pp 28–36
Beebo A et al (2009) Life with and without AtTIP1;1, an Arabidopsis aquaporin preferentially localized in the apposing tonoplasts of adjacent vacuoles. Plant Mol Biol 70:193–209. doi:10.1007/s11103-009-9465-2
Boursiac Y, Chen S, Luu DT, Sorieul M, van den Dries N, Maurel C (2005) Early effects of salinity on water transport in Arabidopsis roots. Molecular and cellular features of aquaporin expression. Plant Physiol 139:790–805
Chaumont F, Barrieu F, Wojcik E, Chrispeels MJ, Jung R (2001) Aquaporins constitute a large and highly divergent protein family in maize. Plant Physiol 125:1206–1215
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676
Danielson JAH, Johanson U (2008) Unexpected complexity of the aquaporin gene family in the moss Physcomitrella patens. BMC Plant Biol 8:45
Danielson JAH, Johanson U (2010) Phylogeny of major intrinsic proteins. Adv Exp Med Biol 679:19–31
Forrest KL, Bhave M (2008) The PIP and TIP aquaporins in wheat form a large and diverse family with unique gene structures and functionally important features. Funct Integr Genom 8:115–133
Gattolin S, Sorieul M, Hunter PR, Khonsari RH, Frigerio L (2009) In vivo imaging of the tonoplast intrinsic protein family in Arabidopsis roots. BMC Plant Biol 9(1):133. doi:10.1186/1471-2229-9-133
Gattolin S, Sorieul M, Frigerio L (2011) Mapping of tonoplast intrinsic proteins in maturing and germinating Arabidopsis seeds reveals dual localization of embryonic TIPs to the tonoplast and plasma membrane. Mol Plant 4(1):180–189. doi:10.1093/mp/ssq051
Gomes D, Agasse A, Thiébaud P, Delrot S, Delrot S, Chaumont F (2009) Aquaporins are multifunctional water and solute transporters highly divergent in living organisms. Biochim Biophys Acta 1788:1213–1228
Goodstein DM et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186. doi:10.1093/nar/gkr944
Gustavsson S, Lebrun AS, Norden K, Chaumont F, Johanson U (2005) A novel plant major intrinsic protein in Physcomitrella patens most similar to bacterial glycerol channels. Plant Physiol 139:287–295
Higuchi T, Suga S, Tsuchiya T, Hisada H, Morishima S, Okada Y, Maeshima M (1998) Molecular cloning, water channel activity and tissue specific expression of two isoforms of radish vacuolar aquaporin. Plant Cell Physiol 39:905–913
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinforma 2008:420747. doi:10.1155/2008/420747
Jami SK, Clark GB, Ayele BT, Ashe P, Kirti PB (2012) Genome-wide comparative analysis of annexin superfamily in plants. PLoS ONE 7:e47801
Johanson U, Karlsson M, Johansson I, Gustavsson S, Sjovall S, Fraysse L, Weig AR, Kjellbom P (2001) The complete set of genes encoding major intrinsic proteins in Arabidopsis provides a framework for a new nomenclature for major intrinsic proteins in plants. Plant Physiol 126:1358–1369
Johnson KD, Hofte H, Chrispeels MJ (1990) An intrinsic tonoplast protein of protein storage vacuoles in seeds is structurally related to a bacterial solute transporter (GIpF). Plant Cell 2:525–532
Kaldenhoff R, Fischer M (2006) Functional aquaporin diversity in plants. Biochim Biophys Acta 1758(8):1134–1141. doi:10.1016/j.bbamem.2006.03.012
Kruse E, Uehlein N, Kaldenhoff R (2006) The aquaporins. Genome Biol 7(2):206. doi:10.1186/gb-2006-7-2-206
Lamesch P et al (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:D1202–D1210. doi:10.1093/nar/gkr1090
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Ludevid D, Hofte H, Himelblau E, Chrispeels MJ (1992) The expression pattern of the tonoplast intrinsic protein gamma-TIP in Arabidopsis thaliana is correlated with cell enlargement. Plant Physiol 100:1633–1639
Maeshima M (2001) Tonoplast transporters: organization and function. Annu Rev Plant Physiol Plant Mol Biol 52:469–497. doi:10.1146/annurev.arplant.52.1.469
Maurel C, Tacnet F, Guclu J, Guern J, Ripoche P (1997) Purified vesicles of tobacco cell vacuolar and plasma membranes exhibit dramatically different water permeability and water channel activity. Proc Natl Acad Sci U S A 94:7103–7108
Maurel C, Verdoucq L, Luu DT, Santoni V (2008) Plant aquaporins: membrane channels with multiple integrated functions. Annu Rev Plant Biol 59:595–624
Pao GM, Wu LF, Johnson KD, Hofte H, Chrispeels MJ, Sweet G, Sandal NN, Saier MH Jr (1991) Evolution of the MIP family of integral membrane transport proteins. Mol Microbiol 5:33–37
Proost S, Van Bel M, Sterck L, Billiau K, Van Parys T, Van de Peer Y, Vandepoele K (2009) PLAZA: a comparative genomics resource to study gene and genome evolution in plants. Plant Cell 21:3718–3731
Quigley F, Rosenberg JM, Shachar-Hill Y, Bohnert HJ (2002) From genome to function: the Arabidopsis aquaporins. Genome Biol 3(1):RESEARCH0001
Rambaut A (2012) FigTree v1.4. Institute of Evolutionary Biology, University of Edinburgh, Edinburgh
Roy SW, Penny D (2007) Patterns of intron loss and gains in plants: intron loss-dominated evolution and genome-wide comparison of O. sativa and A. thaliana. Mol Biol Evol 24(1):171–181
Schultz J et al (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci U S A 95:5857–5864
Silvestro D, Michalak I (2011) raxmlGUI: a graphical front-end for RAxML. Org Divers Evol 12(4):335–337. doi:10.1007/s13127-011-0056-0
Ye CY, Yang X, Xia X, Yin W (2013) Comparative analysis of cation/proton antiporter superfamily in plants. Gene 521:245–251
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1
(DOCX 29 kb)
Supplementary material 2
(GIF 717 kb)
Supplementary material 3
(GIF 93 kb)
Supplementary material 4
(GIF 579 kb)
Supplementary material 5
(GIF 193 kb)
Supplementary material 6
(GIF 137 kb)
Rights and permissions
About this article
Cite this article
Regon, P., Panda, P., Kshetrimayum, E. et al. Genome-wide comparative analysis of tonoplast intrinsic protein (TIP) genes in plants. Funct Integr Genomics 14, 617–629 (2014). https://doi.org/10.1007/s10142-014-0389-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-014-0389-9