Abstract
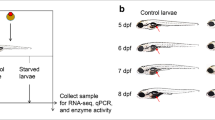
We investigated a time-course larval transcriptional analysis (RNA-seq) in the longfin yellowtail Seriola rivoliana, from hatching to day four at 22 °C, without providing zooplankton as food. Larval starvation is a critical physiological stage that must be prevented to ensure survival. However, the transcriptional mechanisms to endure starvation have not been investigated in marine fish. Differential gene expression showed newly day-specific transcriptome events during larval development. On day 1 (yolk sac absorption), the predominant upregulated developmental processes were larval growth, muscle and vision development, cytoskeletal structure, protein synthesis, protein and fat digestion-absorption, and hormone biosynthesis, whereas the cell cycle was suppressed. On day 2 (yolk sac exhaustion), a new stage of energy regeneration (ATP) was supplied by the oil drop reserve, whereas protein digestion-absorption and growth were suppressed. On day 3 (mouth opening and starvation), stress signals and nutrition deprivation upregulated the p53 signal and triggered autophagy and the AMP-activated protein kinase (AMPK) pathways as an alternative catabolic pathway to enduring starvation, and the circadian rhythm was established. On day 4 (starving and weakened larvae condition), autophagy supported subsequent protein synthesis, activated the immune system, and promoted estrogen signaling and skeleton renovation. However, larvae suppressed muscle development, vision and carbohydrate, and fat digestion-absorption and became lethargic, evidencing limited physiological support by autophagy to maintain survival without exogenous nutrition in this species.






Similar content being viewed by others
Availability of Data and Material
Data are available from the corresponding author, upon a reasonable request.
References
Aggarwal BB (2003) Signaling pathways of the TNF superfamily: a double-edged sword. Nat Rev Immunol 3:745–756
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Cell-cell adhesion. In: Mol Biol Cell, 4th edn. New York, Garland Science
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Aranda PS, LaJoie DM, Jorcyk CL (2012) Bleach gel: a simple agarose gel for analyzing RNA quality. Electrophoresis 33:366–369
Bell G, Sargent JR (2003) Arachidonic acid in aquaculture feeds: current status and future opportunities. Aquaculture 218:491–499
Blackinton JG, Keene JD (2014) Post-transcriptional RNA regulons affecting cell cycle and proliferation. Semin Cell Dev Biol 34:44–54
Blaxter JHS, Ehrlich KF (1974) Changes in behavior during starvation of herring and plaice larvae. In: Blaxter JHS (ed). The early life history of fish. Berlin Heidelberg, New York Springer Verlag.
Blaxter JHS, Hempel G (1963) The influence of egg size on herring larvae (Clupea harengus L.). ICES J Mar Sci 28:211–240
Bonora M, Patergnani S, Rimessi A, De Marchi E, Suski JM, Bononi A, Giorgi C, Marchi S, Missiroli S, Poletti F, Wieckowski MR, Pinton P (2012) ATP Synthesis and Storage Purinergic Signal 8:343–357
Bush A (1996) Transition from endogenous to exogenous nutrition: larval size parameters determining the start of external feeding and size of prey ingested by Ruegen spring herring Clupea harengus. MEPS 130:39–46
Cecconi F, Levine B (2008) The role of autophagy in mammalian development: cell makeover rather than cell death. Dev Cell 15:344–357
Chen BN, Qin JG, Kumar MS, Hutchinson W, Clarke S (2006) Ontogenetic development of the digestive system in yellowtail kingfish Seriola lalandi larvae. Aquaculture 256:489–501
China V, Holzman R (2014) Hydrodynamic starvation in first-feeding larval fishes. PNAS 111:8083
Cuenda A, Rousseau S (2007) P38 MAP-Kinases pathway regulation, function and role in human diseases. Biochim Biophys Acta Mol Cell Res 1773:1358–1375
Crowley SD, Rudemiller NP (2017) Immunologic effects of the renin-angiotensin system. JASN 28:1350–1361
Davidson NM, Oshlack A (2014) Corset: enabling differential gene expression analysis for de novoassembled transcriptomes. Genome Biol 15:410
de Mello NP, Orellana AM, Mazucanti CH, de Morais LG, Scavone C, Kawamoto EM (2019) Insulin and autophagy in neurodegeneration. Front Neurosci 13:491
Doi M, Ohno A, Kohno H, Taki Y, Singhagraiwan T (1997) Development of feeding ability in red snapper Lutjanus argentimaculatus early larvae. Fish Sci 63:845–853
Dou SZ, Masuda R, Tanaka M, Tsukamoto K (2002) Feeding resumption, morphological changes and mortality during starvation in Japanese flounder larvae. J Fish Biol 60:1363–1380
Ebrey T, Koutalos Y (2001) Vertebrate photoreceptors. Prog Retin Eye Res 20:49–94
Faccioli CK, Chedid RA, Mori RH, Amaral AC, Franceschini-Vicentini IB, Vicentini CA (2016) Acid and alkaline phosphatase localization in the digestive tract mucosa of the Hemisorubim platyrhynchos. Acta Histochem 118:722–728
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24:24–41
Fountain JH, Lappin SL (2020) Physiology, renin angiotensin system. In: StatPearls [Internet]. Treasure Island (FL). https://www.ncbi.nlm.nih.gov/books/NBK470410/
Fuentes N, Silveyra P (2019) Estrogen receptor signaling mechanisms. Adv Protein Chem Struct Biol 116:135–170
Gao J, Xu G, Xu P (2020) Comparative transcriptome analysis reveals metabolism transformation in Coilia nasus larvae during the mouth-open period. Comp Biochem Physiol D Genomics Proteomics 36
Gisbert E, Conklin DB, Piedrahita RH (2004) Effects of delayed first feeding on the nutritional condition and mortality of California halibut larvae. J Fish Biol 64:116–132
Gistelinck C, Gioia R, Gagliardi A, Tonelli F, Marchese L, Bianchi L, Landi C, Bini L, Huysseune A, Witten PE, Staes A, Gevaert K, De Rocker N, Menten B, Malfait F, Leikin S, Carra S, Tenni R, Rossi A, De Paepe A, Coucke P, Willaert A, Forlino A (2016) Zebrafish collagen type I: molecular and biochemical characterization of the major structural protein in bone and skin. Sci Rep 15:21540
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Harris S, Levine A (2005) The p53 pathway: positive and negative feedback loops. Oncogene 24:2899–2908
Hansen KD, Brenner SE, Dudoit S (2010) Biases in illumina transcriptome sequencing caused by random hexamer priming. Nucl Acids Res 38
He C, Klionsky DJ (2010) Analyzing autophagy in zebrafish. Autophagy 6:642–645
Hernandez RE, Galitan L, Cameron J, Goodwin N, Ramakrishnan L (2018) Delay of initial feeding of zebrafish larvae until 8 days postfertilization has no impact on survival or growth through the juvenile stage. Zebrafish 15:515–518
Hjort J (1914) Fluctuations in the great fisheries of northern Europe. Rapp P-V Reun Cons int Explor Mer XX: 237 pp.
Jeong CB, Kim BM, Kang HM, Ik-Y C, Rhee J-S, Lee JS (2015) Marine medaka ATP-binding cassette (ABC) superfamily and new insight into teleost Abch nomenclature. Sci Rep 5:15409
Jiang GM, Tan Y, Wang H, Peng L, Chen H-T, Meng X-J, Li L-L, Liu Y, Li W-F, Shan H (2019) The relationship between autophagy and the immune system and its applications for tumor immunotherapy. Mol Cancer 18:17
Kamler E (1992a) Endogenous feeding period. In: Early life History of fish. Springer.
Kamler E (1992b) Mixed feeding period. In: Early life History of fish. Springer.
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucl Acids Res 36:D480–D484
Kim JH, Kim N (2016) Signaling pathways in osteoclast differentiation. Chonnam Med J 52:12–17
Kim YK, Shin JS, Nahm MH (2016) NOD-like receptors in infection, immunity, and diseases. Yonsei Med J 57:5–14
Kotsias F, Cebrian I, Alloatti A (2019) Antigen processing and presentation. Int Rev Cell Mol Biol 348:69–121
Lewis L, Kwong RWM (2018) Zebrafish as a model system for investigating the compensatory regulation of ionic balance during metabolic acidosis. Int J Mol Sci 19:1087
Li B, Dewey C (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. https://doi.org/10.1186/1471-2105-12-323
Mao X, Cai T, Olyarchuk JG, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793
Mawed SA, Zhang J, Ren F, He Y, Mei J (2021) atg7 and beclin1 are essential for energy metabolism and survival during the larval-to-juvenile transition stage of zebrafish. Aquaculture and Fisheries. https://doi.org/10.1016/j.aaf.2021.01.002
Mazurais D, Darias M, Zambonino-Infante J, Cahu CL (2011) Transcriptomics for understanding marine fish larval development. Can J Zool 89:599–611
Mizushima N, Komatsu M (2011) Autophagy: renovation of cells and tissues. Cell 147:728–741
Montaseri A, Giampietri C, Rossi M, Riccioli A, Del Fattore A, Filippini A (2020) The role of autophagy in osteoclast differentiation and bone resorption function. Biomolecules 10:1398
National Center for Biotechnology Information (2021) Pubchem pathway summary for pathway SMP0000006, tyrosine metabolism, source: PathBank. https://pubchem.ncbi.nlm.nih.gov/pathway/PathBank:SMP0000006
Oikonomopoulou K, Ricklin D, Ward PA, Lambris JD (2012) Interactions between coagulation and complement their role in inflammation. Semin Immunopathol 34:151–165
Osse JWM (1990) Form changes in fish larvae in relation to changing demands of function. Neth J Zool 40:362–385
Osse JWM, Van den Boogaart JGM (1999) Dynamic morphology of fish larvae, structural implications of friction forces in swimming, feeding and ventilation. J Fish Biol 55:156–174
Osse JWM, Van den Boogaart JGM, Van Snik GMJ, Van der Sluys L (1997) Priorities during early growth of fish larvae. Aquaculture 155:249–258
Pacheco-Carlón N, Guerrero-Tortolero DA, Cervantes-Montoya LB, Racotta IS, Campos-Ramos R (2021) The effects of constant and oscillating temperature on embryonic development and early larval morphology in longfin yellowtail (Seriola rivoliana Valenciennes). Aquac Res 52:77–93
Patel A, Dettleff P, Hernandez E, Martinez V (2016) A comprehensive transcriptome of early development in yellowtail kingfish (Seriola lalandi). Mol Ecol Resour 16:364–376
Patin F, Corcia P, Vourc’h P, Nadal-Desbarats L, Baranek T, Goossens JF, Marouillat S, Dessein A-F, Descat A, Hounoum BM, Bruno C, Leman S, Andres CR, Blasco H, (2017) Omics to explore amyotrophic lateral sclerosis evolution: the central role of arginine and proline metabolism. Mol Neurobiol 54:5361–5374
Paul AJ (1983) Light, temperature, nauplii concentrations, and prey capture by first feeding pollock larvae Theragra chalcogramma. MEPS 13:175–79
Reichrath J, Reichrath S (2020) Notch signaling and embryonic development: an ancient friend, revisited. Adv Exp Med Biol 1218:9–37
Roo J, Fernández-Palacios H, Hernández-Cruz CM, Mesa-Rodriguez A, Schuchardt D, Izquierdo M (2012) First results of spawning and larval rearing of longfin yellowtail Seriola rivoliana as a fast-growing candidate for European marine finfish aquaculture diversification. Aquac Res 45:689–700
Rønnestad I, Tonheim SK, Fyhn HJ, Rojas-Garcia CR, Kamisaka Y, Koven W, Finn RN, Terjesen BF, Barr Y, Conceição LEC (2003) The supply of amino acids during early feeding stages of marine fish larvae: a review of recent findings. Aquaculture 227:147–164
Rønnestad I, Koven W, Tandler A, Mordechai H, Fyhn HJ (1998) Utilisation of yolk fuels in developing eggs and larvae of European sea bass (Dicentrarchus labrax). Aquaculture 162:157–170
Shan X, Huang W, Cao L, Wu Y (2008a) Advances in studies of the effects of starvation on growth and development of fish larvae. J Ocean Univ China 7:319–326
Shan X, Quan H, Dou S (2008b) Effects of delayed first feeding on growth and survival of rock bream Oplegnathus fasciatus larvae. Aquaculture 277:14–23
Sommer F, Torraca V, Meijer AH (2020) Chemokine receptors and phagocyte biology in Zebrafish. Front Immunol 11:325
Sun Q, Yang Y, Li X, He B, Jia Y, Zhang N, Zhao R (2016) Folate deprivation modulates the expression of autophagy-and circadian-related genes in HT-22 hippocampal neuron cells through GR-mediated pathway. Steroids 112:12–19
Tsibris JCM, McCormick DB, Wright LD (1966) Studies on the binding and function of flavin phosphates with flavin mononucleotide-dependent enzymes. J Biol Chem 241:1138–1143
Wan Q, Liao Z, Rao Y, Yang C, Ji J, Chen X, Su J (2019) Transferrin receptor 1-associated iron accumulation and oxidative stress provides a way for grass carp to fight against reovirus infection. Int J Mol Sci 20:5857
Wendling NC, Bencic DC, Nagler JJ, Cloud JG, Ingermann RL (2004) Adenosine triphosphate levels in steelhead (Oncorhynchus mykiss) eggs: an examination of turnover, localization and role. Comp Biochem Physiol A Mol Integr Physiol 137 739 748
White E (2016) Autophagy and p53. CSH Perspect Med 6:a026120. https://doi.org/10.1101/2Fcshperspect.a026120
Wilson RP (1994) Utilization of dietary carbohydrate by fish. Aquaculture 124:67–80
Wu G, Fang YZ, Yang S, Lupton JR, Turner ND (2004) Glutathione metabolism and its implications for health. J Nutr 134:489–492
Xiang J, Liu X, Ren J, Chen K, Wang HL, Miao YY, Qi MM (2019) How does estrogen work on autophagy? Autophagy 15:197–211
Xu H, Liu E, Li Y, Li X, Ding C (2017) Transcriptome analysis reveals increases in visceral lipogenesis and storage and activation of the antigen processing and presentation pathway during the mouth-opening stage in zebrafish larvae. Int J Mol Sci 18:1634
Yahaya S, Lim L-S, Shaleh SMR, Mukai Y, Anraku K, Kawamura G (2011) Ontogenetic eye development and related behavioural changes in larvae and juveniles of barramundi Lates calcarifer (Bloch). Mar Freshw Behav Physiol 44:339–348
Yin MC, Blaxter JHS (1987) Feeding ability and survival during starvation of marine fish larvae reared in the laboratory. J Exp Mar Biol Ecol 105:73–83
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14
Yúfera M, Perera E, Mata-Sotres JA, Calduch-Giner J, Martínez-Rodríguez G, Pérez-Sánchez J (2017) The circadian transcriptome of marine fish (Sparus aurata) larvae reveals highly synchronized biological processes at the whole organism level. Sci Rep 7:12943
Yúfera M, Darias MJ (2007) The onset of exogenous feeding in marine fish larvae. Aquaculture 268:53–63
Zihni C, Mills C, Matter K, Balda MS (2016) Tight junctions: from simple barriers to multifunctional molecular gates. Nat Rev Mol Cell Biol 17:564–580
Acknowledgements
We thank Kampachi Farms, Mexico, at CIBNOR for providing Seriola rivoliana eggs and Novogene bioinformatics for technical support.
Funding
This study was supported by Centro de Investigaciones Biológicas del Noroeste and Consejo Nacional de Ciencia y Tecnología (CONACYT Grant: 258504 to R.C.R).
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All experimental protocols and procedures employed in fish in this study were ethically reviewed and approved by the Aquaculture Program Animal Welfare Committee of Centro de Investigaciones Biológicas del Noroeste S.C., La Paz, Baja California Sur, Mexico.
Consent for Publication
All authors gave their consent for the publication.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Guerrero-Tortolero, D.A., Vázquez-Islas, G. & Campos-Ramos, R. A Transcriptome Insight During Early Fish Larval Development Followed by Starvation in Seriola rivoliana. Mar Biotechnol 23, 749–765 (2021). https://doi.org/10.1007/s10126-021-10061-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-021-10061-4