Abstract
The study of the soil resistome is important in understanding the evolution of antibiotic resistance and its dissemination between the clinic and the environment. However, very little is known about the soil resistome, especially of those from deserts. Here, we characterize the bacterial communities, using targeted sequencing of the 16S rRNA genes, and both the resistome and the mobilome in Namib Desert soils, using shotgun metagenomics. We detected a variety of antibiotic resistance genes (ARGs) that conferred resistance to antibiotics such as elfamycin, rifampicin, and fluoroquinolones, metal/biocide resistance genes (MRGs/BRGs) conferring resistance to metals such as arsenic and copper, and mobile genetic elements (MGEs) such as the ColE1-like plasmid. The presence of metal/biocide resistance genes in close proximity to ARGs indicated a potential for co-selection of resistance to antibiotics and metals/biocides. The co-existence of MGEs and horizontally acquired ARGs most likely contributed to a decoupling between bacterial community composition and ARG profiles. Overall, this study indicates that soil bacterial communities in Namib Desert soils host a diversity of resistance elements and that horizontal gene transfer, rather than host phylogeny, plays an essential role in their dynamics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The misuse and overuse of antibiotics in the human, animal, and agricultural sectors are an important selective force in the evolution and the dissemination of antibiotic resistance (Bhullar et al., 2012; von Wintersdorff et al., 2016), which is one of the major global challenges of the twenty-first century as it poses serious risks to human health. The One Health concept is a holistic and interdisciplinary approach based on the idea that human and animal health are linked to the health of the ecosystems that they are a part of (Hernando-Amado et al., 2019). The importance of a One Health approach to antibiotic resistance (AR) is clear especially with respect to the dissemination of antibiotic resistance genes that have crossed habitat boundaries (Naidoo et al., 2020).
Soils are perhaps the most significant reservoir of ARGs as many soil microbes produce natural antibiotics to outcompete other microbes (Nesme and Simonet, 2015; Zhu et al., 2019). Indeed, most clinically relevant antibiotics such us streptomycin, tetracycline, and vancomycin originate from soil-dwelling actinomycetes (D’Costa et al., 2006; Cytryn, 2013). Microorganisms have developed a variety of resistance mechanisms (encoded by resistance genes) to prevent them from succumbing to these toxic metabolites (Thaker et al., 2013; Jiang et al., 2017). Many bacteria are naturally resistant to a broad spectrum of antibiotics (Demaneche et al., 2008; Allen et al., 2009), which may reflect their ability to produce more than one antibiotic, or be a by-product of their evolution in proximity to antibiotic-producing neighboring bacteria (Perry and Wright, 2013). Consequently, it is now clear that the soil environment harbors a plethora of both discovered and undiscovered resistance genes, which together constitute the soil resistome (Dantas and Sommer, 2012; Gillings et al., 2017).
Most types of soils contain heavy metals, some of which (at low concentrations) contribute to the metabolism of microorganisms (Knapp et al., 2017; Chen et al., 2019). Interestingly, metals can act as co-selective forces contributing to the increase in antibiotic resistance (Knapp et al., 2017). The relative importance of co-selection of resistance to both antibiotics and metals/biocides is likely to be different in different environments. For instance, some desert soils are known to have low anthropogenic antibiotic input but relatively high levels of metals over long periods of time (Knapp et al., 2011). Consequently in deserts, even in the absence of antibiotics, metals may provide a stronger and more persistent selective pressure for the environmental selection of antibiotic resistance (Zhao et al., 2018).
Resistance genes can move between soil microbial taxa via mobile genetic elements, collectively known as the mobilome (Carr et al., 2020). The mobilome facilitates the acquisition of traits between bacteria through several mechanisms via horizontal gene transfer (HGT), that is, via transformation (involving free DNA), transduction (involving bacteriophages), or conjugation (involving plasmids and integrative conjugative elements) (Peterson and Kaur, 2018). Soil bacteria undergo higher rates of gene transfer in “hotspots” (i.e., areas of higher nutritional content) such as the rhizosphere and manure-treated soil (Perry and Wright, 2013). However, the prevalence of horizontal gene transfer in native soil microbial communities and the effects it may have on soil processes are largely unknown and require further investigation (Fierer, 2017).
It has been hypothesized that many ARGs found in clinical isolates have originated from soil (Forsberg et al., 2012; Walsh and Duffy, 2013; McCann et al., 2019). Therefore, investigating the soil resistome could enable the detection of clinically relevant antibiotic resistance mechanisms (Forsberg et al., 2012; Walsh and Duffy, 2013; McCann et al., 2019). Furthermore, tracking antibiotic resistance genes in less impacted (i.e., deserts) or unimpacted (pristine) soils is important because this allows for the detection of background or intrinsic levels of antibiotic resistance in soil which may aid in estimating potential human health risks (Scott et al., 2020). Additionally, it might help to elucidate the extent of contamination of ARGs due to anthropogenic activity.
In this study, using targeted sequencing of the 16S rRNA genes, we characterized the bacterial communities and, using shotgun metagenomics, the soil resistome and mobilome in surface soils from the Namib Desert. The aims of this study were to investigate (1) the diversity and composition of ARGs and metal resistance genes/biocide resistance genes (MRGs/BRGs), (2) whether or not horizontal gene transfer (HGT) affected the distribution of the resistome, (3) possible co-selection of resistance with metals/biocides and antibiotics, and (4) a possible link between microbial community composition and the resistome.
Experimental procedures
Sampling, soil chemistry, and climate data
Eighteen surface soils (0 to 5 cm deep) were collected across a transect in the Namib Desert. Previously, anthropogenic influences across the desert have been minimal and were mostly limited to scientific expeditions. However, the nature-based tourism has increased drastically in the last decade, possibly increasing the anthropogenic pressure in the Namib Desert (Naidoo et al., 2020). The 18 sampling sites were spaced 5–10 km apart, and at each site, four aliquots of 50g of soil were taken at 100 m spacing and combined in a composite sample. Soils were collected using sterile methods and stored in sterile 50 ml polypropylene Falcon tubes (Grenier, Bio-One) at −80°C within 5 days after collection. Soils were analyzed for soil pH, total carbon, nitrogen phosphorous, and major cations (K, Na, Mg, Ca) at Bemlab, South Africa, using standard procedures.
DNA extraction and sequencing
Metagenomic DNA was extracted from the soil samples using the DNeasy Powersoil Kit (Qiagen, Valencia, CA, USA) as per the manufacturer’s instructions. DNA samples were submitted for sequencing at a commercial supplier for both metagenome and 16SrRNA sequencing (MR DNA Lab, Shallowater, TX, USA, http://www.mrdnalab.com). Shotgun metagenomic sequencing was performed on a HiSeq 2500 Ultra-High-Throughput Sequencing system (Illumina Inc., San Diego, CA, USA) using paired-ends (2 × 250 bp) for 500 cycles as per the manufacturer’s instructions.
Targeted sequencing of the 16S rRNA gene amplicons were amplified using primers 515F (5′-GTGYCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACNVGGGTWTCTAAT-3′). Paired-end 2 × 250 bp sequencing was performed on an Illumina MiSeq instrument according to manufacturer’s instructions (Illumina Inc., San Diego, CA, USA) with the parameters as described (https://support.illumina.com/16s-metagenomic-library-prep-guide-15044223-b.pdf). The metagenome sequence data and 16S amplicon sequence data are available on NCBI (PRJNA592367).
Metagenome assembly and ARG annotation
Raw reads were quality filtered using FastQC (Andrews, 2010) and trimmed using PrinSeq (Schmieder and Edwards, 2012). Quality reads were assembled using SPAdes v3.12 (Bankevich et al., 2012), with default settings and the “meta” parameter specified. The quality of each assembled metagenome (n = 18 ) was assessed using QUAST v5.0.2 (Mikheenko et al., 2018). Gene prediction was performed using Prodigal v2.6.3 (Hyatt et al., 2010) with the “meta” parameter specified. To identify antibiotic resistance genes that may have been acquired by HGT, predicted genes were compared against the ResFinder database (Zankari et al., 2012) by means of BLASTn with an E-value threshold of 1 × 10−6. The filtering parameters used were 100% similarity and a minimum query length of >50 %. Genes predicted by prodigal were also compared to the comprehensive antibiotic resistance database (CARD) (McArthur et al., 2013) by means of BLASTp with an E-value threshold of 1 × 10−6. Results were filtered for hits with a minimum percentage similarity of 87% and a minimum query length of >40%. These parameters were set with BLAST against all other databases used subsequently.
Mobile genetic elements and metal/biocide resistance gene annotation
To identify mobile genetic elements flanking ARGS, contigs were compared to the Mobile Genetic Elements Database (Pärnänen et al., 2018) by means of BLASTn. Detected plasmids were confirmed with the PlasmidFinder database (Carattoli et al., 2014). Metal and biocide resistance genes were detected by running BLASTp with these contigs against the BacMet database (Pal et al., 2014). The BacMet database is a manually curated database of antibacterial biocide and metal resistance genes.
16S rRNA amplicon sequence analysis
Sequence reads were demultiplexed using Sabre (https://github.com/najoshi/sabre), and primers were removed with cutadapt 2.10 (Martin, 2011). Amplicon sequence variants (ASVs) were resolved using DADA2 version 1.14 (Callahan et al., 2016) in R version 3.6.2 (R Core Team, 2013). Quality filtering and trimming were done using MaxEE = c(2,2) and truncLen = c(220, 200); all other parameters were set to default. The error rates were estimated by learnErrors, and sequences were dereplicated using derepFastq with default parameters. removeBimeraDenovo was used to remove chimeric sequences. Taxonomy was assigned against the SILVA non-redundant database version 138 (https://www.arb-silva.de).
Data analyses
The analyses were done in R version 3.6.2 using the packages phyloseq (McMurdie and Holmes, 2013), microbiome (Lahti et al., 2019), tidyverse (Wickham et al., 2019), and vegan (Oksanen et al., 2007). ASV alpha-diversity (richness, Shannon, inverse Simpson, Chao1) was calculated using the vegan package in R. Community data matrices were centered log-ratio transformed, and the Euclidian distance measure was used to generate an Aitchison dissimilarity matrix. To reveal the relationship between microbial composition and the resistome, the pairwise Pearson’s rank correlation (correlate alpha-diversity) was calculated, and a Mantel test was conducted directly from the distance matrices (correlate beta-diversity).
Results and discussion
Diversity and abundance of ARGs
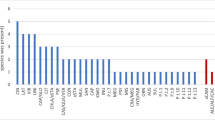
We obtained an average of 660,000 ORFs per sample, and the annotation of those ORFs against the CARD database resulted in a total of 6045 ORF hits with the identification of 46 ARGs. The identified ARGs spanned 26 ARG families exhibiting 17 resistance mechanisms (Supplementary Table S1). A great proportion of the ARGs were in low abundance and found in a single sample (Fig. 1), indicating that most ARGs in Namib Desert soils appear to be rare both in terms of local abundance and habitat specificity. Similar results have been found in other deserts (Van Goethem et al., 2018; McCann et al., 2019). At least two factors may contribute to the low diversity and abundance of ARGs in these environments. First, it has been hypothesized that microbial competition in desert soil microbiomes is low; this is because the allocation of most of the available resources is for survival under stressful environmental conditions and/or to the response of moisture availability (Fierer et al., 2012). Second, compared with other environments, deserts generally have very low human (and animal) population density and therefore are less impacted by human anthropogenic activity. Anthropogenic activity has been shown to increase the diversity and abundance of elements of the soil resistome (Wang et al., 2016).
The most common resistance mechanism detected was target alteration by mutation, followed by target protection, inactivation mechanisms, and various efflux mechanisms (Supplementary figure S1). Target alteration by mutation typically arises via chromosomal mutation and is therefore not laterally transferred. Mutations in the target-encoding genes often confer multidrug-resistance (D’Costa et al., 2006) and resistance to antibiotics such as cephalosporins (Demaneche et al., 2008) and fluoroquinolones (Riesenfeld et al., 2004). This resistance mechanism is essential for the continued evolution of ARGs to natural and synthetic antibiotics (Woodford and Ellington, 2007), especially in the case of clinically relevant isolates (Munita and Arias, 2016).
The most abundant group of ARGs, based on the drug resistance class (Fig. 2), were those that are known to confer resistance to aminoglycosides, elfamycins, glycopeptides, rifamycins, and those that were multi drug resistant. Here, resistance to rifamycins (RIF) was conferred by two different mechanisms (i.e., antibiotic target protection and enzymatic inactivation). Neither of these two mechanisms identified in the Namib soil resistome has been associated with resistance to rifampin in clinical environments (Tomlinson et al., 2016). However, resistance to RIF in these desert soil samples demonstrates that environmental bacteria may also possess multiple mechanisms of resistance to the same antibiotic (Spanogiannopoulos et al., 2012), which is an indication of bacterial adaptation to this environment.
Using the ResFinder database (Zankari et al., 2012), we found two putative horizontally acquired ARGs (Table 1): AAC’3-la and blaTEM-116. AAC’3-la (originally detected in clinical strains of P. aeruginosa) encodes acetyltransferases that mediate enzymatic inactivation of aminoglycosides. blaTEM-116 (originally detected in clinical strains of E. coli ) is a β-lactamase which showed 100% sequence similarity to the original TEM-116 (Song et al., 2005). Neither bacteriophage sequences nor MGEs were found flanking the AAC’3-la genes, and therefore, the vehicle of dissemination could not be established. However, we detected a plasmid (ColE1-like) flanking the blaTEM-116 gene (Naidoo et al., 2020). This particular variant of TEM-116 was first described on plasmids in clinical isolates in Korea and subsequently on plasmids in several clinical and non-clinical environments (Vignoli et al., 2005; Usha et al., 2008; Maravić et al., 2016).
Metal and biocide resistance genes (MRGs and BRGs) and co-selection with ARGs
The annotation of prodigal-predicted ORFs against the BacMet database resulted in a total of 143 ORF hits with the identification of 29 MRGs and BRGs, which spanned 16 families with 4 known resistance mechanisms (Supplementary Table S2). The most abundant MRGs and BRGs were those that are known to confer resistance to iron and triclosan (biocide) respectively (Fig. 3). Other common identified MRGs were those known to confer resistance to three or more compounds (multi-compound resistance); for example, the resistance gene MexK, which is part of a two-component resistance nodulation cell division (RND) efflux system known as MexJK. This system is responsible for the efflux of triclosan (a phenolic compound used in many personal care products as a biocide) and antibiotics such as macrolides and tetracycline (Chuanchuen et al., 2002). Whether the presence of MexK is indicative of anthropogenic impact in this environment needs to be further investigated.
While antibiotics are readily degraded in soil by multiple mechanisms, metals are essentially non-degradable and therefore potentially impose a long-term selection pressure on microbial communities. In fact, metals and biocides might even exert stronger selection pressure for antibiotic resistance than antibiotics themselves (Pal et al., 2017). The two major mechanisms involved in co-selection of resistance are co-resistance and cross-resistance (Murray et al., 2019). Co-resistance is the genetic linkage of resistance genes; that is, that genes responsible for resistance to two or more compounds are co-located on the same genetic element (i.e., plasmid, transposon or integron) (Seiler and Berendonk, 2012; Pal et al., 2017). An example of co-resistance in this study was the presence of an arsC gene on a plasmid that also contained the clinically relevant TEM-116 β-lactamase, suggesting possible co-selection of resistance to antibiotics and arsenic in Rhodococcus ruber (Actinobacteria) (Yashini Naidoo et al., 2020). On the other hand, cross-resistance occurs when a resistance gene or a single resistance mechanism simultaneously encodes for resistance to different compounds (Pal et al., 2017; Imran et al., 2019). In these soils, cross-resistance could occur via the MexJK system, which is able to efflux both triclosan and antibiotics such as tetracycline and macrolides (Chuanchuen et al., 2002). The levels of metals in these samples were not measured, but metal concentrations in nearby surface soils (<100 m distance) seem to be high ((i.e., Fe (17 290 ppm), Ni (29.35 ppm), Cu (25.36 ppm), Zn (59.63 ppm), As (3.02 ppm), Ag (0.20 ppm) and U (2.99 ppm)) (Conti et al., 2018). Thus, it is possible that the metals present in this environment may add a selection pressure that could increase the antibiotic tolerance level of microbes via co-selection mechanisms (Imran et al., 2019).
The occurrence of mobilome-related antibiotic resistance determinants in desert soils
Nine different mobile genetic elements (MGEs) were identified using the Mobile Genetics Elements database. In all metagenomes, insertion sequences were the most abundant MGEs (62% of hits), followed by transposases (33%), integrons (2.7%), and plasmids (2.6%) (Supplementary table S3). The only MGEs linked to any of the ARGs were plasmids (i.e., ColE1 with blaTEM-116). The insertion sequences detected were from the families IS91 and IS10. Of the two IS families, the relative abundance of IS91 sequences was much higher (92%), which might explain the scarcity of MGEs flanking ARGs. Members of the IS91 family of bacterial insertion sequences are primarily associated with pathogenicity determinants in animals (Schleinitz et al., 2010) and have been reported to very rarely flank ARGs, as they are not suited for the rapid dissemination mechanisms that follow antibiotic exposure in clinical environments (Garcillan-Barcia and De la Cruz, 2002). Conversely, members of the IS10 family are more rapidly disseminated and are known to upregulate efflux mechanisms resulting in increased antibiotic resistance (Siguier et al., 2014). Here, we identified class 1 integrons (int1), which have been a major driver in the spread of antibiotic resistance, specifically in clinical environments (Gillings, 2014). Class 1 integrons have been proposed as markers of anthropogenic pollution due to their common association with resistance to antibiotics and metals in both pathogenic and non-pathogenic bacteria (Gillings, 2014). The presence of int1 in these soils may infer a moderate human impact
Consistent with previous reports (Wang et al., 2016; Saenz et al., 2019), the overall abundance of MGEs was low. This suggests that only a small fraction of resistance determinants may be mobilized. The horizontal acquisition of resistance genes is suggested to be rare in environments with both low anthropogenic influence and low nutrient levels (Forsberg et al., 2014) as in the case of the Namib Desert. Furthermore, the regulation of genetic machinery is highly responsive to the environment. For instance, the rates of horizontal gene transfer in soils largely depends on variables such as soil moisture and temperature, pH, and soil type (Aminov, 2011). Desert environments experience extreme fluctuations in temperature and highly erratic precipitation patterns, have low nutrient levels, and are generally subject to low levels of anthropogenic impact, compared to other edaphic environments (Makhalanyane et al., 2015). We suggest that these factors together contribute to the observed low abundance of MGEs in Namib Desert soil metagenomes. Nevertheless, the presence of horizontally acquired resistance genes indicates that mobilization of ARGs is a factor in these soils. Like all deserts, the Namib also is impacted by transient wildlife (Stein et al., 2008) that may contribute to the spread of resistance determinants (Allen et al., 2010). It is also noted that over the past few decades, there has been a substantial growth of tourism in the Namib Desert (Woyo and Amadhila, 2018), which may increase the possible routes of transmission for acquired ARGs.
Decoupling between the microbiome and the resistome
Several studies have shown that bacterial community composition is a key driver that shapes the distribution and abundance of ARGs, for instance, in surface soils (Forsberg et al., 2014), underground coal mine soils (Dunivin and Shade, 2018), and in sewage sludge (Su et al., 2015). However, results from the Mantel test revealed that there were no significant correlations (Mantel r = 0.2, P > 0.05) between the resistome and the microbiome (ASVs derived from 16S rRNA gene sequences). This suggests that microbial community composition was not a major contributor to the variance in resistance genes in Namib Desert soils and suggest that variations in resistance genes abundance in this soil might be driven by other factors, such as the mobilome. Mobile genetic elements are known to play a very important role in shaping ARG dynamics, as they can transfer genes between distantly related taxa, thereby causing a decoupling of bacterial community and ARG profiles (Fang et al., 2019). Alternatively, a decoupling between the microbiome and the resistome can also be expected if most of the mobile genetic elements are hosted by a limited number of taxa widely distributed across the soil. Further research would be necessary to confirm this.
Conclusion
This study represents, to the best of our knowledge, the most comprehensive analyses of soil resistomes conducted to date in hot deserts. We have demonstrated that Namib Desert soils contain diverse ARGs and MRGs/BRGs, some of which seem to be acquired on mobile elements. The presence of these acquired resistance genes supports the assumption that vectors such as terrestrial animals, birds, and humans may be partly responsible for the spread of resistance determinants in these soils which highlights the importance of the One Health approach to the burden of antibiotic resistance. In addition, the co-selection of resistance to antibiotics with metals and/or biocides could lead to the persistence of ARGs in the soil microbiome. The presence of ARGs in this soil ecosystem does not necessarily pose a significant health threat to humans and animals living in or visiting this environment. However, the concern is that, like in other environmental settings, the mobilization of these resistance determinants and their expression in bacterial pathogens could present difficulty for treatment options in the clinic.
Data availability
The metagenome sequence data and 16S amplicon sequence data are available on NCBI (PRJNA592367) and can be accessed at https://www.ncbi.nlm.nih.gov/search/all/?term=PRJNA592367.
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8:251–259
Allen HK, Moe LA, Rodbumrer J, Gaarder A, Handelsman J (2009) Functional metagenomics reveals diverse β-lactamases in a remote Alaskan soil. ISME J 3:243–251
Aminov RI (2011) Horizontal gene exchange in environmental microbiota. Front Microbiol 2
Andrews, S. (2010) Fastqc: a quality control tool for high throughput sequence data.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks ED, Johnston MD et al (2012) Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS One 7(4):e34953
Callahan BJ, Mcmurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Carattoli A, Zankari E, Garciá-Fernández A, Larsen MV, Lund O, Villa L et al (2014) In silico detection and typing of plasmids using plasmid finder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 58:3895–3903
Carr VR, Shkoporov A, Hill C, Mullany P, Moyes DL (2020) Probing the mobilome : discoveries in the dynamic microbiome. Trends Microbiol 29(2):158–170
Chen J, Li J, Zhang H, Shi W, Liu Y (2019) Bacterial heavy-metal and antibiotic resistance genes in a copper tailing dam area in Northern China. Front Microbiol 10:1–12
Chuanchuen R, Narasaki CT, Schweizer HP (2002) The MexJK efflux pump of Pseudomonas aeruginosa requires OprM for antibiotic efflux but not for efflux of triclosan. J Bacteriol 184:5036–5044
Conti E, Costa G, Liberatori G, Vannuccini ML, Protano G, Nannoni F, Corsi I (2018) Ariadna spiders as bioindicator of heavy elements contamination in the Central Namib Desert. Ecol Indic 95:663–672
Cytryn E (2013) The soil resistome: the anthropogenic, the native, and the unknown. Soil Biol Biochem 63:18–23
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Science 311:374–377
Dantas G, Sommer MOA (2012) Context matters - the complex interplay between resistome genotypes and resistance phenotypes. Curr Opin Microbiol 15:577–582
Demaneche S, Sanguin H, Pote J, Navarro E, Bernillon D, Mavingui P et al (2008) Antibiotic-resistant soil bacteria in transgenic plant fields. Proc Natl Acad Sci 105:3957–3962
Dunivin TK, Shade A (2018) Community structure explains antibiotic resistance gene dynamics over a temperature gradient in soil. FEMS Microbiol Ecol 94:fiy016
Fang P, Peng F, Gao X, Xiao P, Yang J (2019) Decoupling the dynamics of bacterial taxonomy and antibiotic resistance function in a subtropical urban reservoir as revealed by high-frequency sampling. Frontiers in Microbiology 10:1448
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590
Fierer N, Leff JW, Adams BJ, Nielsen UN, Bates ST, Lauber CL et al (2012) Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc Natl Acad Sci 109:21390–21395
Forsberg KJ, Patel S, Gibson MK, Lauber CL, Knight R, Fierer N, Dantas G (2014) Bacterial phylogeny structures soil resistomes across habitats. Nature 509:612–616
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Science 337:1107–1111
Garcillan-Barcia MP, De la Cruz F (2002) Distribution of IS 91 family insertion sequences in bacterial genomes : evolutionary implications. FEMS Microbiol Ecol 42:303–313
Gillings MR (2014) Integrons: past, present, and future. Microbiol Mol Biol Rev 78:257–277
Gillings MR, Paulsen IT, Tetu SG (2017) Genomics and the evolution of antibiotic resistance. Ann N Y Acad Sci 1388:92–107
Hernando-Amado S, Coque TM, Baquero F, Martínez JL (2019) Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nat Microbiol 4:1432–1442
Hyatt D, Chen G-L, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119
Imran M, Das KR, Naik MM (2019) Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: an emerging health threat. Chemosphere 215:846–857
Jiang X, Ellabaan MMH, Charusanti P, Munck C, Blin K, Tong Y et al (2017) Dissemination of antibiotic resistance genes from antibiotic producers to pathogens. Nat Commun 8:1–7
Knapp CW, Callan AC, Aitken B, Shearn R, Koenders A, Hinwood A (2017) Relationship between antibiotic resistance genes and metals in residential soil samples from Western Australia. Environ Sci Pollut Res 24:2484–2494
Knapp CW, McCluskey SM, Singh BK, Campbell CD, Hudson G, Graham DW (2011) Antibiotic resistance gene abundances correlate with metal and geochemical conditions in archived Scottish soils. PLoS One 6(11):e27300
Lahti, L., Shetty, S., Blake, T., and Salojarvi, J. (2019) Tools for microbiome analysis in R. Microbiome package version. Bioconductor.
Makhalanyane TP, Valverde A, Gunnigle E, Frossard A, Ramond J, Cowan DA (2015) Microbial ecology of hot desert edaphic systems. FEMS Microbiol Rev 39:203–221
Maravić A, Skočibušić M, Fredotović Ž, Šamanić I, Cvjetan S, Knezović M, Puizina J (2016) Urban riverine environment is a source of multidrug-resistant and ESBL-producing clinically important Acinetobacter spp. Environ Sci Pollut Res 23:3525–3535
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequence reads. EMBnet.journal 17:10–12
McArthur A, Waglechner N, Nizam F, Yan A, Azad MA, Baylay AJ et al (2013) The comprehensive antibiotic resistance database. Antimicrob Agents Chemother 57:3348–3357
McCann CM, Christgen B, Roberts JA, Su J, Arnold KE, Gray ND et al (2019) Understanding drivers of antibiotic resistance genes in High Arctic soil ecosystems. Environ Int 125:497–504
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217
Mikheenko A, Prjibelski A, Saveliev V, Antipov D, Gurevich A (2018) Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 34:i142–i150
Munita JM, Arias CA (2016) Mechanisms of Antibiotic Resistance. Microbiol Spectr 4:1–37
Murray AK, Zhang L, Snape J, Gaze WH (2019) Comparing the selective and co-selective effects of different antimicrobials in bacterial communities. Int J Antimicrob Agents 53:767–773
Naidoo Y, Valverde A, Cason ED, Pierneef RE, Cowan DA (2020) A clinically important, plasmid-borne antibiotic resistance gene (β-lactamase TEM-116 ) present in desert soils. Sci Total Environ 719:1–6
Nesme J, Simonet P (2015) The soil resistome: a critical review on antibiotic resistance origins, ecology and dissemination potential in telluric bacteria. Environ Microbiol 17:913–930
Oksanen J, Kindt R, Legendre P, O’Hara B, Stevens M (2007) The vegan package. Community Ecol Packag 10:631–637
Pal C, Asiani K, Arya S, Rensing C, Stekel DJ, Larsson DGJ, Hobman JL (2017) Metal resistance and its association with antibiotic resistance. In: Advances in Microbial Physiology. Elsevier Ltd., pp 261–313
Pal C, Bengtsson-Palme J, Rensing C, Kristiansson E, Larsson DGJ (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42:737–743
Pärnänen K, Karkman A, Hultman J, Lyra C, Bengtsson-Palme J, Larsson DGJ et al (2018) Maternal gut and breast milk microbiota affect infant gut antibiotic resistome and mobile genetic elements. Nat Commun 9:1–11
Perry JA, Wright GD (2013) The antibiotic resistance “mobilome”: searching for the link between environment and clinic. Front Microbiol 4
Peterson E, Kaur P (2018) Antibiotic resistance mechanisms in bacteria: relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Front Microbiol 9:1–21
R Core Team (2013) A language and environment for statistical computing. R Found Stat Comput
Riesenfeld CS, Goodman RM, Handelsman J (2004) Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ Microbiol 6:981–989
Saenz JS, Airo A, Schulze-Makuch D, Schloter M, Vestergaard G (2019) Functional traits co-occurring with mobile genetic elements in the microbiome of the Atacama Desert. Diversity 11:1–20
Schleinitz KM, Vallaeys T, Kleinsteuber S (2010) Structural Characterization of ISCR8, ISCR22, and ISCR23, subgroups of IS91-like insertion elements. Antimicrob Agents Chemother 54:4321–4328
Schmieder R, Edwards R (2012) Insights into antibiotic resistance through metagenomic approaches. Future Microbiol 7:73–89
Scott LC, Lee N, Aw TG (2020) Antibiotic resistance in minimally human-impacted environments. Environ Res Public Heal 17:1–12
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol 3
Siguier P, Gourbeyre E, Chandler M (2014) Bacterial insertion sequences: their genomic impact and diversity. FEMS Microbiol Rev 38:865–891
Song JS, Jeon JH, Lee JH, Jeong SH, Jeong BC, Kim S-J et al (2005) Molecular characterization of TEM-type B-lactamases identified in cold-seep sediments of Edison seamount. J Microbiol 43:172–178
Spanogiannopoulos P, Thaker M, Koteva K, Waglechner N, Wright GD (2012) Characterization of a rifampin-inactivating glycosyltransferase from a screen of environmental Actinomycetes. Antimicrob Agents Chemother 56:5061–5069
Stein AB, Fuller TK, Marker LL (2008) Opportunistic use of camera traps to assess habitat-specific mammal and bird diversity in northcentral Namibia. Biodivers Conserv 17:3579–3587
Su J-Q, Wei B, Ou-Yang W-Y, Huang F-Y, Xu H-J, Zhu Y-G (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol 49:7356–7363
Thaker MN, Wang W, Spanogiannopoulos P, Waglechner N, King AM, Medina R, Wright GD (2013) Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat Biotechnol 31(10):922–927
Tomlinson JH, Thompson GS, Kalverda AP, Zhuravleva A, O’Neill AJ (2016) A target-protection mechanism of antibiotic resistance at atomic resolution : insights into FusB-type fusidic acid resistance. Sci Rep 6:1–12
Usha G, Chunderika M, Prashini M, Willem SA, Yusuf ES (2008) Characterization of extended-spectrum β-lactamases in Salmonella spp. at a tertiary hospital in Durban, South Africa. Diagn Microbiol Infect Dis 62:86–91
Van Goethem MW, Pierneef R, Bezuidt OKI, Van De Peer Y, Cowan DA, Makhalanyane TP (2018) A reservoir of “historical” antibiotic resistance genes in remote pristine Antarctic soils. Microbiome 6
Vignoli R, Varela G, Mota MI, Cordeiro NF, Power P, Ingold E et al (2005) Enteropathogenic Escherichia coli strains carrying genes encoding the PER-2 and TEM-116 extended-spectrum β-lactamases isolated from children with diarrhea in Uruguay. J Clin Microbiol 43:2940–2943
von Wintersdorff CJH, Penders J, van Niekerk JM, Mills ND, Majumder S, van Alphen LB et al (2016) Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front Microbiol 7:173
Walsh F, Duffy B (2013) The culturable soil antibiotic resistome: a community of multi-drug resistant bacteria. PLoS One 8:e65567
Wang F, Stedtfeld RD, Kim O-S, Chai B, Yang L, Stedtfeld TM et al (2016) Influence of soil characteristics and proximity to antarctic research stations on abundance of antibiotic resistance genes in soils. Environmental science & technology 50(23):12621–12629
Wickham H, Averick M, Bryan J, Chang W, Mcgowan LDA, François R et al (2019) Welcome to the Tidyverse Tidyverse package. J open source Softw 4:1–6
Woodford N, Ellington MJ (2007) The emergence of antibiotic resistance by mutation. Clin Microbiol Infect 13:5–18
Woyo E, Amadhila E (2018) Desert tourists experiences in Namibia: a Netnographic Approach. African J Hosp Tour Leis 7:1–13
Zankari E, Hasman H, Cosentino S, Vestergaard M, Rasmussen S, Lund O et al (2012) Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67:2640–2644
Zhao Y, Cocerva T, Cox S, Tardif S, Su J, Zhu Y, Brandt K (2018) Evidence for co-selection of antibiotic resistance genes and mobile genetic elements in metal polluted urban soils. Sci Total Environ 656:512–520
Zhu YG, Zhao Y, Zhu D, Gillings M, Penuelas J, Ok YS et al (2019) Soil biota , antimicrobial resistance and planetary health. Environ Int 131:1–7
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was funded by the National Research Foundation, South Africa, grant number 112714.
Author information
Authors and Affiliations
Contributions
Data analysis was carried out by Y.N., A.V., and R.P. The first draft of the manuscript was written by Y.N., and all authors reviewed and commented on previous versions. All authors have read an approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
No ethics approval was required for this study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Naidoo, Y., Pierneef, R.E., Cowan, D.A. et al. Characterization of the soil resistome and mobilome in Namib Desert soils. Int Microbiol (2023). https://doi.org/10.1007/s10123-023-00454-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-023-00454-x