Abstract
The widespread of different NDM variants in clinical Enterobacterales isolates poses a serious public health concern, which requires continuous monitoring. In this study, three E. coli strains carrying two novel blaNDM variants of blaNDM-36, -37 were identified from a patient with refractory urinary tract infection (UTI) in China. We conducted antimicrobial susceptibility testing (AST), enzyme kinetics analysis, conjugation experiment, whole-genome sequencing (WGS), and bioinformatics analysis to characterize the blaNDM-36, -37 enzymes and their carrying strains. The blaNDM-36, -37 harboring E. coli isolates belonged to ST227, O9:H10 serotype and exhibited intermediate or resistance to all β-lactams tested except aztreonam and aztreonam/avibactam. The genes of blaNDM-36, -37 were located on a conjugative IncHI2-type plasmid. NDM-37 differed from NDM-5 by a single amino acid substitution (His261Tyr). NDM-36 differed from NDM-37 by an additional missense mutation (Ala233Val). NDM-36 had increased hydrolytic activity toward ampicillin and cefotaxime relative to NDM-37 and NDM-5, while NDM-37 and NDM-36 had lower catalytic activity toward imipenem but higher activity against meropenem in comparison to NDM-5. This is the first report of co-occurrence of two novel blaNDM variants in E. coli isolated from the same patient. The work provides insights into the enzymatic function and demonstrates the ongoing evolution of NDM enzymes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The plasmid-encoded New Delhi metallo-β-lactamase (NDM) is one of the most common carbapenemases worldwide [1]. Its emergence heralds a new era of antibiotic resistance due to the ability to hydrolyze almost all known β-lactam antibiotics and the rapid worldwide dissemination [2]. In 2009, the first NDM variant (NDM-1) was reported in Klebsiella pneumoniae isolated from a Swedish patient of Indian origin who had a urinary tract infection [3]. Following the first report, 41 different NDM variants have been identified in numerous species of Enterobacteriaceae and common nonfermentative Gram-negative bacilli [4] (NDM-1 to NDM-41; NDM-32 is assigned but without any information in NCBI).
The continuous evolution of NDM enzymes under the selection pressure could foster the emergence of new variants that possess different catalytic activities toward β-lactam agents [5]. For example, the NDM-5 variant showed enhanced hydrolytic activity compared with NDM-1 [6], and circular dichroism spectroscopy data revealed significant changes in the secondary structure of NDM variants [7]. Thus, a close surveillance of NDM-producing pathogens should be considered for continuous monitoring of the spread of NDM variants. Here, we describe the first detection of two novel NDM enzymes, designated NDM-36 and NDM-37, recovered from a patient with refractory urinary tract infection in China during 2020.
Methods
Bacterial strains
Three carbapenem-resistant E. coli strains (blaNDM positive) were isolated from the urine samples of a 62-year-old female patient with unilateral indwelling ureteral stents. The patient underwent cystectomy and chemotherapy for recurrence of ovarian and fallopian cancer three months ago. The first E. coli strain (JNQH497-NDM-37) was recovered in an outpatient clinic. Based on the antibiotic susceptibility testing results, empirical levofloxacin treatment (500 mg qd) was then started for urinary tract infection. After 3 weeks, the second strain (JNQH498-NDM-36) was isolated from the urine during hospital admission. The patient was then given meropenem (1000 mg intravenously [i.v.] q8h). The third E. coli strain (JNQH462-NDM-36) was identified from the urine 5 weeks after admission. Her conditions were improved after the removal of ureteral stents via cystoscopy, continuous meropenem treatment as well as implementation of nutritional support. The patient was discharged home on hospital day 16. Ethics committee approval of this study was obtained from the institutional review board of the First Affiliated Hospital of Xiamen University, and informed consent from the patient was also obtained.
Antimicrobial susceptibility testing (AST)
MICs for all the tested strains were determined by broth microdilution method using a bacterial inoculum of 5 × 105 CFU/ml according to CLSI performance standards. For ceftazidime-avibactam (CAZ-AVI) and aztreonam-avibactam (ATM-AVI) MICs evaluation, AVI was tested at a fixed concentration of 4 mg/L, while CAZ and ATM were added at different concentrations ranged from 0.0312 to 64 mg/L and 0.0156 to 32 mg/L, respectively.
Cloning of bla NDM variants
The promoter and full length of the blaNDM genes were amplified with primers NDM-F-EcoRI (5′-CCGGAATTCTTGAAACTGTCGCACCTCAT-3′) and NDM-R-XbaI (5′-CTAGTCTAGAACGCCTCTGTCACATCGAA-3′) using PrimSTAR Max DNA Polymerase (Takara, China). After restriction enzyme digestion, the PCR products were ligated to the vector PET28a to generate PET28a-NDM-5, PET28a-NDM-36, PET28a-NDM-37 respectively. The correct constructs were confirmed by Sanger sequencing, followed by transformation into E. coli DH5α. Antimicrobial susceptibilities of these constructs were determined as described above. The empty pET28a plasmid was used as a control.
Expression of the NDM proteins
The sequences of NDM-5, -36, -37 without peptide signal region were amplified by PCR using primers EcoRI-NDM (29-271AA)-F (5′—CCGGAATTCATGGAATTGCCCAATAT—3′) and HindIII-NDM (29-271AA)-R (5′—CCCAAGCTTTCAGCGCAGCTTGTCGGCC -3′), followed by insertion into plasmid pET28a in E. coli BL21 (Invitrogen). Protein was expressed in E. coli strain BL21 grown in LB at 37 ℃. Once an OD600 of 0.4–0.6 was reached, 0.1 mM ZnCl2 and 0.5 mM isopropyl-β-d-thiogalactoside were added into the LB medium. The temperature was then lowered to 20 ℃, and the expression was allowed to occur overnight. Later, the cells were lysed by sonication, and the supernatant was loaded to a HisTrap™ HP column (GE Healthcare, Little Chalfont, UK). Finally, the purified proteins were dialysised in the buffer (50 mM HEPES, 100 μM ZnCl2, 250 mM NaCl, and 20 mg/L BSA) at 4 °C overnight [8].
Steady-state kinetic parameters
Steady-state kinetic experiments were performed following the hydrolysis of the β-lactams at 25 °C in 50 mM HEPES (pH 7.5) plus 100 μM ZnCl2. The data of the real-time absorbances of meropenem (298 nm), imipenem (297 nm), ceftazidime (257 nm), aztreonam (318 nm), cefotaxime (264 nm), cefepime (254 nm), piperacillin (232 nm), ceftriaxone (240 nm), and ampicillin (235 nm) were collected with a SHIMADZU UV2550 spectrophotometer (Kyoto, Japan). Kinetic parameters were determined under initial-rate conditions using the GraphPad Prism 8.1 software to generate Michaelis–Menten curves or by analyzing the complete hydrolysis time courses [9].
Conjugation experiment
Conjugation experiments were performed using E. coli J53AziR as recipients as previously described [10]. Briefly, overnight cultures of the donor strain (JNQH497, 498) and the recipient strains were mixed (1:1) and applied to 0.45-μm filter paper respectively, which were then placed on an LB agar plate, followed by overnight culture at 37 ℃. Transconjugants were selected on Mueller–Hinton agar containing sodium azide (100 mg/L)/meropenem (2 mg/L) for transconjugates. The selected transconjugants were confirmed by PCR targeting the blaNDM gene. Conjugation frequency was calculated by dividing the number of transconjugants by the number of recipient cells.
Pulsed-field gel electrophoresis (PFGE)
To further explore the relatedness of JNQH497, 498, 462 strains, we used PFGE to analyze the genetic relatedness. PFGE of Xbal-digested genomic DNA samples were performed with a CHEF MAPPER XA apparatus (Bio-Rad, USA), as previously described [11].
Whole-genome sequencing (WGS)
The strains were subject to next generation sequencing using the Illumina HiSeq system (Illumina, San Diego, CA, USA). Genomic DNA was isolated using a WizardR Genomic DNA Purification Kit (Promega, Madison, WI, USA). Sequencing reads were de novo assembled using Spades 3.12.0 [12]. To resolve the complete plasmid sequence carrying blaNDM in JNQH497 and JNQH498, the Oxford Nanopore (MinION system) sequencing was conducted and assembled with Illumina sequences to achieve a high-quality genome assembly. The hybrid assembly was performed using Unicycler v0·5.0 [13]. The whole-genome sequences were annotated by Prokka [14] automatically followed by manual curation.
Genomic analysis
In silico multi-locus sequence typing was performed using MLST 2.0 [15], while the acquired antimicrobial resistance genes were identified using the ABRicate program (https://github.com/tseemann/abricate) to query the CARD database (http://genomicepidemiology.org/). Identification of serotypes was performed using ECTyper (v.1.0) [16] with default parameters (https://github.com/phac-nml/ecoli_serotyping). The plasmid replicons in the sequenced isolates were identified using PlasmidFinder 2.0 [17]. Single-nucleotide polymorphisms (SNPs) and small insertions (INS) were detected using Snippy v3.2 (https://github.com/tseemann/snippy) by mapping the Illumina sequence reads of the JNQH498 and JNQH462 to the complete chromosome sequence of isolate JNQH497. Blastn was used to examine sequences homologous to the sequenced plasmids in the NCBI database. Comparison between homologous plasmids was conducted using CGview server [18]. OriT Finder was used to determine the conjugation module [19]. In order to examine the plasmid replicon distribution of blaNDM-bearing plasmids, plasmid sequences were downloaded from NCBI (https://ftp.ncbi.nlm.nih.gov/refseq/release/plasmid) and compared. Plasmid distance trees were generated using Mashtree [20]. The amino acid sequences of blaNDM were retrieved from BLDB [4] and NCBI database, and were aligned using Clustal Omega [21]. Evolutionary analyses were conducted in MEGA X for inferring maximum-likelihood phylogenies [22].
Results
Antimicrobial susceptibility testing
Broth microdilution susceptibility testing showed that JNQH497, JNQH462, and JNQH498 were resistant to ampicillin, cefazolin, cefotaxime, ceftazidime, meropenem, amikacin, levofloxacin, and ceftazidime/avibactam, but were susceptible to aztreonam (MICs, ≤ 0.25 mg/L), aztreonam/avibactam (MICs, ≤ 0.008 mg/L), and colistin (MICs = 0.125 mg/L) (Table 1). NDM-36-producing isolates (JNQH498, JNQH462) exhibited decreased susceptibility to imipenem and meropenem comparing with the isolate harboring NDM-37 (JNQH497). Of note, JNQH498 and JNQH462 were resistant to imipenem (MIC, 8 mg/L), while JNQH497 was intermediate to imipenem with an MIC value of 2 mg/L.
Identification of bla NDM-36, -37 in E. coli strains
Whole-genome sequencing analysis showed all strains belonged to sequence type 227 (ST227) and O9:H10 serotype. Two novel NDM-encoding genes were identified in JNQH462 and JNQH498, designated blaNDM-36 (NG_076641.1) and blaNDM-37 (NG_076642.1), respectively. Relative to blaNDM-5, blaNDM-37 contained one missense point mutations at positions 781 (C → T), generating amino acid substitution His261Tyr. Relative to NDM-37, NDM-36 contained one additional missense point mutation at position 698 (C → T), resulting in amino acid substitution Ala233Val.
Combination of long-read and short-read sequencing revealed that strain JNQH497 harbored a 4.75-Mb chromosome and two plasmids, designated pJNQH497-1 (263-Kb), and pJNQH497-2 (103.3-Kb). blaNDM-37 was carried by pJNQH497-1 which belongs to IncHI2-type plasmid. JNQH498 harbored a 4.75-Mb chromosome and one plasmid, designated pJNQH498-1 (263-Kb). blaNDM-36 was carried by pJNQH498-1 which also belongs to IncHI2-type plasmid. blaNDM-36 and blaNDM-37 were located in an ΔTn125-like region with the structure of “tnpA-IS5-blaNDM-37-bleMBL-trpF-tat,” containing Tn3 family transposase IS3000 upstream and IS1380 family transposase ISEcp1 downstream (Fig. 1). In addition to the truncated Tn125 cluster, multiple aminoglycoside resistance genes [APH(4)-Ia, AAC(3)-IV, APH(3′)-Ia, APH(3′)-Ib, APH(6)-Id, ANT(3′)-IIa, AAC(6′)-Ib], fluroquinolone [rmpA, AAC(6′)-Ib-cr), and sulphonamide resistance genes (sul1, sul2) were also found on the same plasmid (Fig. 2). Further, OXA-1 and TEM-150 genes producing ESBL were located on pJNQH497-1 and pJNQH497-2 respectively in JNQH497. OXA-1 gene was located on pJNQH498-1 while TEM-150 was absent in JNQH498.
Comparison of pJNQH497-1 (CP091926), pJNQH498-1 (CP104385), p8c59NDM (NZ_MT407547.1), pHNGD64-NDM (NZ_MW296099.1), pNDM33-1 (NZ_CP076648.1), and draft genome sequences of JNQH462. Open reading frames (ORFs) of pJNQH497-1 are shown as the outermost ring, with plasmid replicons, plasmid transfer associated (oriT, T4SS, T4CP, relaxase), and antimicrobial resistance genes highlighted. pJNQH497-1 was used as the reference for Blastn comparison
Comparison of sequences and PFGE patterns
Comparison of complete sequences of JNQH497 and JNQH498 showed the blaNDM-36 harboring plasmid (pJNQH498-1) was almost the same as the blaNDM-37 harboring plasmid (pJNQH497-1, 99.99% nucleotide identity and 100% coverage) (Fig. 2). There were only 13 SNPs between the two plasmids including the c.698C > T in blaNDM gene. Blastn analysis revealed that pJNQH497-1 was almost identical (100% query coverage and 99% identity) to the plasmid p8C59-NDM (NZ_MT407547.1) and displayed high similarity with pHNGD64-NDM [23] (NZ_MW296099.1, 89% query coverage and 99.8% identity) and pNDM33-1 [24] (NZ_CP076648.1, 86% query coverage and 99.9% identity). p8C59-NDM, pHNGD64-NDM, and pNDM33-1 also belonged to IncHI2-type and were carried by E. coli isolated from animal sources in China; however, they all harbored blaNDM-5, and the host E. coli strains belonged to ST10, ST4063, and ST48, respectively. In addition, a premature stop codon was introduced into the coding region of ompD (c.238G > T p.Glu80*) in JNQH498 and JNQH462 strains while it was absent in JNQH497 (Table S1).
The major plasmid types carrying blaNDM from reference NCBI database (n = 876) included IncX3 (29.68%), IncFII (15.41%), IncFIB (12.79%), and IncC (9.59%) (Fig. 3). Twelve blaNDM harboring plasmids were found to be IncHI2 type (1.37%), which were mainly found in mainland China with the exception of two plasmids from Taiwan and Nepal respectively. Notably, three blaNDM bearing IncHI2 plasmids also coharbored mcr9.1 colistin resistance genes (Fig. 4).
The PFGE showed JNQH497, 498, 462 had highly similar band patterns which indicated they were closely related (Fig. 5).
Transferability of bla NDM harboring plasmids and conjugation module analysis
Conjugation assays showed the blaNDM-36 and blaNDM-37 harboring IncHI2-type plasmids were successfully transferred into E. coli J53 from JNQH498 and JNQH497 strains. E. coli J53 transconjugants acquired resistance to levofloxacin and most β-lactam antibiotics except aztreonam/avibactam (Table 1), which indicated the resistant markers to fluroquinolones were co-transferred with blaNDM genes. The conjugation frequency was 10−3 per recipient cell for JNQH497, whereas it was only 10−8 for JNQH498. Conjugation module analysis revealed the conjugation genes are in two separate regions: transfer region 1 carries the origin of transfer site (oriT), type IV coupling protein gene (T4CP), and genes encoding the relaxase and some type IV secretion components. Region 2 encodes most type IV secretion proteins (Fig. 2). Blastn analysis revealed all the conjugation modules were also found in JNQH498, JNQH462 strains.
Expression of the NDM proteins and enzyme activity analysis
Susceptibility testing of pET28a constructs showed that expression of the blaNDM-36 and blaNDM-37 genes in E. coli DH5α conferred resistance to most of the tested β-lactams except aztreonam and ATM/AVI. Kinetic data showed that NDM-36 had higher affinity to cefotaxime than that of NDM-37, with the Km value reduced by 82.62 μM, whereas NDM-36 displayed slightly lower affinity than those of NDM-5, -37 for imipenem and meropenem. The kcat/Km ratio for ampicillin and cefotaxime of NDM-36 was higher than those of NDM-37, -5, but imipenem kcat/Km ratio was slighter higher than those of NDM-37, -5. In comparison to NDM-5, although NDM-36, -37 had lower kcat/Km ratio for imipenem, they had higher kcat/Km ratio for meropenem. These results suggested NDM-36 had higher hydrolytic activity toward ampicillin and cefotaxime relative to NDM-37, -5, and that NDM-37, -36 had lower catalytic activity against imipenem but higher activity against meropenem relative to NDM-5 (Table 2).
Phylogenetic analysis of NDM protein sequences
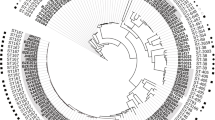
Phylogenetic analysis of the protein sequence of NDM variants is represented in Fig. 6. Evolutionary analysis of the amino acid sequences showed the amino acids were substituted at 30 polymorphic sites except for NDM-18, which has five amino acids tandem repeat (QRFGD) at positions 44 to 48 relative to NDM-1 [25]. The NDM variants had most hotspot mutation at amino acid positions 88, 154, and 130 (Fig. 6).
Evolutionary analysis and genetic variations among the NDM variants and its first source of spread. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches. Schematic representation of blaNDM-1 gene in the alignment showing the mutations at various nucleotide positions leading to the occurrence of NDM variants. Each unique color of NDM variants in the column showing mutant residues at different position
Discussion
The widespread of NDM variants among E. coli strains and other Enterobacteriaceae isolates represents a large threat to the public health globally [26]. To date, a total of 41 NDM variants have been named and described, of which 40 sequences have been deposited in the GenBank database. In this study, two novel blaNDM variants were carried by E. coli strains isolated from the same patient. Sequence analysis showed that NDM-37 differed from NDM-5 by a single amino acid substitution (His261Tyr) due to one missense point mutations at positions 781 (C → T). One additional missense point mutation at position 698 (C → T) in blaNDM-37 resulting in NDM-36. As such, this study provides a good explanation to increase our understanding that blaNDM variants are undergoing continuous evolution and thus need to be closely monitored.
WGS revealed the new variants of blaNDM-36, -37 were located on a conjugational IncHI2-type plasmid. IncHI2 plasmids are larger than most of the other conjugative plasmids and have been found to be associated with various resistance genes including mcr, ESBL, and carbapenemase encoding genes in Enterobacteriaceae [27,28,29]. Complete transfer operons were identified in the plasmids, which is consistent with the finding that the blaNDM-36 and blaNDM-37 harboring IncHI2-type plasmids can be transferred by conjugation. In addition, the identification of highly similar IncHI2 plasmids in different E. coli STs suggested this plasmid had horizontal inter-species transfer between different E. coli clones, probably due to various antibiotic selection pressures as the plasmid contained multiple resistance genes.
It is noteworthy to mention, as shown in Table 1, most of the antibiotic susceptibility profiles of NDM-carrying clinical isolates were consistent with those of the corresponding E. coli DH5α transformants and J53 transconjugants, except for imipenem and meropenem. Compared with JNQH497, a premature stop codon was introduced into the coding region of ompD (c.238G > T p.Glu80*) in JNQH498 and JNQH462 strains. OmpD had been reported to be the main mechanism that mediated reduced susceptibility to imipenem in Enterobacter spp [30]. As such, it is speculated the premature stop codon likely accounts for the inconsistency of imipenem and meropenem susceptibility between pET28a-NDM-36 and pET28a-NDM-37 as compared to the corresponding source isolates. In addition, considering these isolates having MICs for meropenem ≤ 8 mg/L, the patient was given high-dose extended-infusion meropenem for urinary tract infection [31]. However, due to the excellent in vitro activity of ATM/AVI and the carriage of ESBL encoding genes, utility of aztreonam in combination with ceftazidime-avibactam might be one promising treatment strategy [32].
In summary, this study identified two novel NDM-type β-lactamases, NDM-36 and NDM-37, from E. coli strains isolated from a patient with refractory urinary tract infection. To the best of our knowledge, this is the first report describing two novel NDM variants detected from the same patient. This work extended our understanding of enzymatic function and demonstrated the ongoing evolution of NDM enzymes. Emergence of new NDM variants could be driven by de novo resistance evolution. A close surveillance of NDM-producing pathogens should be enacted for continued monitoring of the spread of NDM variants.
Data availability
The draft genome sequences of JNQH497, JNQH498, JNQH462 were deposited into NCBI Genome database under BioProject PRJNA702614. The complete genome sequences of JNQH497 and JNQH498 have been submitted to GenBank under the accession numbers of CP091925 ~ CP091927 and CP104384 ~ CP104385 respectively.
Code availability
Not applicable.
References
Wu W, Feng Y, Tang G, Qiao F, McNally A, Zong Z (2019) NDM metallo-beta-lactamases and their bacterial producers in health care settings. Clin Microbiol Rev 32(2):e00115-18
Feng H, Liu X, Wang S, Fleming J, Wang DC, Liu W (2017) The mechanism of NDM-1-catalyzed carbapenem hydrolysis is distinct from that of penicillin or cephalosporin hydrolysis. Nat Commun 8(1):2242
Yong D, Toleman MA, Giske CG, Cho HS, Sundman K, Lee K, Walsh TR (2009) Characterization of a new metallo-beta-lactamase gene, bla(NDM-1), and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother 53(12):5046–5054
Naas T, Oueslati S, Bonnin RA, Dabos ML, Zavala A, Dortet L, Retailleau P, Iorga BI (2017) Beta-lactamase database (BLDB)—structure and function. J Enzyme Inhib Med Chem 32(1):917–919
Wang T, Zhou Y, Zou C, Zhu Z, Zhu J, Lv J, Xie X, Chen L, Niu S, Du H (2021) Identification of a novel blaNDM Variant, blaNDM-33, in an Escherichia coli isolate from hospital wastewater in China. mSphere 6(5):e0077621
Hornsey M, Phee L, Wareham DW (2011) A novel variant, NDM-5, of the New Delhi metallo-beta-lactamase in a multidrug-resistant Escherichia coli ST648 isolate recovered from a patient in the United Kingdom. Antimicrob Agents Chemother 55(12):5952–5954
Ali A, Gupta D, Srivastava G, Sharma A, Khan AU (2019) Molecular and computational approaches to understand resistance of New Delhi metallo beta-lactamase variants (NDM-1, NDM-4, NDM-5, NDM-6, NDM-7)-producing strains against carbapenems. J Biomol Struct Dyn 37(8):2061–2071
Rehman MT, AlAjmi MF, Hussain A, Rather GM, Khan MA (2019) High-throughput virtual screening, molecular dynamics simulation, and enzyme kinetics identified ZINC84525623 as a potential inhibitor of NDM-1. Int J Mol Sci 20(4):819
Celenza G, Luzi C, Aschi M, Segatore B, Setacci D, Pellegrini C, Forcella C, Amicosante G, Perilli M (2008) Natural D240G Toho-1 mutant conferring resistance to ceftazidime: biochemical characterization of CTX-M-43. J Antimicrob Chemother 62(5):991–997
Hao M, Schuyler J, Zhang H, Shashkina E, Du H, Fouts DE, Satlin M, Kreiswirth BN, Chen L (2021) Apramycin resistance in epidemic carbapenem-resistant Klebsiella pneumoniae ST258 strains. J Antimicrob Chemother 76(8):2017–2023
Hao M, Shi X, Lv J, Niu S, Cheng S, Du H, Yu F, Tang YW, Kreiswirth BN, Zhang H, Chen L (2020) In vitro activity of apramycin against carbapenem-resistant and hypervirulent Klebsiella pneumoniae isolates. Front Microbiol 11:425
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19(5):455–477
Wick RR, Judd LM, Gorrie CL, Holt KE (2017) Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol 13(6):e1005595
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069
Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL, Jelsbak L, Sicheritz-Ponten T, Ussery DW, Aarestrup FM, Lund O (2012) Multilocus sequence typing of total-genome-sequenced bacteria. J Clin Microbiol 50(4):1355–1361
Bessonov K, Laing C, Robertson J, Yong I, Ziebell K, Gannon VPJ, Nichani A, Arya G, Nash JHE, Christianson S (2021) ECTyper: in silico Escherichia coli serotype and species prediction from raw and assembled whole-genome sequence data. Microb Genom 7(12):728
Carattoli A, Zankari E, Garcia-Fernandez A, Voldby Larsen M, Lund O, Villa L, Moller Aarestrup F, Hasman H (2014) In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 58(7):3895–3903
Grant JR, Stothard P (2008) The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res 36(Web Server issue):W181-184
Li X, Xie Y, Liu M, Tai C, Sun J, Deng Z, Ou HY (2018) oriTfinder: a web-based tool for the identification of origin of transfers in DNA sequences of bacterial mobile genetic elements. Nucleic Acids Res 46(W1):W229–W234
Katz LS, Griswold T, Morrison SS, Caravas JA, Zhang S, den Bakker HC, Deng X, Carleton HA (2019) Mashtree: a rapid comparison of whole genome sequence files. J Open Source Softw 4(44):1762
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Soding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549
Ma Z, Zeng Z, Liu J, Liu C, Pan Y, Zhang Y, Li Y (2021) Emergence of IncHI2 plasmid-harboring blaNDM-5 from porcine Escherichia coli isolates in Guangdong, China. Pathogens 10(8):954
Zhao QY, Zhu JH, Cai RM, Zheng XR, Zhang LJ, Chang MX, Lu YW, Fang LX, Sun J, Jiang HX (2021) IS26 is responsible for the evolution and transmission of blaNDM-harboring plasmids in Escherichia coli of poultry origin in China. mSystems 6(4):e0064621
Farhat N, Khan AU (2020) Evolving trends of New Delhi Metallo-betalactamse (NDM) variants: a threat to antimicrobial resistance. Infect Genet Evol 86:104588
Dadashi M, Yaslianifard S, Hajikhani B, Kabir K, Owlia P, Goudarzi M, Hakemivala M, Darban-Sarokhalil D (2019) Frequency distribution, genotypes and prevalent sequence types of New Delhi metallo-beta-lactamase-producing Escherichia coli among clinical isolates around the world: a review. J Glob Antimicrob Resist 19:284–293
Sidjabat HE, Townell N, Nimmo GR, George NM, Robson J, Vohra R, Davis L, Heney C, Paterson DL (2015) Dominance of IMP-4-producing enterobacter cloacae among carbapenemase-producing Enterobacteriaceae in Australia. Antimicrob Agents Chemother 59(7):4059–4066
Ma K, Feng Y, Zong Z (2018) Fitness cost of a mcr-1-carrying IncHI2 plasmid. PLoS ONE 13(12):e0209706
Huang H, Dong N, Shu L, Lu J, Sun Q, Chan EW, Chen S, Zhang R (2020) Colistin-resistance gene mcr in clinical carbapenem-resistant Enterobacteriaceae strains in China, 2014–2019. Emerg Microbes Infect 9(1):237–245
Lee JY, Hong YK, Lee H, Ko KS (2017) High prevalence of non-clonal imipenem-nonsusceptible Enterobacter spp. isolates in Korea and their association with porin down-regulation. Diagn Microbiol Infect Dis 87(1):53–59
Pascale R, Giannella M, Bartoletti M, Viale P, Pea F (2019) Use of meropenem in treating carbapenem-resistant Enterobacteriaceae infections. Expert Rev Anti Infect Ther 17(10):819–827
Shields RK, Doi Y (2020) Aztreonam combination therapy: an answer to metallo-β-lactamase–producing gram-negative bacteria? Oxford University Press US, pp 1099–1101
Funding
This work was supported by the Shandong Provincial Natural Science Foundation [Grant number ZR2021MH078]; Clinical & Medical Science and Technology Innovation Program of Jinan, Shandong Province [Grant number 202134040]; and Cultivate Fund from The First Affiliated Hospital of Shandong First Medical University & Shandong Provincial Qianfoshan Hospital [Grant number QYPY2022NSFC0802, 2020NSFC0808]. The above funding bodies play a role in the design of the study and collection, analysis, and interpretation of data.
Author information
Authors and Affiliations
Contributions
MH and SN: conceptualization, investigation, funding acquisition, writing—original draft. BZ enrolled the subjects. WM, WW: methodology, acquisition of data. QW, XC, YW, XD, XL, JM, FC: analysis and interpretation of data. LC, XS: review and editing. All authors agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Ethics committee approval of this study was granted by the institutional review board of the First Affiliated Hospital of Xiamen University, and informed consent from the patient was obtained.
Consent to participate
Authors had sought consent from the individual to publish the data in a journal article.
Consent for publication
All authors have contributed to the creation of this manuscript for important intellectual content and read and approved the final manuscript to be published.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Wanshan Ma, Bo Zhu, and Wen Wang contributed equally to this work. Author order was determined by contributions to the study.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, W., Zhu, B., Wang, W. et al. Genetic and enzymatic characterization of two novel blaNDM-36, -37 variants in Escherichia coli strains. Eur J Clin Microbiol Infect Dis 42, 471–480 (2023). https://doi.org/10.1007/s10096-023-04576-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-023-04576-y