Abstract
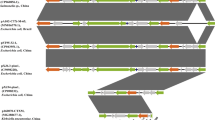
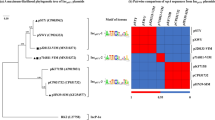
In Enterobacteriaceae, the blaOXA-48-like genes have been identified on plasmids in different regions of the world. The OXA-370 is a plasmid-encoded OXA-48-like enzyme reported in two distinct regions of Brazil. Recently, we demonstrate that the blaOXA-370 gene is disseminated among several Enterobacteriaceae species and clones, indicating a high potential for dissemination. In this work, we described for the first time the complete nucleotide sequence of six plasmids harboring the blaOXA-370 gene. Complete DNA sequencing using the Illumina platform and annotation of the plasmids showed that they belonged to incompatibility groups IncX and had in average 70 kbp. The blaOXA-370 gene is located in a composite transposon containing four genes encoding transposases, named Tn6435. In this study, highly similar plasmids were detected in different Enterobacteriaceae genera.
Similar content being viewed by others
References
Bush K, Jacoby GA (2010) Updated functional classification of β-lactamases. Antimicrob Agents Chemother 54:969–976
Sampaio JL, Ribeiro VB, Campos JC, Rozales FP, Magagnin CM, Falci DR, da Silva RC, Dalarosa MG, Luz DI, Vieira FJ, Antochevis LC, Barth AL, Zavascki AP (2014) Detection of OXA-370, an OXA-48-related class D β-lactamase, in Enterobacter hormaechei from Brazil. Antimicrob Agents Chemother 58:3566–3567
Pereira PS, Borghi M, de Araujo CF, Aires CA, Oliveira JC, Asensi MD, Carvalho-Assef AP (2015) Clonal dissemination of OXA-370-producing Klebsiella pneumoniae in Rio de Janeiro, Brazil. Antimicrob Agents Chemother 59:4453–4456
Magagnin CM, Rozales FP, Antochevis L, Nunes LS, Martins AS, Barth AL, Sampaio JM, Zavascki AP (2017) Dissemination of bla OXA-370 gene among several Enterobacteriaceae species in Brazil. Eur J Clin Microbiol Infect Dis 36:1907–1910
Maiwald M (2004) Broad-range PCR for detection and identification of bacteria. In: Persing DH (ed) Molecular microbiology diagnostic principles and practice, 2rd edn. American Society for Microbiology, Washington, DC, pp 379–390
Hoffmann H, Stindl S, Ludwig W, Stumpf A, Mehlen A, Monget D, Pierard D, Ziesing S, Heesemann J, Roggenkamp A, Schleifer KH (2005) Enterobacter hormaechei subsp. oharae subsp. nov., E. hormaechei subsp. hormaechei comb. nov., and E. hormaechei subsp. steigerwaltii subsp. nov., three new subspecies of clinical importance. J Clin Microbiol 43:3297–3303
Hoffmann H, Roggenkamp A (2003) Population genetics of the nomenspecies Enterobacter cloacae. Appl Environ Microbiol 69:5306–5318
Campos JC, da Silva MJ, dos Santos PR, Barros EM, Pereira Mde O, Seco BM, Magagnin CM, Leiroz LK, de Oliveira TG, de Faria-Junior C, Cerdeira LT, Barth AL, Sampaio SC, Zavascki AP, Poirel L, Sampaio JL (2015) Characterization of Tn3000, a transposon responsible for bla NDM-1 dissemination among Enterobacteriaceae in Brazil, Nepal, Morocco, and India. Antimicrob Agents Chemother 59:7387–7395
Birnboim HC (1983) A rapid alkaline extraction method for the isolation of plasmid DNA. Methods Enzymol 100:243–255
Casali N, Preston A(ed) (2003) E. coli plasmid vectors: methods and applications. Humana Press, Totowa
Poirel L, Walsh TR, Cuvillier V, Nordmann P (2011) Multiplex PCR for detection of acquired carbapenemase genes. Diagn Microbiol Infect Dis 70:119–123
Macrina FL, Kopecko DJ, Jones KR, Ayers DJ, McCowen SM (1978) A multiple plasmid-containing Escherichia coli strain: convenient source of size reference plasmid molecules. Plasmid 1:417–420
Clinical and Laboratory Standards Institute (2016) Performance standards for antimicrobial susceptibility testing. Document M100-S26. CLSI, Wayne, PA
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75
Carver T, Harris SR, Berriman M, Parkhill J, McQuillan JA (2012) Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 28:464–469
Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M (2006) ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res 34:D32–D36
Carattoli A, Zankari E, Garcia-Fernandez A, Voldby Larsen M, Lund O, Villa L, Moller Aarestrup F, Hasman H (2014) In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 58:3895–3903
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41
Pullinger GD, Lax AJ (1992) A Salmonella dublin virulence plasmid locus that affects bacterial growth under nutrient-limited conditions. Mol Microbiol 6:1631–1643
Hibi M, Sonoki T, Mori H (2005) Functional coupling between vanillate-O-demethylase and formaldehyde detoxification pathway. FEMS Microbiol Lett 253:237–242
Ho PL, Li Z, Lo WU, Cheung YY, Lin CH, Sham PC, Cheng VC, Ng TK, Que TL, Chow KH (2012) Identification and characterization of a novel incompatibility group X3 plasmid carrying bla NDM-1 in Enterobacteriaceae isolates with epidemiological links to multiple geographical areas in China. Emerg Microbes Infect 1:e39
Sonnevend A, Al Baloushi A, Ghazawi A, Hashmey R, Girgis S, Hamadeh MB, Al Haj M, Pal T (2013) Emergence and spread of NDM-1 producer Enterobacteriaceae with contribution of IncX3 plasmids in the United Arab Emirates. J Med Microbiol 62:1044–1050
Cerdeira LT, Cunha MPV, Francisco GR, Bueno MFC, Araujo BF, Ribas RM, Gontijo-Filho PP, Knobl T, de Oliveira GD, Lincopan N (2017) IncX3 plasmid harboring a non-Tn4401 genetic element (NTEKPC) in a hospital-associated clone of KPC-2-producing Klebsiella pneumoniae ST340/CG258. Diagn Microbiol Infect Dis 89:164–167
Shen P, Zhang Y, Li G, Jiang X (2016) Characterization of the genetic environment of the bla KPC-2 gene among Klebsiella pneumoniae isolates from a Chinese hospital. Braz J Infect Dis 20:384–388
Ho PL, Cheung YY, Lo WU, Li Z, Chow KH, Lin CH, Chan JF, Cheng VC (2013) Molecular characterization of an atypical IncX3 plasmid pKPC-NY79 carrying bla KPC-2 in a Klebsiella pneumoniae. Curr Microbiol 67:493–498
Kassis-Chikhani N, Frangeul L, Drieux L, Sengelin C, Jarlier V, Brisse S, Arlet G, Decre D (2013) Complete nucleotide sequence of the first KPC-2- and SHV-12-encoding IncX plasmid, pKpS90, from Klebsiella pneumoniae. Antimicrob Agents Chemother 57:618–620
Garcia-Fernandez A, Villa L, Carta C, Venditti C, Giordano A, Venditti M, Mancini C, Carattoli A (2012) Klebsiella pneumoniae ST258 producing KPC-3 identified in Italy carries novel plasmids and OmpK36/OmpK35 porin variants. Antimicrob Agents Chemother 56:2143–2145
Chen L, Chavda KD, Fraimow HS, Mediavilla JR, Melano RG, Jacobs MR, Bonomo RA, Kreiswirth BN (2013) Complete nucleotide sequences of bla KPC-4 and bla KPC-5 harboring IncN and IncX plasmids from Klebsiella pneumoniae strains isolated in New Jersey. Antimicrob Agents Chemother 57:269–276
Fernandes MR, McCulloch JA, Vianello MA, Moura Q, Perez-Chaparro PJ, Esposito F, Sartori L, Dropa M, Matte MH, Lira DP, Mamizuka EM, Lincopan N (2016) First report of the globally disseminated IncX4 plasmid carrying the mcr-1 gene in a colistin-resistant Escherichia coli sequence type 101 isolate from a human infection in Brazil. Antimicrob Agents Chemother 60:6415–6417
Sellera FP, Fernandes MR, Sartori L, Carvalho MP, Esposito F, Nascimento CL, Dutra GH, Mamizuka EM, Perez-Chaparro PJ, McCulloch JA, Lincopan N (2017) Escherichia coli carrying IncX4 plasmid-mediated mcr-1 and bla CTX-M genes in infected migratory Magellanic penguins (Spheniscus magellanicus). J Antimicrob Chemother 72:1255–1256
Batchelor RA, Pearson BM, Friis LM, Guerry P, Wells JM (2004) Nucleotide sequences and comparison of two large conjugative plasmids from different Campylobacter species. Microbiology 150:3507–3517
Cao TB, Saier MH Jr (2001) Conjugal type IV macromolecular transfer systems of Gram-negative bacteria: organismal distribution, structural constraints and evolutionary conclusions. Microbiology147:3201–3214
Funding
This work was supported by the Fundo de Incentivo à Pesquisa e Eventos do Hospital de Clínicas de Porto Alegre, Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul, National Council for Scientific and Technological Development (CNPq), Ministry of Science and Technology, Brazil, and Fleury Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A. P. Z. has received honoraria for speaking engagements and consultancy from Merck, AstraZeneca, Pfizer, and United Pharmaceuticals. All other authors have no conflicts of interest to declare.
Ethical approval
The study was approved by the Ethical Committee of Hospital de Clínicas de Porto Alegre (14-0046).
Informed consent
Not applicable.
Rights and permissions
About this article
Cite this article
Magagnin, C.M., Campos, J.C., da Rocha, D.A. et al. Dissemination of blaOXA-370 is mediated by IncX plasmids and the Tn6435 transposon. Eur J Clin Microbiol Infect Dis 37, 2165–2169 (2018). https://doi.org/10.1007/s10096-018-3356-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-018-3356-x