Abstract
Inhibitors of the interaction between Neuropilin-1 (NRP-1) and Vascular Endothelial Growth Factor-A165 (VEGF-A165) hold significant promise as therapeutic and diagnostic agents directed against cancers overexpressing NRP-1. In our efforts in this field, a few series of strong and fairly stable peptide-like inhibitors of the general formula Lys(Har)1-Xaa2-Xaa3-Arg4 have been previously discovered. In the current work, we focused on Lys(Har)-Dap/Dab-Pro-Arg sequence. The aim was to examine whether replacing C-terminal Arg with its homologs and mimetics would yield more stable yet still potent inhibitors. Upon considering the results of modelling and other factors, ten novel analogues with Xaa4 = homoarginine (Har), 2-amino-4-guanidino-butyric acid (Agb), 2-amino-3-guanidino-propionic acid (Agp), citrulline (Cit), 4-aminomethyl-phenylalanine [Phe(4-CH2-NH2)] were designed, synthesized and evaluated. Two of the proposed modifications resulted in inhibitors with activity slightly lower [e.g. IC50 = 14.3 μM for Lys(Har)-Dab-Pro-Har and IC50 = 19.8 μM for Lys(Har)-Dab-Pro-Phe(4-CH2-NH2)] than the parent compounds [e.g. IC50 = 4.7 μM for Lys(Har)-Dab-Pro-Arg]. What was a surprise to us, the proteolytic stability depended more on position two of the sequence than on position four. The Dab2-analogues exhibited half-life times beyond 60 h. Our results build up the knowledge on the structural requirements that effective VEGF-A165/NRP-1 inhibitors should fulfil.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Neuropilin-1 (NRP-1) is a cell surface receptor involved, inter alia, in the development of axon guidance and also in physiological, as well as pathological angiogenesis processes, including cancer (He and Tessier-Lavigne 1997; Lee et al. 2002; Staton et al. 2007). This single-pass transmembrane glycoprotein plays a crucial role in the formation of capillaries in the angiogenesis process. Its overexpression is associated with tumor aggressiveness and metastasis as observed e.g. in breast (Stephenson et al. 2002), pancreas (Parikh et al. 2003), prostate (Latil et al. 2000) or colon (Parikh et al. 2004) cancers. One of the most important ligands of NRP-1 and the main mediators of angiogenesis processes is the vascular endothelial growth factor-A165 (VEGF-A165), which acts as a proangiogenic factor that interacts with b1 and b2 of the NRP-1 subdomains. Compounds that block this interaction are potential inhibitors of the VEGF-A165/NRP-1 complex that may find application in the diagnosis and therapy of cancer (Peng et al. 2019). NRP-1 is investigated as a promising target for the targeted delivery of diagnostic or therapeutic radionuclides (Adhikari et al. 2019; Moussaron et al. 2021; Masłowska et al. 2022, 2023). Outside of the oncological field, recent data point to the possibility of using VEGF-A165/NRP-1 inhibitors as anti-SARS-CoV-2 agents (Cantuti-Castelvetri et al. 2020; Daly et al. 2020; Hu et al. 2023) or as a novel treatment strategy for neuropathic pain (Stratton et al. 2023).
The so far proposed inhibitors of the VEGF-A165/NRP-1 complex include cyclic (Jia et al. 2006, 2014; Grabowska et al. 2016, 2017) or linear (von Wronski et al. 2006; Starzec et al. 2006, 2007; Vander Kooi et al. 2007; Kamarulzaman et al. 2017; Fedorczyk et al. 2017; Tymecka et al. 2017, 2018; Puszko et al. 2019, 2020, 2023) peptides/peptidomimetics and small molecules (Jarvis et al. 2010; Novoa et al. 2010; Borriello et al. 2014; Starzec et al. 2014; Liu et al. 2014, 2018; Richard et al. 2016; Powell et al. 2018). In the group of small molecules, the significant result was the design of EG00229 that has been shown by in vivo assays to be a promising anticancer compound (Jarvis et al. 2010). Another small molecular inhibitor with demonstrated in vivo anticancer activity is NRPa-308 (Liu et al. 2018). In the peptides group, the significant achievement was the identification (by a mutated phage library screening) of heptapeptide Ala-Thr-Trp-Leu-Pro-Pro-Arg (A7R) which selectively inhibits VEGF-A165 binding to NRP-1 and decreases breast cancer angiogenesis and growth in vivo (Starzec et al. 2006, 2007).
Structure–Activity Relationship (SAR) studies around A7R showed that the shortest active fragment is its C-terminal tetrapeptide Leu-Pro-Pro-Arg. In the course of further studies, we found that substitution of Leu in the first position by Lys, and additionally the extension of the side chain of Lys by attachment of homoarginine (Har) residue, provides more active and more stable analogues. Moreover, increasing the flexibility of the middle part of molecule, in particular with simultaneous introduction of additional receptor interacting elements, at the second or third position, produced branched pentapeptides up to 30-fold more active than the A7R (Tymecka et al. 2018).
Another key SAR finding, relevant to all peptidic inhibitors of the VEGF-A165/NRP-1 interactions, is that the C-terminal arginine is crucial for binding to the NRP-1 and its lack is associated with a drastic decrease or even loss of inhibitory activity (Starzec et al. 2007; Jarvis et al. 2010). On the other hand, the metabolic stability studies of our branched pentapeptides Lys(Har)1-Xaa2-Xaa3-Arg4 have shown that among the first cleavage sites is the detachment of this key arginine (Tymecka et al. 2018; Puszko et al. 2023).
In view of the above information, we wondered whether exchange of C-terminal Arg for its homologs and mimetics in our two previously presented compounds, Lys(Har)-Dap-Pro-Arg (1) and Lys(Har)-Dab-Pro-Arg (2), would yield more stable yet still potent inhibitors. This problem was of practical interest in our works towards optimization of these analogues. It is a common knowledge that insufficient stability of peptide active substances is their major liability, significantly hampering their practical applications. Our inhibitors 1 and 2 displayed good stabilities with t½’s beyond 24 h (Tymecka et al. 2018), that nonetheless were deemed desirable to improve. In this paper we report on design, synthesis and serum metabolic stability of several C-terminally modified analogues (compounds 3–12) of our leads.
Materials and methods
Molecular modelling
Compounds contemplated to be synthesized, including analogues 3–12, were modelled in the NRP-1 binding site by manual docking followed with scoring using AutoDock Vina (Trott and Olson 2010). The basis for the modelling were the 2ORZ structure (Vander Kooi et al. 2007) from the PDB database (Yin et al. 2018) and two snapshot structures from our molecular dynamics (MD) simulations performed for compound 2 bound to NRP-1 (started also from the 2ORZ structure), described previously (Tymecka et al. 2018). The frames were representative of two binding poses (BP1 and BP2) that were significantly sampled during the MD runs of NRP-1 bound with our branched pentapeptide inhibitors.
The C-terminally modified analogues of 1 and 2 were sculpted manually (by modifying the structure of 2 from the MD snapshots) in the BP1 or BP2 binding orientation using the Discovery Studio (Biovia Discovery Studio Visualizer 2018). The manually prepared conformations were minimized (cleaned), followed by minimization of protein hydrogens. Alternatively, model fragments Ac-Pro-Xaa-OH (where Xaa stands for the contemplated C-terminal residue) were similarly prepared in the NRP-1 binding site. After the preparations, the ligands and proteins structures were converted to suitable input files using AutoDock Tools (Morris et al. 2009). The obtained complexes were scored with AutoDock Vina (Trott and Olson 2010) (using the score_only option, thus omitting the search procedure).
In all dockings, the protonation states for both ligands and the protein were set as expected at pH 7.4. The post-modelling analyses included visual inspection (Fischer et al. 2021) and comparison of the designs’ scores to those of the parent compounds. Molecular graphics were prepared in Pymol (Schrödinger LLC 2018) and Chimera (Pettersen et al. 2004).
Peptidomimetic synthesis
Unless otherwise specified, reagents and solvents were obtained from commercial suppliers and used without further purification. Fmoc-Arg(Pbf) Wang resin, Fmoc-Har(Pbf)-OH, Fmoc-Cit-OH and 4-(Boc-aminomethyl)-N-Fmoc-l-phenylalanine were purchased from Merck (Poland). 2-Chlorotrityl chloride resin, other amino acids, coupling reagents (DIC/Oxyma) and solvents were purchased from Iris Biotech (Marktredwitz, Germany). The Oxy-B and TBEC reagent were given by Luxembourg Bio Technologies LTD (Israel).
The designed compounds were synthesized by a standard 9-fluorenylmethoxycarbonyl (Fmoc) solid phase peptide synthesis methodology as previously reported (Fedorczyk et al. 2017). Preloaded Fmoc-Arg(Pbf)-Wang resin was used for the synthesis of parent compounds (1–2). The synthesis of all other peptidomimetics 3–12 was carried out on 2-Chlorotrityl chloride resin, using standard loading protocol for attachment of the first amino acid to the resin (Fmoc-Xaa-OH: DIPEA 0.6 eq./4 eq. in dry DCM). Final compounds were cleaved from the resin with the use of cocktail TFA:phenol:H2O:TIS (88:5:5:2, v/v). The crude peptidomimetics were purified by reversed phase high performance liquid chromatography (RP-HPLC), then analysed with liquid chromatography coupled with mass spectrometry (LC–MS), (Supplemental Figure SI-25 and Supplemental Table SI-3). The structure of each peptide was confirmed by high resolution mass spectrometry (HR-MS), (Supplemental Table SI-2).
Inhibition of VEGF-A165/NRP-1 binding assay
Inhibitory activity of the synthesized peptides was measured by a competitive ELISA test, following a protocol similar to the one previously described (Puszko et al. 2019). Briefly, the first step was to coat the bottom surface of a 96-well plate using 100 µl (200 ng/well) of recombinant human NRP-1 (BioLegend, San Diego, CA, USA) and incubate overnight at 4 °C. Nonspecific binding was blocked with 0.5% BSA (BioShop, Canada) in PBS (Bioshop, Canada). Then, 50 µl peptide in PBS in the selected concentrations and 50 µl (400 ng/ml) human (bt)-(bt) VEGF-A165 (Abcam, Cambridge, UK) in PBS containing 4 µg/ml heparin (Bioshop, Canada) were added. After 2 h of incubation at RT, the plate was washed and treated with ECL Streptavidin–Horseradish Peroxidase conjugate (GE Healthcare, Little Chalfont, UK) in PBS (1:100) for 45 min. After the final wash the chemiluminescence was quantified immediately after addition of 100 μl chemiluminescent substrate (SuperSignal ELISA Pico Chemiluminescent Substrate; Pierce Biotechnology, Rockford, IL, USA). Wells incubated with (bt)-VEGF-A165 served as a positive control (P), while wells not-coated with NRP-1 were used as a negative control (NS). Percentages of inhibition were calculated according to the following formula:
where S is the signal intensity obtained from wells incubated with the studied compound. Half-maximal inhibitory concentration (IC50, [µM]) values (mean ± SD of three independent experiments were determined by nonlinear regression analysis (log(inhibitor) vs. normalized response—variable slope) generated in GraphPad Prism 9.0.0 program (Graph Pad, San Diego, CA, USA).
Degradation assay in human serum
Serum stability studies of compounds 1–4 and 11–12 were performed in pooled human serum (off the clot) obtained from Innovative Research, Inc (Novi, MI, USA). Stock solutions of the peptidomimetics (10 µl, C = 168 µmol/ml) were diluted into human serum (690 µl) to give an incubation concentration of 2.4 µmol/ml of each peptidomimetic. All samples were incubated at 37 °C and 50 µl aliquots were withdrawn at selected time intervals (for 1, 3 and 11: 0 min, 4 h, 8 h, 12 h, 24 h, 30 h, 36 h, 48 h, 60 h, 72 h and 96 h; for 2, 4 and 12: 0 min, 12 h, 24 h, 36 h, 48 h, 60 h, 72 h, 84 h, 96 h, 120 h and 144 h). Then 200 µl of ACN/H2O/FA (89:9:2, v/v) was added to the samples in order to precipitate the serum proteins. The cloudy mixtures were vortexed, cooled to 4 °C and subsequently centrifuged (14,000 rpm) for 15 min in 4 °C to pellet the proteins. Every time, 150 µl of supernatant was collected, then diluted with water and freeze-dried. The freeze-dried samples were redissolved in 0.1% TFA (or FA for LC–MS analyses), then analysed by RP-HPLC or RP-HPLC/MS (for selected samples), (Supplemental Figures SI-26 to SI-31). As internal standards, H-Trp-OH and Z-Lys-OH were used. The samples were tested in four independent experiments, each performed once (3× for RP-HPLC and one for LC–MS). All experiments were conducted in sterile Eppendorf tubes and sampled with sterile tips, using high-purity organic solvents and Milli-Q quality water. Before the peptidomimetics were tested, the metabolic activity of human serum was checked first using endomorphin-2 (EM-2: Tyr-Pro-Phe-Phe-NH2), at the concentration identical to that for peptidomimetics, (Supplemental Figure SI-32). In the case of EM-2, the metabolic activity was tested in three independent experiments.
Results
Design rationale
As mentioned, in the presented studies we wanted to obtain more stable and active analogues of our branched peptidomimetic inhibitors 1 and 2 (Table 1). To this end, we considered replacements of the C-terminal arginine with its mimetics, including those in Table 1. Our designs were supported by structure-based considerations and molecular docking.
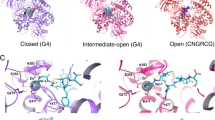
In our previous work (Tymecka et al. 2018), based on molecular dynamics simulations, we proposed an interaction model for compounds 1 and 2 bound to NRP-1. The model had the following key features: (1) the branched peptidomimetics adopt more than one binding pose (BP), with two poses (BP1 and BP2, Fig. 1) being dominant and in mutual equilibrium; (2) the C-terminal Arg residue is inserted in the shallow cleft at the NRP-1 surface and forms several interactions, including H-bonds to Asp320, Ser346, Thr349 (Fig. 1); (3) the middle and N-terminal parts of the peptidomimetic retain some residual mobility and switch between positioning BP1 and BP2; (4) they form permanent or transient H-bonds to several residues out of Gly318, Glu319, Glu324, Ser294, Tyr297, Glu348 (the exact set of interaction partners depends on the binding pose).
Two dominant binding poses (BP1 and BP2) found in molecular dynamics simulations for 1 in complex with NRP-1. The protein is depicted as an electrostatic color-coded surface (red: negative charges, white: neutral, blue: positive). Colors of the ligand are green, red and blue for carbon, oxygen and nitrogen, respectively
In the light of this model, we contemplated extending the Arg4 residue to Har. According to crystal structure 5IJR (Mota et al. 2018), Har can be accommodated in the cleft although with some displacement of the binding mode compared to if Arg is present in the cleft. Based on our modelling (Supplemental Figures SI-5 to SI-8), we assumed that interactions of Lys(Har)1 and Dap/Dab2 would not be much affected, so they should allow retaining significant affinity.
Furthermore, we speculated that it should be also possible to shorten the Arg side chain to 2-amino-4-guanidino-butyric acid (Agb) or even 2-amino-3-guanidino-propionic acid (Agp). Given that Lys(Har)1 and Dap/Dab2 side-chains are long and flexible, it was envisaged that some shortening in Xaa4 should be tolerated. In that case, the first and second residue should still be able to reach their interaction partners. Meanwhile, the shorter Xaa4 should be still able to interact with Asp320 and/or Ser346/Thr349 (Supplemental Figures SI-9 to SI-16).
A kind of an acid-test to the importance of guanidine-Asp320 interactions was provided by a Cit4-analogues. According to our modelling, Cit4 side chain should enable forming one H-bond to Asp320 (but contrary to other analogues, without charge-assistance; Supplemental Figures SI-17 to SI-20). The Cit4-analogues were furthermore quite decently scored by AutoDock Vina (Supplemental Table SI-1), so it was expected that significant affinity would be retained.
In another attempt, it was interesting to see if the C-terminal residue with an aromatic ring could gain some affinity due to partial rigidification and formation of aromatic interactions with Tyr297 (Supplemental Figures SI-21 to SI-24). This consideration led to analogues with C-terminal Phe(4-CH2-NH2).
Overall, our conclusion was that the proposed modifications do not disturb the overall binding mode to much extent (Fig. 2), so drastic affinity drops would not occur. At the same time, we believed the unnatural amino-acid in the C-terminal position could improve the metabolic stability.
Inhibitory activity
In order to verify the accuracy of the design assumptions, a modified competitive enzyme-linked immunosorbent assay (ELISA) was used to evaluate the inhibitory activity of the designed peptidomimetics against VEGF-A165/NRP-1 complex formation. In addition to the parent compounds (1 and 2), heptapeptide A7R was used for the sake of comparison. The results are summarized in Table 2.
Obtained results showed that extension of Arg4 side chain (1 and 2) by introducing one methylene group to obtain Har4 (3 and 4, respectively), gives only a slight decrease in inhibitory activity (threefold) as compared to the parent compounds. On the other hand, replacing Arg4 with Agb that has side chain shorter by one methylene group leads to compounds 5 and 6 with a 10- and 20-fold weaker inhibition of VEGF-A165 binding to NRP-1, respectively (IC50 = 87 and 106 μM). Unfortunately, further shortening of Arg4 side chain to Agp (− 2 × CH2), for 7 and 8, results in a further significant decrease in inhibitory activity (IC50 = 148 and 165 μM, respectively). Moreover, a substantial decrease in the inhibition of VEGF-A165/NRP-1 complex formation (close to that found for 7 and 8) was also observed for compounds 9 and 10 where the guanidine group of Arg was replaced by urea group of Cit (IC50 = 170 and 193 μM, respectively). Interestingly, the replacement of the alkyl side chain of Arg4 by the aromatic ring of Phe(4-CH2-NH2)4 with a simultaneous change of the guanidine to an amino group (1 and 2 vs. 11 and 12, respectively), resulted in a slight decrease in activity, comparable to that found for Har4-analogues.
Proteolytic stability in human serum
To evaluate the effect of arginine substitution with its mimetics on proteolytic stability, the most potent novel compounds (3–4 and 11–12) and the parent compounds (1–2) were incubated in human serum at 37 °C. The enzymatic degradation progress was checked for each sample at specific time intervals by RP-HPLC and the changes in the area of peptide peak were determined, where the area at time t0 was treated as 100%. The results are depicted in Fig. 3.
In general, all the tested compounds were degraded, but the process occurred at different rates depending on the structure of the compound. The peptidomimetics containing Dap residue at position 2 (compounds 1, 3 and 11) were less stable. Their estimated half-life times were t½ = 29 h (1), t½ = 26 h (3) and t½ = 19 h (11). Interestingly, the compounds containing Dab residue at the second position turned out to be more stable with half-life times of 62 or 66 h for 2 (Arg4) or 4 (Har4), respectively, and even 85 h for 12 (Phe(4-CH2-NH2)4).
To identify the formed metabolites, and therefore possible degradation pathways, the taken samples were analysed by LC–MS. In case of all compounds, we were able to find similar and/or identical metabolites suggesting three independent and parallel metabolic pathways (Fig. 4). In the first one (IA, Fig. 4), as we previously observed (Tymecka et al. 2018), proteolytic enzymes hydrolysed the N-terminal peptide bond between the Lys(Har)1 and Dap2/Dab2 residues. This led to the release of the Lys(Har) and the corresponding Dap/Dab-Pro-Xaa tripeptide should be generated. However, in none of these cases, we were able to identify the expected tripeptides. This is probably because they were rapidly degraded to single amino acids, at a rate faster than the initial study sampling time of 4 or 12 h (for 1, 3 or 11 and for 2, 4 or 12, respectively). Conversely, in the second pathway (IB, Fig. 4), the C-terminal peptide bond was cleaved to produce Lys(Har)-Dap/Dab-Pro peptide and the corresponding amino acid Xaa (Arg, Har or Phe(4-CH2-NH2)). Also for this degradation pathway, we did not observe the expected branched tetrapeptides. However, here we were able to detect the products of their further degradation (IIB, Fig. 4), i.e. Lys(Har)-Dap/Dab peptides and proline. In the case of the third pathway (IC, Fig. 4), we assume that the epsilon-peptide bond between the Lys and Har was cleaved first, since we found Har and Lys-Dap/Dab-Pro-Xaa tetrapeptides as metabolites. Moreover, the resulting tetrapeptides were further degraded to Lys-Dap/Dab-Pro and the corresponding amino acid Xaa (IIC, Fig. 4).
Probable degradation pathways and cleavage sites of 1–4 and of 11–12. The general structure of these compounds is shown with marked locations of the first major enzymatic cleavage sites (for pathway IA, IB and IC). The second position of Dap or Dab (important in view of the half-life time) is highlighted in pale grey. Peptide sequences in grey have not been identified by mass spectrometry. IIB and IIC indicate further degradation pathways for Lys(Har)-Dap/Dab-Pro and Lys-Dap/Dab-Pro-Xaa peptides, respectively
Discussion
Three major themes emerge from our results. The first one pertains to the role of the arginine residue in binding to NRP-1. C-terminal Arg is present in endogenous interaction partners for NPR-1, including VEGF-A165 or certain semaphorins. Almost all active peptide inhibitors reported so far have the C-terminal Arg. Removing this residue or modifying it has been many times shown to bring about drastic activity losses (refer to Supplemental Table SI-4 for structures and activity data relevant for the discussion below). This was the case for such A7R-analogues as des-Arg7-A7R, [Lys7]-A7R, [Ala7]-A7R or A7R-Ala8 (Starzec et al. 2007). The tetrapeptides Lys-Pro-Ala-D-Arg and Lys-Pro-Ala-Lys are significantly less potent than the Lys-Pro-Ala-Arg parent inhibitor (Jarvis et al. 2010). Furthermore, even some simple Nα-substituted arginines (e.g. carbamates like Nα-Boc-Arg-OH or Nα-Cbz-Arg-OH) exhibit measurable binding to NRP-1 (Mota et al. 2018). Arg fragment is also present in peptidomimetics designed based directly on the peptide ligands, including the EG00229 (Jarvis et al. 2010) and EG01377 (Powell et al. 2018). On the contrary, a number of small molecular inhibitors, discovered by virtual screening approaches, lack an Arg substructure (or guanidine moiety), yet still exhibit significant inhibitory potencies (Borriello et al. 2014; Starzec et al. 2014; Liu et al. 2018; Peng et al. 2020; Perez-Miller et al. 2021).
With this background, our desire was checking if Arg4 in the structures of our leads 1 and 2 could be replaced with Arg analogues (homologs and mimetics) and still allow retaining reasonable levels of inhibitory activity. The modelled binding poses of our designs suggested that these modifications should enable keeping majority of the interactions of the N-terminal and middle part of the structure, so our reasoning was that inhibitory activity drops should be minor, if any. The experimental data turned out more diversified than this expectation. The Arg4-parent compounds were not surpassed in inhibitory activity by any of the novel analogues. The shorter Agb4 or Agp4 residues gave large decreases in activity and so did the Cit4 residue lacking the charged side chain. The elongated Har4 residue and the aromatic Phe(4-CH2-NH2)4 residue turned out to be rather tolerated, with the activity decreases being moderate. The presented modifications have not been probed in the peptide inhibitors reported so far. In the peptidomimetics area (Supplemental Table SI-4), analogues close to EG00229 having (S)-Phe(4-NH-CH(NH2) = NH) or (S)-Har pieces instead of (S)-Arg fragment exhibited major activity reduction (Jarvis et al. 2010), which is in some way (though of course this is not exact comparison) different than what is seen in our peptide inhibitors.
Overall, our experimental data together with the modelled binding poses once again corroborate the importance of the interactions that take place within the Arg binding cleft on the NRP-1 surface. It seems that in the case of peptide inhibitors these intermolecular contacts are “more important” than others that may take place beyond this cleft.
In the second place, let us note that our modelling approach for the designs had several limitations. First, we utilized only the static binding poses and omitted the search element of the docking, assuming that the close analogues should adapt poses that we saw previously in molecular dynamics (Tymecka et al. 2018). This simplification was dictated by the time requirements needed to run MD simulations. On the other hand, it needs to be admitted that it could lead to incorrect description of the interactions because of the limited interaction sampling or inability to capture dynamic processes (like switching the rotameric states of the binding site), and both could affect the energetics estimations. Another limitation of the modelling approach taken is that we did not consider water molecules that are present in the binding site. As a result, our binding poses cannot capture water-mediated interactions. Furthermore, any solvation effects that could be found in MD are also lost. Relevantly to the current problem, Mota et al. showed that the water molecules play important role in the interactions with NRP-1 binding site (Mota et al. 2018). These limitations might significantly affect the docking outcomes and result in the lack of quantitative agreement between the experimental activity and the scoring results. It seems that further work on VEGF-A165/NRP-1 inhibitors could benefit from developing a design approach middle-way between full MD and static, local docking.
The third discussion topic pertains to the observed trends in the stability of our inhibitors. Our anticipation was that upon replacing the C-terminal arginine with its mimetics, we should observe significant changes in the half-life times of novel inhibitors. Surprisingly, a much more pronounced influence on the proteolytic stability was associated with the presence of either Dab or Dap in the second position of the sequence. The Dab2-analogues had t1/2 at least two times greater (t1/2 > 60 h) than their Dap2-counterparts (t1/2 around 20-30 h). On the other hand, Arg4/Har4 pairs display half-life differences of only a few hours (3–4 h). That may indicate that the introduction of an additional methylene group into the side chain at the C-terminal position (Arg vs. Har) has a negligible effect on the interaction of particular inhibitor with the active site of enzymes responsible for the hydrolysis of our compounds. Unexpectedly, in the Arg4/Phe(4-CH2-NH2)4 pairs, the stability change is also not regular however, over a much wider range, being a 10-h-loss in the case of Dap2-analogues (1 vs 11) while a more than a 20-h-gain in the case of Dab2-analogues (2 vs 12). This variability causes that stability of compounds in the Dap2 series changes as follows Arg4 > Har4 > Phe(4-CH2-NH2)4 while in the case of the Dab2 series it changes in the opposite direction, i.e. Phe(4-CH2-NH2)4 > Har4 > Arg4. This reversed order of stability surprised us, and we do not know exactly what causes it. A putative explanation could be that the active site of a certain enzyme (or enzymes), that plays the key role in the degradation pathways seen with our inhibitors, adopts its substrates in separate binding modes depending on what is present in position two. These binding modes are associated with the placement of C-terminal residue in two separate sub-pockets of the site. As a result, the stability trends in the two series are notably different. In view of the above results, further systematic studies on the structure-stability relationships of our lead structures are warranted, including an in-depth exploration of the influence of the second position.
Conclusions
In conclusion, ten novel analogues of Lys(Har)-Dab/Dap-Pro-Arg were designed, synthesized and tested. The scope of molecular modification included variation of the C-terminal residue with Arg mimetics/derivatives. None of the novel analogues was more potent than the parent compounds, but Har4 and Phe(4-CH2-NH2)4 residues turned out to be tolerated, yielding only moderate decreases in affinity. Strikingly, in the tested analogues it was position two rather than four that influenced the proteolytic stability more. The Dab2-analogues exhibited half-life times beyond 60 h. Our study builds up further SAR knowledge likely to be useful in future research. In our case, when more stable and active inhibitors are developed, we then plan to use them as targeting vector (molecule) in radioconjugates, that can be used as theranostic-like radiopharmaceuticals for the imaging and therapy of cancers that overexpress NRP-1. So far, using Lys(Har)-Dab-Pro-Arg for this purpose turned out suboptimal because of unsatisfactory nano-scale stability in human serum, especially for use as therapeutic radioagents (Masłowska et. al. 2022, 2023). Defining the border pharmacophoric requirements may be helpful in works on both ‘classic’ single-function VEGF-A165/NRP-1 inhibitors but also on the recently sought-after multifunctional/multitarget compounds that utilize the NRP-1 binding component, like dual inhibitors of NRP-1 and receptor-binding domain of Spike protein of SARS-CoV-2 (Hu et al. 2023) or dual inhibitors of NRP-1 and tubulin (Zheng et al. 2023).
Data availability
No datasets were generated or analysed during the current study.
References
Adhikari A, Tiwari AK, Shukla A et al (2019) Synthesis and preclinical evaluation of radioligand, 99m Tc-DO3A-Et-RPAR for imaging NRP-1 specific tumor. ChemistrySelect 4:12950–12954. https://doi.org/10.1002/slct.201902556
Biovia Discovery Studio Visualizer (2018) Biovia discovery studio visualizer
Borriello L, Montès M, Lepelletier Y et al (2014) Structure-based discovery of a small non-peptidic Neuropilins antagonist exerting in vitro and in vivo anti-tumor activity on breast cancer model. Cancer Lett 349:120–127. https://doi.org/10.1016/j.canlet.2014.04.004
Cantuti-Castelvetri L, Ojha R, Pedro LD et al (2020) Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science (80-) 370:856–860. https://doi.org/10.1126/science.abd2985
Daly JL, Simonetti B, Klein K et al (2020) Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science (80-) 370:861–865. https://doi.org/10.1126/science.abd3072
Fedorczyk B, Lipiński PFJ, Tymecka D et al (2017) Conformational latitude—activity relationship of KPPR tetrapeptide analogues toward their ability to inhibit binding of vascular endothelial growth factor 165 to neuropilin-1. J Pept Sci 23:445–454. https://doi.org/10.1002/psc.3009
Fischer A, Smieško M, Sellner M, Lill MA (2021) Decision making in structure-based drug discovery: visual inspection of docking results. J Med Chem 64:2489–2500. https://doi.org/10.1021/acs.jmedchem.0c02227
Grabowska K, Puszko AK, Lipiński PFJ et al (2016) Design, synthesis and in vitro biological evaluation of a small cyclic peptide as inhibitor of vascular endothelial growth factor binding to neuropilin-1. Bioorg Med Chem Lett. https://doi.org/10.1016/j.bmcl.2016.04.059
Grabowska K, Puszko AK, Lipiński PFJ et al (2017) Structure-activity relationship study of a small cyclic peptide H-c[Lys-Pro-Glu]-Arg-OH: a potent inhibitor of Vascular Endothelial Growth Factor interaction with Neuropilin-1. Bioorg Med Chem 25:597–602. https://doi.org/10.1016/j.bmc.2016.11.024
He Z, Tessier-Lavigne M (1997) Neuropilin is a receptor for the axonal chemorepellent semaphorin III. Cell 90:739–751. https://doi.org/10.1016/S0092-8674(00)80534-6
Hu C, Guo T, Zou Y et al (2023) Discovery of dual S-RBD/NRP1-targeting peptides: structure-based virtual screening, synthesis, biological evaluation, and molecular dynamics simulation studies. J Enzyme Inhib Med Chem. https://doi.org/10.1080/14756366.2023.2212327
Jarvis A, Allerston CK, Jia H et al (2010) Small molecule inhibitors of the neuropilin-1 vascular endothelial growth factor A (VEGF-A) interaction. J Med Chem 53:2215–2226. https://doi.org/10.1021/jm901755g
Jia H, Bagherzadeh A, Hartzoulakis B et al (2006) Characterization of a bicyclic peptide neuropilin-1 (NP-1) antagonist (EG3287) reveals importance of vascular endothelial growth factor exon 8 for NP-1 binding and role of NP-1 in KDR signaling. J Biol Chem 281:13493–13502. https://doi.org/10.1074/jbc.M512121200
Jia H, Aqil R, Cheng L et al (2014) N-Terminal modification of VEGF-A C terminus-derived peptides delineates structural features involved in neuropilin-1 binding and functional activity. ChemBioChem 15:1161–1170. https://doi.org/10.1002/cbic.201300658
Kamarulzaman EE, Vanderesse R, Gazzali AM et al (2017) Molecular modelling, synthesis and biological evaluation of peptide inhibitors as anti-angiogenic agent targeting neuropilin-1 for anticancer application. J Biomol Struct Dyn 35:26–45. https://doi.org/10.1080/07391102.2015.1131196
Latil A, Bièche I, Pesche S et al (2000) VEGF overexpression in clinically localized prostate tumors and neuropilin-1 overexpression in metastatic forms. Int J Cancer 89:167–171. https://doi.org/10.1002/(SICI)1097-0215(20000320)89:2%3c167::AID-IJC11%3e3.0.CO;2-9
Lee P, Goishi K, Davidson AJ et al (2002) Neuropilin-1 is required for vascular development and is a mediator of VEGF-dependent angiogenesis in zebrafish. Proc Natl Acad Sci 99:10470–10475. https://doi.org/10.1073/pnas.162366299
Liu W-Q, Megale V, Borriello L et al (2014) Synthesis and structure–activity relationship of non-peptidic antagonists of neuropilin-1 receptor. Bioorg Med Chem Lett 24:4254–4259. https://doi.org/10.1016/.2014.07.028
Liu W-Q, Lepelletier Y, Montès M et al (2018) NRPa-308, a new neuropilin-1 antagonist, exerts in vitro anti-angiogenic and anti-proliferative effects and in vivo anti-cancer effects in a mouse xenograft model. Cancer Lett 414:88–98. https://doi.org/10.1016/j.canlet.2017.10.039
Masłowska K, Witkowska E, Tymecka D et al (2022) Synthesis, physicochemical and biological study of gallium-68- and lutetium-177-labeled VEGF-A165/NRP-1 complex inhibitors based on peptide A7R and branched peptidomimetic. Pharmaceutics 14:100. https://doi.org/10.3390/pharmaceutics14010100
Masłowska K, Redkiewicz P, Halik PK et al (2023) Scandium-44 radiolabeled peptide and peptidomimetic conjugates targeting neuropilin-1 co-receptor as potential tools for cancer diagnosis and anti-angiogenic therapy. Biomedicines 11:564. https://doi.org/10.3390/biomedicines11020564
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Mota F, Fotinou C, Rana RR et al (2018) Architecture and hydration of the arginine-binding site of neuropilin-1. FEBS J 285:1290–1304. https://doi.org/10.1111/febs.14405
Moussaron A, Jouan-Hureaux V, Collet C et al (2021) Preliminary study of new gallium-68 radiolabeled peptide targeting NRP-1 to detect brain metastases by positron emission tomography. Molecules 26:7273. https://doi.org/10.3390/molecules26237273
Novoa A, Pellegrini-Moïse N, Bechet D et al (2010) Sugar-based peptidomimetics as potential inhibitors of the vascular endothelium growth factor binding to neuropilin-1. Bioorg Med Chem 18:3285–3298. https://doi.org/10.1016/j.bmc.2010.03.012
Parikh AA, Liu WB, Fan F et al (2003) Expression and regulation of the novel vascular endothelial growth factor receptor neuropilin-1 by epidermal growth factor in human pancreatic carcinoma. Cancer 98:720–729. https://doi.org/10.1002/cncr.11560
Parikh AA, Fan F, Liu WB et al (2004) Neuropilin-1 in human colon cancer. Am J Pathol 164:2139–2151. https://doi.org/10.1016/S0002-9440(10)63772-8
Peng K, Bai Y, Zhu Q et al (2019) Targeting VEGF–neuropilin interactions: a promising antitumor strategy. Drug Discov Today 24:656–664. https://doi.org/10.1016/j.drudis.2018.10.004
Peng K, Li Y, Bai Y et al (2020) Discovery of novel nonpeptide small-molecule NRP1 antagonists: Virtual screening, molecular simulation and structural modification. Bioorg Med Chem 28:115183. https://doi.org/10.1016/j.bmc.2019.115183
Perez-Miller S, Patek M, Moutal A et al (2021) Novel compounds targeting neuropilin receptor 1 with potential to interfere with SARS-CoV-2 virus entry. ACS Chem Neurosci 12:1299–1312. https://doi.org/10.1021/acschemneuro.0c00619
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera? A visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Powell J, Mota F, Steadman D et al (2018) Small molecule neuropilin-1 antagonists combine antiangiogenic and antitumor activity with immune modulation through reduction of transforming growth factor beta (TGFβ) production in regulatory T-cells. J Med Chem 61:4135–4154. https://doi.org/10.1021/acs.jmedchem.8b00210
Puszko AK, Sosnowski P, Tymecka D et al (2019) Neuropilin-1 peptide-like ligands with proline mimetics, tested using the improved chemiluminescence affinity detection method. Medchemcomm 10:332–340. https://doi.org/10.1039/C8MD00537K
Puszko AK, Sosnowski P, Raynaud F et al (2020) Does cysteine rule (CysR) complete the CendR principle? Increase in affinity of peptide ligands for NRP-1 through the presence of N-terminal cysteine. Biomolecules 10:448. https://doi.org/10.3390/biom10030448
Puszko AK, Sosnowski P, Hermine O et al (2023) Structure-activity relationship studies and biological properties evaluation of peptidic NRP-1 ligands: investigation of N-terminal cysteine importance. Bioorg Med Chem 94:117482. https://doi.org/10.1016/j.bmc.2023.117482
Richard M, Chateau A, Jelsch C et al (2016) Carbohydrate-based peptidomimetics targeting neuropilin-1: synthesis, molecular docking study and in vitro biological activities. Bioorg Med Chem 24:5315–5325. https://doi.org/10.1016/J.BMC.2016.08.052
Schrödinger LLC (2018) The PyMOL molecular graphics system. sourceforge.net/p/pymol/code/HEAD/tree/trunk/pymol
Starzec A, Vassy R, Martin A et al (2006) Antiangiogenic and antitumor activities of peptide inhibiting the vascular endothelial growth factor binding to neuropilin-1. Life Sci 79:2370–2381. https://doi.org/10.1016/j.lfs.2006.08.005
Starzec A, Ladam P, Vassy R et al (2007) Structure–function analysis of the antiangiogenic ATWLPPR peptide inhibiting VEGF165 binding to neuropilin-1 and molecular dynamics simulations of the ATWLPPR/neuropilin-1 complex. Peptides 28:2397–2402
Starzec A, Miteva MA, Ladam P et al (2014) Discovery of novel inhibitors of vascular endothelial growth factor-A–Neuropilin-1 interaction by structure-based virtual screening. Bioorg Med Chem 22:4042–4048. https://doi.org/10.1016/J.BMC.2014.05.068
Staton C, Kumar I, Reed M, Brown N (2007) Neuropilins in physiological and pathological angiogenesis. J Pathol 212:237–248. https://doi.org/10.1002/path.2182
Stephenson JM, Banerjee S, Saxena NK et al (2002) Neuropilin-1 is differentially expressed in myoepithelial cells and vascular smooth muscle cells in preneoplastic and neoplastic human breast: a possible marker for the progression of breast cancer. Int J Cancer 101:409–414. https://doi.org/10.1002/ijc.10611
Stratton HJ, Boinon L, Gomez K et al (2023) Targeting the vascular endothelial growth factor A/neuropilin 1 axis for relief of neuropathic pain. Pain 164:1473–1488. https://doi.org/10.1097/j.pain.0000000000002850
Trott O, Olson AJ (2010) AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
Tymecka D, Lipiński FB et al (2017) Structure-activity relationship study of tetrapeptide inhibitors of the Vascular Endothelial Growth Factor A binding to Neuropilin-1. Peptides 94:25–32. https://doi.org/10.1016/j.peptides.2017.06.003
Tymecka D, Puszko AK, Lipiński PFJ et al (2018) Branched pentapeptides as potent inhibitors of the vascular endothelial growth factor 165 binding to Neuropilin-1: design, synthesis and biological activity. Eur J Med Chem 158:453–462. https://doi.org/10.1016/j.ejmech.2018.08.083
Vander Kooi CW, Jusino MA, Perman B et al (2007) Structural basis for ligand and heparin binding to neuropilin B domains. Proc Natl Acad Sci USA 104:6152–6157. https://doi.org/10.1073/pnas.0700043104
von Wronski MA, Raju N, Pillai R et al (2006) Tuftsin binds neuropilin-1 through a sequence similar to that encoded by exon 8 of vascular endothelial growth factor. J Biol Chem 281:5702–5710. https://doi.org/10.1074/jbc.M511941200
Yin J, Chapman K, Clark LD et al (2018) Crystal structure of the human NK 1 tachykinin receptor. Proc Natl Acad Sci 115:13264–13269. https://doi.org/10.1073/pnas.1812717115
Zheng L, Zou Y, Xie T et al (2023) Discovery of a dual tubulin and neuropilin-1 (NRP1) inhibitor with potent in vivo anti-tumor activity via pharmacophore-based docking screening, structure optimization, and biological evaluation. J Med Chem 66:16187–16200. https://doi.org/10.1021/acs.jmedchem.3c01572
Acknowledgements
We would like to thank Luxembourg Bio Technologies LTD for donating Oxy-B and TBEC. We also thank Kacper Blaziak (CNBCh UW) for HR-MS and MS analysis.
Funding
This research was carried out within grant 2019/33/B/NZ7/02818, supported by the National Science Centre (Poland).
Author information
Authors and Affiliations
Contributions
Conceptualization: DT, PFJL, AM; methodology: DT, PR, PFJL and AM; investigation: DT (synthesis, metabolic stability), PR (NRP-1-binding assay), PFJL (molecular modelling); resources: AM; writing: DT, PFJL and AM; writing—review and editing: DT, AM. Figures 1, 2: PFJL; Figs. 3, 4 and Tables 1, 2: DT; All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical approval
The experiments in this manuscript did not involve animal or human experiments.
Additional information
Handling editor: F. Albericio.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tymecka, D., Redkiewicz, P., Lipiński, P.F.J. et al. Peptidomimetic inhibitors of the VEGF-A165/NRP-1 complex obtained by modification of the C-terminal arginine. Amino Acids 56, 49 (2024). https://doi.org/10.1007/s00726-024-03411-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00726-024-03411-8