Abstract
Severe fever with thrombocytopenia syndrome virus (SFTSV) has been reported in many countries in Southeast Asia, which expands the original geographic range of China, Korea, and Japan. Here, we report the complete genome sequences of two Thai SFTSV strains previously identified in patients with undifferentiated febrile illness in 2020. Phylogenetically, both clustered with SFTSV genotype B strains and were most closely related to those previously reported in central China (≥99.0% nucleotide sequence identity) in the L, M, and S gene segments. Nine amino acid residues encoded by one or more Thai SFTSV genomes differed from those found in global strains. Interestingly, the observed differences in numerous residues between the Thai strains suggest possible separate introductions of different variants into the region.

Similar content being viewed by others
Data availability
The nucleotide sequences of SFTSV strains in this study were deposited in the GenBank database under the accession numbers OQ341221-OQ341227.
References
Yu X-J, Liang M-F, Zhang S-Y et al (2011) Fever with thrombocytopenia associated with a novel bunyavirus in China. N Engl J Med 364:1523–1532. https://doi.org/10.1056/NEJMoa1010095
International committee on taxonomy of viruses. Taxon details: severe fever with thrombocytopenia syndrome virus. https://ictv.global/taxonomy/taxondetails?taxnode_id=202100166
Luo L-M, Zhao L, Wen H-L et al (2015) Haemaphysalis longicornis ticks as reservoir and vector of severe fever with thrombocytopenia syndrome virus in China. Emerg Infect Dis 21(10):1770–1776. https://doi.org/10.3201/eid2110.150126
Yun S-M, Lee W-G, Ryou J et al (2014) Severe fever with thrombocytopenia syndrome virus in ticks collected from humans, South Korea 2013. Emerg Infect Dis 20(8):1358–1361. https://doi.org/10.3201/eid2008.131857
Fang L-Z, Xiao X, Lei S-C et al (2023) Haemaphysalis flava ticks as a competent vector of severe fever with thrombocytopenia syndrome virus. Ticks Tick Borne Dis 14:102100. https://doi.org/10.1016/j.ttbdis.2022.102100
Niu G, Li J, Liang M et al (2013) Severe fever with thrombocytopenia syndrome virus among domesticated animals, China. Emerg Infect Dis 19(5):756–763. https://doi.org/10.3201/eid1905.120245
Ando T, Nabeshima T, Inoue S et al (2021) Severe fever with thrombocytopenia syndrome in cats and its prevalence among veterinarian staff members in Nagasaki, Japan. Viruses 13:1142. https://doi.org/10.3390/v13061142
Guu TSY, Zheng W, Tao YJ (2012) Bunyavirus: structure and replication. In: Rossmann MG, Rao VB (eds) Viral molecular machines. Springer, Boston, pp 245–266
Takahashi T, Maeda K, Suzuki T et al (2014) The first identification and retrospective study of severe fever with thrombocytopenia syndrome in Japan. J Infect Dis 209:816–827. https://doi.org/10.1093/infdis/jit603
Kim YR, Yun Y, Bae SG et al (2018) Severe fever with thrombocytopenia syndrome virus infection, South Korea. Emerg Infect Dis 24(11):2103–2105. https://doi.org/10.3201/eid2411.170756
Tran XC, Yun Y, An LV et al (2019) Endemic severe fever with thrombocytopenia syndrome, Vietnam. Emerg Infect Dis 25(5):1029–1031. https://doi.org/10.3201/eid2505.181463
Win AM, Nguyen YTH, Kim Y et al (2020) Genotypic heterogeneity of Orientia tsutsugamushi in scrub typhus patients and thrombocytopenia syndrome co-infection, Myanmar. Emerg Infect Dis 26(8):1878–1881. https://doi.org/10.3201/eid2608.200135
Peng S-H, Yang S-L, Tang S-E et al (2019) (2020) Human case of severe fever with thrombocytopenia syndrome virus infection, Taiwan. Emerg Infect Dis 26(7):1612–1614. https://doi.org/10.3201/eid2607.200104
Rattanakomol P, Khongwichit S, Linsuwanon P et al (2022) Severe fever with thrombocytopenia syndrome virus infection, Thailand, 2019–2020. Emerg Infect Dis 28(12):2572–2574. https://doi.org/10.3201/eid2812.221183
Zu Z, Lin H, Hu Y et al (2022) The genetic evolution and codon usage pattern of severe fever with thrombocytopenia syndrome virus. Infect Genet Evol 99:105238. https://doi.org/10.1016/j.meegid.2022.105238
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis Version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Huang X, Ding S, Jiang X et al (2019) Detection of SFTS virus RNA and antibodies in severe fever with thrombocytopenia syndrome surveillance cases in endemic areas of China. BMC Infect Dis 19:476. https://doi.org/10.1186/s12879-019-4068-2
Hu B, Cai K, Liu M et al (2018) Laboratory detection and molecular phylogenetic analysis of severe fever with thrombocytopenia syndrome virus in Hubei province, central China. Arch Virol 163:3243–3254. https://doi.org/10.1007/s00705-018-3993-5
Wang P, Liu L, Liu A et al (2020) Structure of severe fever with thrombocytopenia syndrome virus L protein elucidates the mechanisms of viral transcription initiation. Nat Microbiol 5:864–871. https://doi.org/10.1038/s41564-020-0712-2
Vogel D, Thorkelsson SR, Quemin ERJ et al (2020) Structural and functional characterization of the severe fever with thrombocytopenia syndrome virus L protein. Nucleic Acids Res 48:5749–5765. https://doi.org/10.1093/nar/gkaa253
Halldorsson S, Behrens A-J, Harlos K et al (2016) Structure of a phleboviral envelope glycoprotein reveals a consolidated model of membrane fusion. Proc Natl Acad Sci USA 113:7154–7159. https://doi.org/10.1073/pnas.1603827113
Acknowledgements
This work was funded by the Center of Excellence in Clinical Virology of the Faculty of Medicine of Chulalongkorn University and Hospital. PR, SK, and WC are supported by the Second Century Fund (C2F) of Chulalongkorn University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling Editor: Hideki Ebihara.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
705_2023_5897_MOESM1_ESM.pdf
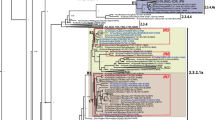
Supplementary file1 (PDF 894 KB) Supplementary Fig. S1 Additional phylogenetic analysis of the Thai SFTSV. (A) Partial M segment (549 nt). (B) Partial S segment (476 nt). The trees were generated using the maximum-likelihood method based on the Kimura 2-parameter model with 1,000 bootstrap replicates. Genotype assignments (A to F) were based on six distinct clusters with member strains identified by their GenBank accession number, country, and year of identification. Bootstrap values >70% are indicated at the branch nodes. The scale bar indicates the number of substitutions per site. Thai strains are shown with dots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rattanakomol, P., Khongwichit, S., Chuchaona, W. et al. Severe fever with thrombocytopenia syndrome virus genotype B in Thailand. Arch Virol 168, 271 (2023). https://doi.org/10.1007/s00705-023-05897-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05897-1