Abstract
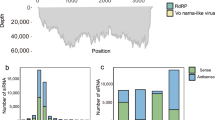
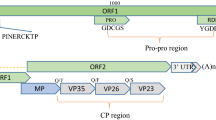
Cicada flower, Cordyceps chanhua, is a precious edible and medicinal mushroom with uses in both medicine and food in China. In this study, Cordyceps chanhua strain RCEF5995 was found to be coinfected by a previously characterized alternavirus, Cordyceps chanhua alternavirus 1 (CcAV1), and a novel victorivirus, tentatively named "Cordyceps chanhua victorivirus 1" (CcV1). Molecular characterization of CcV1 showed that its complete genome is 5,232 nucleotides long with a GC content of 57.5%. Sequence analysis indicated that CcV1 contains two overlapping open reading frames (ORFs), ORF1 and ORF2, encoding a putative coat protein (CP) of 742 amino acids (aa) and a putative RNA-dependent RNA polymerase (RdRp) of 836 aa, respectively. The termination codon of the CP ORF overlaps with the initiation codon of the RdRp ORF at the tetranucleotide sequence AUGA. Homolog searches, sequence comparisons, and phylogenetic analysis based on deduced amino acid sequences of RdRp indicated that CcV1 is a new member of the genus Victorivirus, family Totiviridae.


Similar content being viewed by others
References
Li ZZ, Chen ZA, Chen YP (2014) Chanhua, a National Treasure cordyceps in China. Hefei University of Technology Press, Hefei
Fan WW, Zgang S, Zhang YJ (2019) The complete mitochondrial genome of the Chan-hua fungus Isaria cicadae: a tale of intron evolution in Cordycipitace. Environ Microbiol 21(2):864–879
Chunyu Y, Lu Z, Luo Z, Li S, Li H, Geng Y, Xu Hong XuZ, Shi J (2019) Promotion of metabolite synthesis in Isaria cicadae, a dominant species in the cicada flower microbiota, by Cicada Pupae. J Agric Food Chem 67:8476–8484
Dong C, Li W, Li Z, Yan W, Li T, Liu X (2016) Cordyceps industry in China: current status, challenges and perspectives-Jinhu declaration for cordyceps industry development. J Mycosystema 35:1–15
Ghabrial SA, Caston JR, Jiang D, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 479–480:356–368
King AM, Adams MJ, Lefkowitz EJ et al (eds) (2011) Virus taxonomy. In: IXth report of the International Committee on Taxonomy of Viruses, 9th edn. Elsevier, Waltham
Jiang Y, Zhang T, Luo C, Jiang D, Li G, Li Q, Hsiang T, Huang J (2015) Prevalence and diversity of mycoviruses infecting the plant pathogen Ustilaginoidea virens. Virus Res 195:47–56
Li H, Havens WM, Nibert ML, Ghabrial SA (2011) RNA sequence determinants of a coupled termination-reinitiation strategy for downstream open reading frame translation in Helminthosporium victoriae virus 190S and other victoriviruses (Family Totiviridae). J Virol 85:7343–7352
Wickner RB, Ghabrial SA, Nibert ML, Patterson JL, Wang CC (2011) Family Totiviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowits EJ (eds) Virus taxonomy: ninth report of the international committee for the taxonomy of viruses. Elsevier, Academic Press, New York, pp 639–650
Zhang Y, Shi N, Wang P, Zhu Q, Yang G, Huang B (2022) Molecular characterization of a novel alternavirus infecting the entomopathogenic fungus Cordyceps chanhua. Arch Virol 167:1467–1470
Shi N, Yang G, Wang P, Wang Y, Yu D, Huang B (2019) Complete genome sequence of a novel partitivirus from the entomogenous fungus Beauveria bassiana in China. Arch Virol 164:3141–3144
Herrero N, Duenas E, Quesada-Moraga E, Zabalgogeazcoa I (2012) Prevalence and diversity of viruses in the entomopathogenic fungus Beauveria bassiana. Appl Environ Microbiol 78:8523–8530
Coutts RHA, Livieratos IC (2003) A rapid method for sequencing the 5′- and 3′-termini of double-stranded RNA viral templates using RLM-RACE. J Phytopathol 151:525–527
Kibbe WA (2007) OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res 35:W43-46
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Liu MM, Ni YX, Zhao H, Liu XT, Jia M, Liu HY, Tian BM (2022) Molecular characterization of a novel victorivirus infecting Corynespora cassiicola. Adv Virol 167:1365–1368
Funding
This work was supported by the National Natural Science Foundation of China (Grant no. 32172473) and the Anhui Natural Science Foundation (Grant no. 1908085MC56).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Stephen John Wylie.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, Q., Shi, N., Zhang, Y. et al. Complete genome sequence of a novel victorivirus infecting cicada flower (Cordyceps chanhua). Arch Virol 168, 4 (2023). https://doi.org/10.1007/s00705-022-05640-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-022-05640-2