Abstract
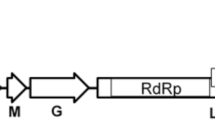
A cytorhabdovirus, tentatively named “strawberry-associated virus 1” (SaV1), was identified in strawberry (Fragaria ananassa Duch.), and its complete genome sequence was determined. Its negative-sense single-stranded RNA genome is composed of 14,159 nucleotides and contains eight open reading frames (ORFs) in the canonical order 3′-N-P-P3-M-G-P6-P7-L-5. The ORFs are separated by conserved intergenic sequences, and the genome coding region is flanked by 3ʹ and 5ʹ untranslated regions of 179 and 856 nt, respectively. SaV1 N and L genes shares 32–57% and 38–64% amino acid sequence identity with those of nine reported cytorhabdoviruses, respectively. Phylogenetic analysis showed that SaV1 clustered with high confidence with representative cytorhabdoviruses and is most closely related to tomato yellow mottle-associated virus. There are two additional small genes of unknown function between the G and L genes. We propose that SaV1 should be considered a member of a novel species in the genus Cytorhabdovirus, family Rhabdoviridae.


Similar content being viewed by others
References
Martin RR, Tzanetakis IE (2006) Characterization and recent advances in detection of strawberry viruses. Plant Dis 90:384–396
Walker PJ, Blasdell KR, Calisher CH, Dietzgen RG, Kondo H, Kurath G, Longdon B, Stone DM, Tesh RB, Tordo N, Vasilakis N, Whitfield AE (2018) ICTV virus taxonomy profile: Rhabdoviridae. J Gen Virol 99:447–448
Wang GP, Liu FC, Guo JX (1991) Identification of strawberry viruses in China. Acta Phyto Sin 21(1):9–14
Ding XL, Chen D, Wang AM, Zhang J, Wu ZJ (2017) First report of strawberry pallidosis-associated virus on strawberry in China. Plant Dis 101(12):2154
Shi Y, Han XY, Sun H, Wei Y, Wang ZY, Li HL, Chen LL, Sun BJ (2018) First report of Strawberry pallidosis associated virus Infecting Strawberry in China. Plant Dis 102(1):257
Chen D, Ding XL, Wang AM, Zhang J, Wu ZJ (2018) First report of strawberry crinivirus 3 and strawberry crinivirus 4 on strawberry in China. New Dis Rep 37:24
Xu CX, Sun XP, Taylor A, Jiao C, Xu YM, Cai XF, Wang XL, Ge CH, Pan GH, Wang QX, Fei ZJ, Wang QH (2017) Diversity, distribution, and evolution of tomato viruses in china uncovered by small RNA sequencing. J Virol 91(11):e00173-17
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359
Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ (2014) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kalyaanamoorthy S, Minh BQ, Wong TKF, Von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Minh BQ, Nguyen MA, Von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Bejerman N, Giolitti F, De Breuil S, Trucco V, Nome C, Lenardon S, Dietzgen RG (2015) Complete genome sequence and integrated protein localization and interaction map for alfalfa dwarf virus, which combines properties of both cytoplasmic and nuclear plant rhabdoviruses. Virology 483:275–283
Koloniuk I, Fránová J, Sarkisova T, Přibylová J (2018) Complete genome sequences of two divergent isolates of strawberry crinkle virus coinfecting a single strawberry plant. Arch Virol 163:2539–2542
Wetzel T, Dietzgen RG, Dale JL (1994) Genomic organization of lettuce necrotic yellows rhabdovirus. Virology 200:401–412
Yang X, Huang J, Liu C, Chen B, Zhang T, Zhou G (2017) Rice stripe mosaic virus, a novel cytorhabdovirus infecting rice via leafhopper transmission. Front Microbiol 7:2140
Acknowledgements
We thank Dr. Fangluan Gao for his assistance in the preparation of Figure 2, and Dr. Hongguang Cui (Hainan University) for review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Elvira Fiallo-Olivé.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, X., Chen, D., Du, Z. et al. The complete genome sequence of a novel cytorhabdovirus identified in strawberry (Fragaria ananassa Duch.). Arch Virol 164, 3127–3131 (2019). https://doi.org/10.1007/s00705-019-04390-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04390-y