Abstract
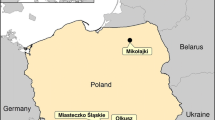
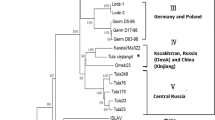
Tula virus (TULV) is a vole-associated hantavirus with low or no pathogenicity to humans. In the present study, 686 common voles (Microtus arvalis), 249 field voles (Microtus agrestis) and 30 water voles (Arvicola spec.) were collected at 79 sites in Germany, Luxembourg and France and screened by RT-PCR and TULV-IgG ELISA. TULV-specific RNA and/or antibodies were detected at 43 of the sites, demonstrating a geographically widespread distribution of the virus in the studied area. The TULV prevalence in common voles (16.7 %) was higher than that in field voles (9.2 %) and water voles (10.0 %). Time series data at ten trapping sites showed evidence of a lasting presence of TULV RNA within common vole populations for up to 34 months, although usually at low prevalence. Phylogenetic analysis demonstrated a strong genetic structuring of TULV sequences according to geography and independent of the rodent species, confirming the common vole as the preferential host, with spillover infections to co-occurring field and water voles. TULV phylogenetic clades showed a general association with evolutionary lineages in the common vole as assessed by mitochondrial DNA sequences on a large geographical scale, but with local-scale discrepancies in the contact areas.





Similar content being viewed by others
References
Plyusnin A, Elliott RM (2011) Bunyaviridae: molecular and cellular biology. Caister Academic Press, Norfolk
Henttonen H, Buchy P, Suputtamongkol Y, Jittapalapong S, Herbreteau V, Laakkonen J, Chaval Y, Galan M, Dobigny G, Charbonnel N, Michaux J, Cosson JF, Morand S, Hugot JP (2008) Recent discoveries of new hantaviruses widen their range and question their origins. Ann N Y Acad Sci 1149:84–89
Yanagihara R, Gu SH, Arai S, Kang HJ, Song JW (2014) Hantaviruses: rediscovery and new beginnings. Virus Res 187:6–14
Vaheri A, Strandin T, Hepojoki J, Sironen T, Henttonen H, Mäkelä S, Mustonen J (2013) Uncovering the mysteries of hantavirus infections. Nat Rev Microbiol 11:539–550
Heyman P, Ceianu CS, Christova I, Tordo N, Beersma M, João Alves M, Lundkvist A, Hukic M, Papa A, Tenorio A, Zelená H, Essbauer S, Visontai I, Golovljova I, Connell J, Nicoletti L, Esbroeck M Van, Gjeruldsen Dudman S, Aberle SW, Avšić-Županc T, Korukluoglu G, Nowakowska A, Klempa B, Ulrich RG, Bino S, Engler O, Opp M, Vaheri A (2011) A five-year perspective on the situation of haemorrhagic fever with renal syndrome and status of the hantavirus reservoirs in Europe, 2005–2010. Euro Surveill 16(36):pii=19961
Avsic-Zupanc T, Toney A, Anderson K, Chu YK, Schmaljohn C (1995) Genetic and antigenic properties of Dobrava virus: a unique member of the Hantavirus genus, family Bunyaviridae. J Gen Virol 76:2801–2808
Plyusnin A, Morzunov SP (2001) Virus evolution and genetic diversity of hantaviruses and their rodent hosts. Curr Top Microbiol Immunol 256:47–75
Cong M, Guo W, Wang J, Wang W, Zhou R, Li M, Zhang J, Zhang Y (2012) Genetic characteristics of hantaviruses carried by Microtus maximowixzii in Yakeshi of Inner Mongolia, China. Zhonghua Liu Xing Bing Xue Za Zhi 33:832–835
Hörling J, Chizhikov V, Lundkvist A, Jonsson M, Ivanov L, Dekonenko A, Niklasson B, Dzagurova T, Peters CJ, Tkachenko E, Nichol S (1996) Khabarovsk virus: a phylogenetically and serologically distinct hantavirus isolated from Microtus fortis trapped in far-east Russia. J Gen Virol 77:687–694
Plyusnina A, Laakkonen J, Niemimaa J, Nemirov K, Muruyeva G, Pohodiev B, Lundkvist Å, Vaheri A, Henttonen H, Vapalahti O, Plyusnin A (2008) Genetic analysis of hantaviruses carried by Myodes and Microtus rodents in Buryatia. Virol J 5:4
Zou Y, Wang JB, Gaowa HS, Yao LS, Hu G-W, Li MH, Chen HX, Plyusnin A, Shao R, Zhang YZ (2008) Isolation and genetic characterization of hantaviruses carried by Microtus voles in China. J Med Virol 80:680–688
Zou Y, Xiao QY, Dong X, Lv W, Zhang SP, Li MH, Plyusnin A, Zhang YZ (2008) Genetic analysis of hantaviruses carried by reed voles Microtus fortis in China. Virus Res 137:122–128
Plyusnin A, Vapalahti O, Lankinen H, Lehväslaiho H, Apekina N, Myasnikov Y, Kallio-Kokko H, Henttonen H, Lundkvist A, Brummer-Korvenkontio M (1994) Tula virus: a newly detected hantavirus carried by European common voles. J Virol 68:7833–7839
Sibold C, Sparr S, Schulz A, Labuda M, Kozuch O, Lysý J, Krüger DH, Meisel H (1995) Genetic characterization of a new hantavirus detected in Microtus arvalis from Slovakia. Virus Genes 10:277–281
Tkachenko EA, Witkowski PT, Radosa L, Dzagurova TK, Okulova NM, Yunicheva YV, Vasilenko L, Morozov VG, Malkin GA, Krüger DH, Klempa B (2015) Adler hantavirus, a new genetic variant of Tula virus identified in Major’s pine voles (Microtus majori) sampled in southern European Russia. Infect Genet Evol 29:156–163
Pounder KC, Begon M, Sironen T, Henttonen H, Watts PC, Voutilainen L, Vapalahti O, Klempa B, Fooks AR, McElhinney LM (2013) Novel hantavirus in field vole, United Kingdom. Emerg Infect Dis 19:673–675
Kukkonen SK, Vaheri A, Plyusnin A (1998) Completion of the Tula hantavirus genome sequence: properties of the L segment and heterogeneity found in the 3’ termini of S and L genome RNAs. J Gen Virol 79:2615–2622
Plyusnin A, Cheng Y, Vapalahti O, Pejcoch M, Unar J, Jelinkova Z, Lehväslaiho H, Lundkvist A, Vaheri A (1995) Genetic variation in Tula hantaviruses: sequence analysis of the S and M segments of strains from Central Europe. Virus Res 39:237–250
Vapalahti O, Lundkvist A, Kukkonen SK, Cheng Y, Gilljam M, Kanerva M, Manni T, Pejcoch M, Niemimaa J, Kaikusalo A, Henttonen H, Vaheri A, Plyusnin A (1996) Isolation and characterization of Tula virus, a distinct serotype in the genus Hantavirus, family Bunyaviridae. J Gen Virol 77:3063–3067
Jääskeläinen KM, Kaukinen P, Minskaya ES, Plyusnina A, Vapalahti O, Elliott RM, Weber F, Vaheri A, Plyusnin A (2007) Tula and Puumala hantavirus NSs ORFs are functional and the products inhibit activation of the interferon-beta promoter. J Med Virol 79:1527–1536
Jääskeläinen KM, Plyusnina A, Lundkvist A, Vaheri A, Plyusnin A (2008) Tula hantavirus isolate with the full-length ORF for nonstructural protein NSs survives for more consequent passages in interferon-competent cells than the isolate having truncated NSs ORF. Virol J 5:3
Mertens M, Hofmann J, Petraityte-Burneikiene R, Ziller M, Sasnauskas K, Friedrich R, Niederstrasser O, Krüger DH, Groschup MH, Petri E, Werdermann S, Ulrich RG (2011) Seroprevalence study in forestry workers of a non-endemic region in eastern Germany reveals infections by Tula and Dobrava-Belgrade hantaviruses. Med Microbiol Immunol 200:263–268
Klempa B, Meisel H, Rath S, Bartel J, Ulrich R, Kruger DH (2003) Occurrence of renal and pulmonary syndrome in a region of northeast Germany where Tula hantavirus circulates. J Clin Microbiol 41:4894–4897
Zelená H, Mrázek J, Kuhn T (2013) Tula hantavirus infection in immunocompromised host, Czech Republic. Emerg Infect Dis 19:1873–1876
Klempa B, Radosa L, Kruger DH (2013) The broad spectrum of hantaviruses and their hosts in Central Europe. Acta Virol 57:130–137
Michalski A, Niemcewicz M, Bielawska-Drózd A, Nowakowska A, Gaweł J, Pitucha G, Joniec J, Zielonka K, Marciniak-Niemcewicz A, Kocik J (2014) Surveillance of hantaviruses in Poland: a study of animal reservoirs and human hantavirus disease in subcarpathia. Vector Borne Zoonotic Dis 14:514–522
Nikolic V, Stajkovic N, Stamenkovic G, Cekanac R, Marusic P, Siljic M, Gligic A, Stanojevic M (2014) Evidence of recombination in Tula virus strains from Serbia. Infect Genet Evol 21:472–478
Olsson GE, Leirs H, Henttonen H (2010) Hantaviruses and their hosts in Europe: reservoirs here and there, but not everywhere? Vector-Borne Zoonotic Dis 10:549–561
Scharninghausen JJ, Pfeffer M, Meyer H, Davis DS, Honeycutt RL, Faulde M (2002) Genetic evidence for Tula virus in Microtus arvalis and Microtus agrestis populations in Croatia. Vector Borne Zoonotic Dis 2:19–27
Schmidt S, Essbauer SS, Mayer-Scholl A, Poppert S, Schmidt-Chanasit J, Klempa B, Henning K, Schares G, Groschup MH, Spitzenberger F, Richter D, Heckel G, Ulrich RG (2014) Multiple infections of rodents with zoonotic pathogens in Austria. Vector Borne Zoonotic Dis 14:467–475
Iakimenko VV, Dekonenko AE, Mal’kova MG, Kuz’min IV, Tantsev AK, Dzagurova TK, Tkachenko EA (2000) The spread of hantaviruses in western Siberia. Med Parazitol (Mosk) 3:21–28
Korva M, Duh D, Puterle A, Trilar T, Županc TA (2009) First molecular evidence of Tula hantavirus in Microtus voles in Slovenia. Virus Res 144:318–322
Song JW, Gligic A, Yanagihara R (2002) Identification of Tula hantavirus in Pitymys subterraneus captured in the Cacak region of Serbia-Yugoslavia. Int J Infect Dis 6:31–36
Schmidt-Chanasit J, Essbauer S, Petraityte R, Yoshimatsu K, Tackmann K, Conraths FJ, Sasnauskas K, Arikawa J, Thomas A, Pfeffer M, Scharninghausen JJ, Splettstoesser W, Wenk M, Heckel G, Ulrich RG (2010) Extensive host sharing of central European Tula virus. J Virol 84:459–474
Schlegel M, Kindler E, Essbauer SS, Wolf R, Thiel J, Groschup MH, Heckel G, Oehme RM, Ulrich RG (2012) Tula virus infections in the Eurasian water vole in central Europe. Vector Borne Zoonotic Dis 12:503–513
Guo WP, Lin XD, Wang W, Tian JH, Cong ML, Zhang HL, Wang MR, Zhou RH, Wang JB, Li MH, Xu J, Holmes EC, Zhang YZ (2013) Phylogeny and origins of hantaviruses harbored by bats, insectivores, and rodents. PLoS Pathogens 9:e1003159
Nemirov K, Leirs H, Lundkvist A, Olsson GE (2010) Puumala hantavirus and Myodes glareolus in northern Europe: no evidence of co-divergence between genetic lineages of virus and host. J Gen Virol 91:1262–1274
Ramsden C, Melo FL, Figueiredo LM, Holmes EC, Zanotto PMA, VGDN Consortium (2008) High rates of molecular evolution in hantaviruses. Mol Biol Evol 25:1488–1492
Fink S, Excoffier L, Heckel G (2004) Mitochondrial gene diversity in the common vole Microtus arvalis shaped by historical divergence and local adaptations. Mol Ecol 13:3501–3514
Heckel G, Burri R, Fink S, Desmet JF, Excoffier L (2005) Genetic structure and colonization processes in European populations of the common vole, Microtus arvalis. Evolution 59:2231–2242
Lischer HEL, Excoffier L, Heckel G (2014) Ignoring heterozygous sites biases phylogenomic estimates of divergence times: implications for the evolutionary history of Microtus voles. Mol Biol Evol 31:817–831
Drexler JF, Corman VM, Müller MA, Maganga GD, Vallo P, Binger T, Gloza-Rausch F, Rasche A, Yordanov S, Seebens A, Oppong S, Sarkodie YA, Pongombo C, Lukashev AN, Schmidt-Chanasit J, Stöcker A, Carneiro AJB, Erbar S, Maisner A, Fronhoffs F, Buettner R, Kalko EKV, Kruppa T, Franke CR, Kallies R, Yandoko ERN, Herrler G, Reusken C, Hassanin A, Krüger DH, Matthee S, Ulrich RG, Leroy EM, Drosten C (2012) Bats host major mammalian paramyxoviruses. Nat Commun 3:796
Drexler JF, Corman VM, Müller MA, Lukashev AN, Gmyl A, Coutard B, Adam A, Ritz D, Leijten LM, van Riel D, Kallies R, Klose SM, Gloza-Rausch F, Binger T, Annan A, Adu-Sarkodie Y, Oppong S, Bourgarel M, Rupp D, Hoffmann B, Schlegel M, Kümmerer BM, Krüger DH, Schmidt-Chanasit J, Setién AA, Cottontail VM, Hemachudha T, Wacharapluesadee S, Osterrieder K, Bartenschlager R, Matthee S, Beer M, Kuiken T, Reusken C, Leroy EM, Ulrich RG, Drosten C (2013) Evidence for Novel Hepaciviruses in Rodents. PLoS Pathogens 9:e1003438
Guenther S, Grobbel M, Heidemanns K, Schlegel M, Ulrich RG, Ewers C, Wieler LH (2010) First insights into antimicrobial resistance among faecal Escherichia coli isolates from small wild mammals in rural areas. Sci Total Environ 408:3519–3522
Mayer-Scholl A, Hammerl J, Schmidt S, Ulrich R, Pfeffer M, Woll D, Scholz H, Thomas A, Nöckler K (2014) Leptospira spp. in Rodents and Shrews in Germany. Int J Environ Res Public Health 11:7562–7574
Ulrich RG, Schmidt-Chanasit J, Schlegel M, Jacob J, Pelz H-J, Mertens M, Wenk M, Büchner T, Masur D, Sevke K, Groschup MH, Gerstengarbe F-W, Pfeffer M, Oehme R, Wegener W, Bemmann M, Ohlmeyer L, Wolf R, Zoller H, Koch J, Brockmann S, Heckel G, Essbauer SS (2008) Network “Rodent-borne pathogens” in Germany: longitudinal studies on the geographical distribution and prevalence of hantavirus infections. Parasitol Res 103:121–129
Schlegel M, Ali HS, Stieger N, Groschup MH, Wolf R, Ulrich RG (2012) Molecular identification of small mammal species using novel cytochrome b gene-derived degenerated primers. Biochem Genet 50:440–447
Aasen E, Medrano JF (1990) Amplification of the ZFY and ZFX genes for sex identification in humans, cattle, sheep and goats. Biotechnology (N Y) 8:1279–1281
Bryja J, Konecny A (2003) Fast sex identification in wild mammals using PCR amplification of the Sry gene. Folia Zoologica 52:269–274
Schlegel M, Sheikh Ali H, Keller M, Groschup M, Ulrich R (2013) Molekulare Artbestimmung von Kleinsäugern. LabLoeffler 7:16–17
Essbauer SS, Schmidt J, Conraths FJ, Friedrich R, Koch J, Hautmann W, Pfeffer M, Wölfel R, Finke J, Dobler G, Ulrich R (2006) A new Puumala hantavirus subtype in rodents associated with an outbreak of Nephropathia epidemica in South-East Germany in 2004. Epidemiol Infect 134:1333–1344
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symp Ser 41:95–98
Braaker S, Heckel G (2009) Transalpine colonisation and partial phylogeographic erosion by dispersal in the common vole Microtus arvalis. Mol Ecol 18:2518–2531
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biol 61:539–542
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Gateway Computing Environments Workshop (GCE), 2010, pp 1–8
Rambaut A (2012) FigTree v1. 4. Molecular evolution, phylogenetics and epidemiology. University of Edinburgh, Institute of Evolutionary Biology, Edinburgh
Song JW, Baek LJ, Song KJ, Skrok A, Markowski J, Bratosiewicz-Wasik J, Kordek R, Liberski PP, Yanagihara R (2004) Characterization of Tula virus from common voles (Microtus arvalis) in Poland: evidence for geographic-specific phylogenetic clustering. Virus Genes 29:239–247
Plyusnin A, Beaty BJ, Elliott RM, Goldbach R, Kormelink R, Lundkvist A, Schmaljohn CS, Tesh RB (2011) Bunyaviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ (eds) Virus taxonomy: ninth report of the international committee on taxonomy of viruses. Elsevier Inc., San Diego, pp 725–741
Jacob J, Manson P, Barfknecht R, Fredricks T (2014) Common vole (Microtus arvalis) ecology and management: implications for risk assessment of plant protection products. Pest Manag Sci 70:869–878
Yletyinen S, Norrdahl K (2008) Habitat use of field voles (Microtus agrestis) in wide and narrow buffer zones. Agr Ecosyst Environ 123:194–200
Myllymäki A (1977) Interactions between the field vole Microtus agrestis and its microtine competitors in Central-Scandinavian populations. Oikos 29:570–580
Jakab F, Horváth G, Ferenczi E, Sebők J, Szűcs G (2008) First detection of Tula hantaviruses in Microtus arvalis voles in Hungary. Arch Virol 153:2093–2096
Essbauer SS, Schmidt-Chanasit J, Madeja EL, Wegener W, Friedrich R, Petraityte R, Sasnauskas K, Jacob J, Koch J, Dobler G, Conraths FJ, Pfeffer M, Pitra C, Ulrich RG (2007) Nephropathia epidemica in metropolitan area, Germany. Emerg Infect Dis 13:1271–1273
Weber de Melo V, Sheikh Ali H, Freise J, Kühnert D, Essbauer S, Mertens M, Wanka KM, Drewes S, Ulrich RG, Heckel G (2015) Spatiotemporal dynamics of Puumala hantavirus associated with its rodent host, Myodes glareolus. Evol Appl 8:545–559
Engelthaler DM, Levy CE, Fink TM, Tanda D, Davis T (1998) Short report: Decrease in seroprevalence of antibodies to hantavirus in rodents from 1993–1994 hantavirus pulmonary syndrome case sites. Am J Trop Med Hyg 58:737–738
Garanina SB, Platonov AE, Zhuravlev VI, Murashkina AN, Yakimenko VV, Korneev AG, Shipulin GA (2009) Genetic diversity and geographic distribution of hantaviruses in Russia. Zoonoses Public Health 56:297–309
Klein TA, Kang HJ, Gu SH, Moon S, Shim SH, Park YM, Lee SY, Kim HC, Chong ST, O’Guinn M, Lee JS, Turell MJ, Song JW (2011) Hantaan virus surveillance targeting small mammals at Dagmar North Training Area, Gyeonggi Province, Republic of Korea, 2001–2005. J Vector Ecol 36:373–381
Tadin A, Turk N, Korva M, Margaletić J, Beck R, Vucelja M, Habuš J, Svoboda P, Županc TA, Henttonen H, Markotić A (2012) Multiple co-infections of rodents with hantaviruses, Leptospira, and Babesia in Croatia. Vector Borne Zoonotic Dis 12:388–392
Yanagihara R, Daum CA, Lee PW, Baek LJ, Amyx HL, Gajdusek DC, Gibbs CJ Jr (1987) Serological survey of Prospect Hill virus infection in indigenous wild rodents in the USA. Trans R Soc Trop Med Hyg 81:42–45
Deter J, Chaval Y, Galan M, Gauffre B, Morand S, Henttonen H, Laakkonen J, Voutilainen L, Charbonnel N, Cosson JF (2008) Kinship, dispersal and hantavirus transmission in bank and common voles. Arch Virol 153:435–444
Handke W, Oelschlegel R, Franke R, Krüger DH, Rang A (2009) Hantaan virus triggers TLR3-dependent innate immune responses. J Immunol 182:2849–2858
Alexeyev OA, Ahlm C, Elgh F, Aava B, Palo T, Settergren B, Tärnvik A, Wadell G, Juto P (1998) A minority of seropositive wild bank voles (Clethrionomys glareolus) show evidence of current Puumala virus infection. Epidemiol Infect 121:419–425
Kuenzi AJ, Douglass RJ, Bond CW, Calisher CH, Mills JN (2005) Long-term dynamics of Sin Nombre viral RNA and antibody in deer mice in Montana. J Wildl Dis 41:473–481
Meyer BJ, Schmaljohn CS (2000) Persistent hantavirus infections: characteristics and mechanisms. Trends Microbiol 8:61–67
Beysard M, Heckel G (2014) Structure and dynamics of hybrid zones at different stages of speciation in the common vole (Microtus arvalis). Mol Ecol 23:673–687
Fischer MC, Foll M, Heckel G, Excoffier L (2014) Continental-scale footprint of balancing and positive selection in a small rodent (Microtus arvalis). PLoS One 9:e112332
Martinkova M, Barnett R, Cucchi T, Struchen R, Pascal M, Pascal M, Fischer MC, Higham T, Brace S, Ho SYW, Quere JP, O′Higgins J, Excoffier L, Heckel G, Hoelzel AR, Dobney KM, Searle JB (2013) Divergent evolutionary processes associated with colonization of offshore islands. Mol Ecol 22:5205–5220
Beysard M, Krebs-Wheaton R, Heckel G (2015) Tracing reinforcement through asymmetrical partner preference in the European common vole Microtus arvalis. BMC Evol Biol 15:170
Schweizer M, Excoffier L, Heckel G (2007) Fine-scale genetic structure and dispersal patterns in the common vole Microtus arvalis. Mol Ecol 16:2463–2473
Acknowledgments
The work in the laboratory of RGU was supported by the Robert Koch-Institut with funds from the German Ministry of Public Health (grant no. 1362/1-924 and 1362/1-980 to RGU), Deutsche Forschungsgemeinschaft (SPP 1596 “Ecology and Species Barriers in Emerging Viral Diseases”, UL 405/1-1 to RGU), and the EU grants APHAEA (grant no. 2811ERA117) and FP7-261504 EDENext and is catalogued by the EDENext Steering Committee as EDENext368 (http://www.edenext.eu). The contents of this publication are the sole responsibility of the authors and do not necessarily reflect the views of the European Commission. Collection of samples in Jeeser, Mecklenburg-Western Pomerania, were done in the frame of the UFOPLAN project 3709 41 401 granted to Jens Jacob. Moritz Saxenhofer was supported by grant 31003A-149585 from the Swiss National Science Foundation to Gerald Heckel. The technical assistance of Sybille Herzog, Jorge A. Encarnação, Séverine Murri, Daniel Nobach, Markus Eickmann, Manon Bourg, Kathrin Baumann, Samuel Bernstein, Thomas Büchner, Angele Breithaupt, Fabian Deutskens, Paul Dremsek, Ulrike Duve, Sylvia Ferguson, Henrike Gregersen, Bärbel Hammerschmidt, Dörte Kaufmann, Robert Klopfleisch, Nastasja G. Kratzmann, Christian Kretzschmar, Jens Lewitzki, Christina Maresch, Marc Mertens, Joachim Meyer, Ulrike M. Rosenfeld, Josephine Schlosser, Julia Schneider, Josephine Schröder, Kati Sevke, Hanan Sheikh Ali, Jens P. Teifke, Franziska Thomas, Ute Wessels, Daniel Windolph, and Theres Wollny in rodent dissection and the collection and provision of rodents by Sabine Atger, Laurent Crespin, Anja Globig, Torsten Heidecke, Dietrich Heidecke, Anika Hellemann, Christian Imholt, Martin Kaatz, Joachim Kramer, Thilo Liesenjohann, Paul-Walter Löhr, Jennifer Maiano, Sébastien Masseglia, Lutz Ohlmeyer, Angélique Pion, Daniela Reil, Dirk Steinhauer, Alexandre Teynier, Jörg Thiel, Cornelia Triebenbacher, Elise Vaumourin, Matthias Wenk and the RIFCON Field Study Team is kindly acknowledged. The authors would like to thank Martin H. Groschup and Noel Tordo for support, and Nicole Neumann for generation of Figures 1 and 4.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Schmidt, S., Saxenhofer, M., Drewes, S. et al. High genetic structuring of Tula hantavirus. Arch Virol 161, 1135–1149 (2016). https://doi.org/10.1007/s00705-016-2762-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-016-2762-6