Abstract
H6 influenza viruses are prevalent in domestic and wild birds in Eurasian countries and have been isolated from pigs and a human. To prepare for an influenza pandemic, we have established an influenza virus library consisting of more than 1,300 influenza virus strains, including 144 combinations of 16 hemagglutinin and 9 neuraminidase subtypes. H6 viruses in the library were classified into Early, Group II, Group III, and W312 sublineages and the North America lineage on the basis of their phylogenetic features. Chicken antisera to A/duck/Hong Kong/960/1980 (H6N2) of the Early sublineage broadly reacted with viruses of different sublineages in a hemagglutinin inhibition test. A whole inactivated virus particle vaccine was prepared from A/duck/Hong Kong/960/1980 (H6N2) which was stocked in the influenza virus library. The potency of this vaccine against A/duck/Vietnam/OIE-0033/2012 (H6N2), which belongs to a different sublineage, was evaluated in mice. The test vaccine was sufficiently potent to induce an immune response that reduced the impact of disease caused by a challenge with A/duck/Vietnam/OIE-0033/2012 (H6N2) in mice. The present results indicate that the whole inactivated virus particle vaccine prepared from a virus strain in the influenza virus library is useful as a vaccine against pandemic influenza.
Similar content being viewed by others
Introduction
Influenza A viruses infect a large variety of birds and mammals, including humans. In humans, H1 and H3 viruses cause seasonal influenza. Since neutralizing (NT) antibodies to the hemagglutinin (HA) or neuraminidase (NA) are subtype-specific, the human population is immunologically naïve to infection with viruses of novel subtypes. Influenza viruses of each of the HA subtypes (H1–H13) replicate in the respiratory tract of pigs [13]. In addition, avian influenza viruses acquire the potential to infect humans after multiple infections in pigs [25]. Cases of human infection with H1, H2, H3, H5, H6, H7, H9, and H10 influenza viruses have been reported [1, 3, 4, 21, 24, 30]. Therefore, it is recommended to prepare vaccines against viruses of each of the HA subtype for preparedness for future pandemics.
To provide information on the precursor genes of future pandemic influenza viruses, global surveillance of avian influenza has been conducted since 1977 in Japan, Alaska, Siberia, Mongolia, Vietnam and Australia [9, 15, 18, 20, 23, 29]. Virus isolates in the surveillance have been stocked in the influenza virus library in our laboratory for vaccine and diagnostic use. Their biological characteristics have been analyzed, and the data are available at http://virusdb.czc.hokudai.ac.jp/. In our previous study, vaccines against infection with H1, H5, H7, and H9 influenza viruses were prepared from viruses in the library, and they induced effective immunity against challenge viruses in mice and cynomolgus macaques [10, 11, 17, 19, 27]. It was also revealed that inactivated whole-particle virus vaccines were more effective than ether-split vaccines [6, 19].
H6 influenza viruses are prevalent in poultry populations in Eurasian countries [8, 14]. The HA genes of Eurasian H6 viruses have been phylogenetically divided into Early, Europe, Group I, Group II, Group III, and W312 sublineages [8]. H6 viruses were also isolated from domestic pigs in southern China [31]. In May 2013, an H6N1 virus was isolated from a 20-year-old woman with symptoms including fever, cough, headache, and muscle ache in Taiwan [30]. In the present study, to prepare for a potential influenza pandemic, H6 viruses from the influenza virus library were genetically and antigenically analyzed to select a vaccine strain. The efficacy of an inactivated whole-virus-particle vaccine prepared from this strain was evaluated in mice.
Materials and methods
Viruses and cells
A/duck/Hong Kong/960/1980 (H6N2) (Dk/Hong Kong/960) and A/teal/Hong Kong/W312/1997 (H6N1) were kindly provided by Dr. Shortridge K. F., and Dr. Guan Y., University of Hong Kong, Hong Kong SAR, China, respectively. A/duck/Vietnam/OIE-0033/2012 (H6N2) (Dk/Vietnam/2012) was isolated from a domestic duck in our surveillance study in Vietnam [18]. The HA sequence data of the representative viruses of each lineage, Dk/Hong Kong/960, Dk/Vietnam/2012, A/teal/Hong Kong/W312/1997 (H6N1), A/shearwater/S. Australia/1/1972 (H6N5), and A/turkey/Massachusetts/3740/1965 (H6N2), have been deposited at GenBank/EMBL/DDBJ (accession no. AB294213, AB727927, AF250479, AB278600, and AB296072). All viruses used in the present study were propagated in 10-day-old embryonated chicken eggs at 35 °C for 48 h, and infectious allantoic fluids were stored at −80 °C until use.
Madin-Darby canine kidney (MDCK) cells were grown in MEM (Nissui Pharmaceutical, Tokyo, Japan) supplemented with calf serum and used to titrate virus infectivity by plaque assay [25].
Sequencing and phylogenetic analysis
Viral RNA was extracted from the allantoic fluids of embryonated chicken eggs using TRIzol LS Reagent (Life Technologies, Carlsbad, California, U.S.A.) and reverse-transcribed with the Uni 12 primer (5’-AGCAAAAGCAGG-3’) [26] and M-MLV reverse transcriptase (Life Technologies). The full-length cDNAs of the eight gene segments were amplified by polymerase chain reaction with Ex-Taq (TaKaRa, Shiga, Japan) and gene-specific primer sets [7]. Direct sequencing of each gene segment was performed using a 3500 Genetic Analyzer (Life Technologies). For phylogenetic analysis, sequence data of the genes together with those from the public database were analyzed by the maximum-likelihood method using MEGA 5.0 software (http://www.megasoftware.net/).
Serological tests
Hemagglutinin inhibition (HI) and NT tests were performed basically according to Sakabe et al. [22]. The HI titer was expressed as the reciprocal of the highest serum dilution showing complete inhibition of hemagglutination when using 8 HA units of the virus. In NT tests, titers were determined as the reciprocal of the serum dilution at which complete inhibition of the cytopathic effect was achieved in MDCK cells when using 100 times the 50 % tissue culture infectious dose (TCID50) of virus.
Virus growth in embryonated chicken eggs
One hundred times the 50 % egg infectious dose (EID50) of the virus was used to inoculate 10-day-old embryonated chicken eggs, which were then incubated at 35 °C. Allantoic fluids were collected at 12, 24, 36, 48, 60, and 72 h after inoculation to determine virus titers by plaque assay in MDCK cells.
Vaccine preparation
Dk/Hong Kong/960 and Dk/Vietnam/2012 were inoculated into the allantoic cavities of 10-day-old embryonated chicken eggs and propagated at 35 °C for 48 h. The viruses in the allantoic fluids were purified by differential centrifugation and sedimentation through a sucrose gradient [12]. The purified virus was inactivated with 0.2 % formalin at 4 °C for 7 days. The protein concentration was measured using BCA Protein Assay Reagent (Thermo Fisher Scientific, Waltham, Massachusetts, U.S.A.).
Potency test of vaccines in mice
Each of the whole inactivated vaccines of Dk/Hong Kong/960 (2, 10, or 50 µg of protein) or Dk/Vietnam/2012 (2 or 10 µg of protein) was injected once intraperitoneally into ten 4-week-old female BALB/c mice (Japan SLC, Shizuoka, Japan). PBS was injected into the control mice. Three weeks later, serum samples were collected and mice were challenged with 106.0 PFU/40 µl of Dk/Vietnam/2012 intranasally under anesthesia. At 3 days post-challenge, five mice of each group were sacrificed, and their lungs were collected. Virus titers in the lung homogenates were quantified by plaque assay in MDCK cells. Five other mice of each group were observed for weight loss for 14 days. Each of the inactivated vaccines of Dk/Hong Kong/960 (2, 10 or 50 µg of protein) was also injected twice into mice with a 2-week interval. Two weeks after the second vaccination, serum samples were collected, and mice were challenged with 106.0 PFU/40 µl of Dk/Vietnam/2012. The statistical significance of weight loss and virus titers in the lungs of mice was calculated by Student’s t-test. Animal experiments were authorized by the Institutional Animal Care and Use Committee of the Graduate School of Veterinary Medicine, Hokkaido University (approval numbers: 13-0104), and all experiments were performed according to the guidelines of this committee.
Results
Genetic analysis of H6 influenza viruses
Nucleotide sequences of the HA genes of the 94 H6 viruses in the influenza virus library were phylogenetically analyzed by the maximum-likelihood method with other representative H6 strains of each lineage. On the basis of the results of phylogenetic analysis, 10 H6 HA genes were classified into the Early sublineage, 58 into Group II, 21 into Group III, one into the W312 sublineage, and one into the North America lineage. Three H6 viruses, including A/shearwater/S. Australia/1/1972 (H6N5), were classified into the Eurasian lineage but not classified into any sublineage. A phylogenetic tree of 20 representative strains of the library is shown in Fig. 1, and these strains are indicated in bold. The H6 viruses isolated from poultry in Hong Kong in 1980 were classified into the Early sublineage. The H6 viruses isolated from poultry in Asia in the past decade were classified into Group II, and the isolates from wild birds in the East Asia and poultry in Taiwan were classified into Group III.
Phylogenetic tree of H6 HA genes of influenza viruses. Nucleotides 123–857 (735 bp) of the HA gene were used for maximum-likelihood phylogenetic analysis. Horizontal distances are proportional to the minimum number of nucleotide differences required to join nodes and sequences. Numbers at each node indicate the confidence level in bootstrap analysis with 1,000 replications. The representative viruses of each sublineage are underlined. The viruses in the influenza virus library are presented in bold. The vaccine and challenge strains are highlighted
Antigenic analysis of H6 influenza viruses
From 94 H6 influenza virus strains in the library, 18 genetically distinct strains were selected from each lineage and antigenically analyzed using the HI test (Table 1). H6 viruses of the Group II sublineage displayed low reactivity with antisera against H6 viruses of other sublineages. Antiserum against Dk/Hong Kong/960 of the Early sublineage reacted with H6 viruses of all sublineages. These results indicate that the antigenicity of H6 viruses of Group II is different from that of other H6 viruses, and the vaccine strain for H6 virus should be selected from the viruses of the Early sublineage.
Selection of a vaccine strain from H6 influenza viruses in the influenza virus library
To select a vaccine strain, the growth in embryonated chicken eggs of six H6 viruses of the Early sublineage, A/duck/Hong Kong/834/1980 (H6N1), A/duck/Hong Kong/882/1980 (H6N2) (Dk/Hong Kong/882), A/duck/Hong Kong/926/1980 (H6N2), A/duck/Hong Kong/941/1980 (H6N1), A/duck/Hong Kong/943/1980 (H6N5), and Dk/Hong Kong/960 was assessed. Dk/Hong Kong/882 and Dk/Hong Kong/960 replicated most effectively in chicken embryos. The highest virus titers for 72 h were 107.4 and 107.1 PFU/ml, respectively. Their replication had reached a plateau by 36 and 24 h, respectively (data not shown). Following this, protein yields after virus purification were determined. Dk/Hong Kong/882 was purified from 30 embryonated chicken eggs, and 650 μg of purified virus was obtained. Dk/Hong Kong/960 was purified from the same number of embryonated chicken eggs, and 990 μg of purified virus was obtained, suggesting that Dk/Hong Kong/960 exhibited a higher protein yield after virus purification than Dk/Hong Kong/882. Therefore, Dk/Hong Kong/960 was selected as a vaccine strain.
Potency test of the vaccine against an H6 influenza virus strain in mice
The inactivated vaccine of Dk/Hong Kong/960 or Dk/Vietnam/2012 was intraperitoneally injected once into mice. The serum antibody titers of mice against the vaccine and the challenge strain were examined (Table 2). The NT antibodies to the homologous vaccine strain were detected in the sera of all mice vaccinated with 10 or 50 μg of protein of the Dk/Hong Kong/960 vaccine and 2 or 10 μg of protein of the Dk/Vietnam/2012 vaccine. The NT antibodies to Dk/Vietnam/2012 were also detected in the sera of more than half of the mice vaccinated with 10 or 50 μg of protein of the Dk/Hong Kong/960 vaccine.
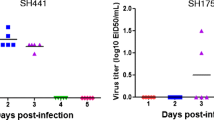
Further, to assess the potency of the vaccine against a challenge with Dk/Vietnam/2012, 106.0 PFU of the virus was intranasally inoculated into mice that had been vaccinated once. The virus titers in the lungs after the challenge were measured to assess the protective immunity induced by the vaccines (Table 2). The virus titers in the lungs were <101.0−3.8 PFU/g in mice injected with 2 or 10 μg of protein of the Dk/Vietnam/2012 vaccine. The virus titers in the lungs of the mice injected with the Dk/Hong Kong/960 vaccine containing 10 or 50 μg of protein were 104.2−5.4 PFU/g, and those of mice injected with 2 μg of protein were not significantly different from those of control mice. These results indicate that the Dk/Vietnam/2012 and Dk/Hong Kong/960 vaccines induced immunity in mice to suppress virus replication in the lungs. The potency of the vaccine was also evaluated by observing weight loss of mice after the challenge (Fig. 2 A and B). The mice injected with 2 or 10 µg of the Dk/Vietnam/2012 vaccine or 50 µg of the Dk/Hong Kong/960 vaccine did not display significant weight loss. The body weight of mice injected with 2 or 10 µg of the Dk/Hong Kong/960 vaccine recovered faster than the control group mice. These results indicate that the Dk/Vietnam/2012 and Dk/Hong Kong/960 vaccines induced immunity in mice and lessened the impact of disease caused by the challenge strain.
Changes in the body weight of mice vaccinated intraperitoneally once with the Dk/Hong Kong/960 (A) or Dk/Vietnam/2012 (B), and twice with the Dk/Hong Kong/960 (C) vaccine after challenge with Dk/Vietnam/2012. Data are shown as the mean body weight of five mice ± standard error. Statistical significance was calculated by Student’s t-test. *, P < 0.05 versus the group of mice injected with PBS
To improve the efficacy of the Dk/Hong Kong/960 vaccine, the vaccine was intraperitoneally injected twice into mice. At 2 weeks after the second injection, the serum NT antibody titers against Dk/Hong Kong/960 were higher than those of mice injected once (Table 2). NT antibodies against Dk/Vietnam/2012 were also detected in the sera of all vaccinated mice (Table 2). Virus replication in the lungs of all of the vaccinated mice was significantly suppressed compared with that of the control group (Table 2). The mice injected twice with 50 µg of the Dk/Hong Kong/960 vaccine did not display significant weight loss, and those injected with 2 or 10 µg of the Dk/Hong Kong/960 vaccine recovered their weight faster than the mice in the control group (Fig. 2 C). These results indicate that the potency of the Dk/Hong Kong/960 vaccine against the challenge strain was enhanced by two intraperitoneal injections in mice.
Discussion
The HA genes of H6 virus strains in the influenza virus library were phylogenetically divided into five groups: the Early, Group II, Group III, and W312 sublineages and the North America lineage (Fig. 1). Antigenic analysis revealed that the antigenicity of the viruses of each sublineage differed from that of the others (Table 1). Since it is not possible to predict which virus of any sublineage would cause pandemic influenza in humans, it is important to prepare a vaccine that broadly cross-reacts with H6 viruses of all sublineages. It was reported that the live attenuated A/teal/Hong Kong/W312/1997 (H6N1) vaccine induced cross-protective immunity against the viruses of the Early, and W312 sublineages and the North America lineage in mice and ferrets [2, 28]. In the present study, it was observed that an antiserum to an Early sublineage virus broadly reacted with viruses of all sublineages, including Group II and Group III (Table 1). A comparison of the antigenic sites of the HAs of different H6 lineages may reveal the reason for this broad reactivity; however, the antigenic sites of the HA of H6 virus have not been defined yet. In the present study, therefore, it is difficult to analyze the antigenic sites of the HA of H6 viruses. From the six H6 viruses of the Early sublineage in the library, Dk/Hong Kong/960 was selected as a vaccine strain because this strain exhibited the best growth potential in embryonated chicken eggs and the highest protein yield. To evaluate the potency of this vaccine, Dk/Vietnam/2012 from the H6 viruses of Group II, which are antigenically different from other H6 viruses, was selected as a challenge strain.
It has been reported that whole-virus-particle vaccines induced strong immune responses, and an H5N1 whole-particle vaccine induced protective immunity against a challenge with an antigenically distinct virus [6, 10, 16]. In the present study, an inactivated whole-particle vaccine was prepared from an H6 avian influenza virus, Dk/Hong Kong/960, from the influenza virus library. The potency of this vaccine against Dk/Vietnam/2012 was evaluated in mice. The Dk/Hong Kong/960 vaccine conferred immunity in mice and reduced the severity of disease caused by a challenge with Dk/Vietnam/2012 (Fig. 2, Table 2). These results indicate that the whole-particle vaccine has potency even against the antigenically distinct H6 virus in mice.
Vaccination is one of the important control measures for human influenza; however, approximately 6 months are required to produce vaccines [5]. To prepare for future influenza pandemics, avian influenza viruses of 144 combinations of 16 HA and 9 NA subtypes have been stocked in the influenza virus library. Since the pathogenicity, antigenicity, genetic information, and yield in embryonated chicken eggs of the viruses in the influenza virus library were assessed previously, we can provide vaccine strains immediately. The present study indicates that the whole-virus-particle vaccine prepared from a virus strain from the influenza virus library is useful as a vaccine against pandemic influenza.
References
Arzey GG, Kirkland PD, Arzey KE, Frost M, Maywood P, Conaty S, Hurt AC, Deng YM, Iannello P, Barr I, Dwyer DE, Ratnamohan M, McPhie K, Selleck P (2012) Influenza virus A (H10N7) in chickens and poultry abattoir workers, Australia. Emerg Infect Dis 18:814–816
Chen Z, Santos C, Aspelund A, Gillim-Ross L, Jin H, Kemble G, Subbarao K (2009) Evaluation of live attenuated influenza a virus h6 vaccines in mice and ferrets. J Virol 83:65–72
Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, Garten RJ, Gubareva LV, Xu X, Bridges CB, Uyeki TM, Team NS-OIAHNVI (2009) Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med 360:2605–2615
DeLay PD, Casey HL, Tubiash HS (1967) Comparative study of fowl plague virus and a virus isolated from man. Public Health Rep 82:615–620
Gerdil C (2003) The annual production cycle for influenza vaccine. Vaccine 21:1776–1779
Hagenaars N, Mastrobattista E, Glansbeek H, Heldens J, van den Bosch H, Schijns V, Betbeder D, Vromans H, Jiskoot W (2008) Head-to-head comparison of four nonadjuvanted inactivated cell culture-derived influenza vaccines: effect of composition, spatial organization and immunization route on the immunogenicity in a murine challenge model. Vaccine 26:6555–6563
Hoffmann E, Stech J, Guan Y, Webster RG, Perez DR (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146:2275–2289
Huang K, Bahl J, Fan XH, Vijaykrishna D, Cheung CL, Webby RJ, Webster RG, Chen H, Smith GJ, Peiris JS, Guan Y (2010) Establishment of an H6N2 influenza virus lineage in domestic ducks in southern China. J Virol 84:6978–6986
Ito T, Okazaki K, Kawaoka Y, Takada A, Webster RG, Kida H (1995) Perpetuation of influenza A viruses in Alaskan waterfowl reservoirs. Arch Virol 140:1163–1172
Itoh Y, Ozaki H, Tsuchiya H, Okamoto K, Torii R, Sakoda Y, Kawaoka Y, Ogasawara K, Kida H (2008) A vaccine prepared from a non-pathogenic H5N1 avian influenza virus strain confers protective immunity against highly pathogenic avian influenza virus infection in cynomolgus macaques. Vaccine 26:562–572
Itoh Y, Ozaki H, Ishigaki H, Sakoda Y, Nagata T, Soda K, Isoda N, Miyake T, Ishida H, Okamoto K, Nakayama M, Tsuchiya H, Torii R, Kida H, Ogasawara K (2010) Subcutaneous inoculation of a whole virus particle vaccine prepared from a non-pathogenic virus library induces protective immunity against H7N7 highly pathogenic avian influenza virus in cynomolgus macaques. Vaccine 28:780–789
Kida H, Yanagawa R (1979) Isolation and characterization of influenza a viruses from wild free-flying ducks in Hokkaido, Japan. Zentralbl Bakteriol Orig A 244:135–143
Kida H, Ito T, Yasuda J, Shimizu Y, Itakura C, Shortridge KF, Kawaoka Y, Webster RG (1994) Potential for transmission of avian influenza viruses to pigs. J Gen Virol 75(Pt 9):2183–2188
Kim HR, Lee YJ, Lee KK, Oem JK, Kim SH, Lee MH, Lee OS, Park CK (2010) Genetic relatedness of H6 subtype avian influenza viruses isolated from wild birds and domestic ducks in Korea and their pathogenicity in animals. J Gen Virol 91:208–219
Kishida N, Sakoda Y, Shiromoto M, Bai GR, Isoda N, Takada A, Laver G, Kida H (2008) H2N5 influenza virus isolates from terns in Australia: genetic reassortants between those of the Eurasian and American lineages. Virus Genes 37:16–21
Lu X, Edwards LE, Desheva JA, Nguyen DC, Rekstin A, Stephenson I, Szretter K, Cox NJ, Rudenko LG, Klimov A, Katz JM (2006) Cross-protective immunity in mice induced by live-attenuated or inactivated vaccines against highly pathogenic influenza A (H5N1) viruses. Vaccine 24:6588–6593
Nomura N, Sakoda Y, Soda K, Okamatsu M, Kida H (2012) An H9N2 influenza virus vaccine prepared from a non-pathogenic isolate from a migratory duck confers protective immunity in mice against challenge with an H9N2 virus isolated from a girl in Hong Kong. J Vet Med Sci 74:441–447
Okamatsu M, Nishi T, Nomura N, Yamamoto N, Sakoda Y, Sakurai K, Chu HD, Thanh LP, Van Nguyen L, Van Hoang N, Tien TN, Yoshida R, Takada A, Kida H (2013) The genetic and antigenic diversity of avian influenza viruses isolated from domestic ducks, muscovy ducks, and chickens in northern and southern Vietnam, 2010-2012. Virus Genes 47:317–329
Okamatsu M, Sakoda Y, Hiono T, Yamamoto N, Kida H (2013) Potency of a vaccine prepared from A/swine/Hokkaido/2/1981 (H1N1) against A/Narita/1/2009 (H1N1) pandemic influenza virus strain. Virol J 10:47
Okazaki K, Takada A, Ito T, Imai M, Takakuwa H, Hatta M, Ozaki H, Tanizaki T, Nagano T, Ninomiya A, Demenev VA, Tyaptirganov MM, Karatayeva TD, Yamnikova SS, Lvov DK, Kida H (2000) Precursor genes of future pandemic influenza viruses are perpetuated in ducks nesting in Siberia. Arch Virol 145:885–893
Peiris M, Yam WC, Chan KH, Ghose P, Shortridge KF (1999) Influenza A H9N2: aspects of laboratory diagnosis. J Clin Microbiol 37:3426–3427
Sakabe S, Sakoda Y, Haraguchi Y, Isoda N, Soda K, Takakuwa H, Saijo K, Sawata A, Kume K, Hagiwara J, Tuchiya K, Lin Z, Sakamoto R, Imamura T, Sasaki T, Kokumai N, Kawaoka Y, Kida H (2008) A vaccine prepared from a non-pathogenic H7N7 virus isolated from natural reservoir conferred protective immunity against the challenge with lethal dose of highly pathogenic avian influenza virus in chickens. Vaccine 26:2127–2134
Sakoda Y, Sugar S, Batchluun D, Erdene-Ochir TO, Okamatsu M, Isoda N, Soda K, Takakuwa H, Tsuda Y, Yamamoto N, Kishida N, Matsuno K, Nakayama E, Kajihara M, Yokoyama A, Takada A, Sodnomdarjaa R, Kida H (2010) Characterization of H5N1 highly pathogenic avian influenza virus strains isolated from migratory waterfowl in Mongolia on the way back from the southern Asia to their northern territory. Virology 406:88–94
Schäfer JR, Kawaoka Y, Bean WJ, Süss J, Senne D, Webster RG (1993) Origin of the pandemic 1957 H2 influenza A virus and the persistence of its possible progenitors in the avian reservoir. Virology 194:781–788
Shichinohe S, Okamatsu M, Sakoda Y, Kida H (2013) Selection of H3 avian influenza viruses with SAα2,6Gal receptor specificity in pigs. Virology 444:404–408
Skehel JJ, Hay AJ (1978) Nucleotide sequences at the 5’ termini of influenza virus RNAs and their transcripts. Nucleic Acids Res 5:1207–1219
Soda K, Sakoda Y, Isoda N, Kajihara M, Haraguchi Y, Shibuya H, Yoshida H, Sasaki T, Sakamoto R, Saijo K, Hagiwara J, Kida H (2008) Development of vaccine strains of H5 and H7 influenza viruses. Jpn J Vet Res 55:93–98
Talaat KR, Karron RA, Luke CJ, Thumar B, McMahon BA, Chen GL, Lamirande EW, Jin H, Coelingh KL, Kemble G, Subbarao K (2011) An open label Phase I trial of a live attenuated H6N1 influenza virus vaccine in healthy adults. Vaccine 29:3144–3148
Yamamoto N, Sakoda Y, Motoshima M, Yoshino F, Soda K, Okamatsu M, Kida H (2011) Characterization of a non-pathogenic H5N1 influenza virus isolated from a migratory duck flying from Siberia in Hokkaido, Japan, in October 2009. Virol J 8:65
Yuan J, Zhang L, Kan X, Jiang L, Yang J, Guo Z, Ren Q (2013) Origin and molecular characteristics of a Novel 2013 Avian influenza A(H6N1) virus causing human infection in Taiwan. Clin Infect Dis 57:1367–1368
Zhang G, Kong W, Qi W, Long LP, Cao Z, Huang L, Qi H, Cao N, Wang W, Zhao F, Ning Z, Liao M, Wan XF (2011) Identification of an H6N6 swine influenza virus in southern China. Infect Genet Evol 11:1174–1177
Acknowledgements
We thank Dr. Shortridge K. F. and Dr. Guan Y. for providing viruses. This study was supported by the Strategic Funds for the Promotion of Science and Technology (2011–2013), Japan (JST), the Global Center of Excellence (GCOE) Program of Hokkaido University, and the Japan Initiative for Global Research Network on Infectious Disease (J-GRID).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nishi, T., Sakoda, Y., Okamatsu, M. et al. Potency of an inactivated influenza vaccine prepared from A/duck/Hong Kong/960/1980 (H6N2) against a challenge with A/duck/Vietnam/OIE-0033/2012 (H6N2) in mice. Arch Virol 159, 2567–2574 (2014). https://doi.org/10.1007/s00705-014-2107-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-014-2107-2