Abstract
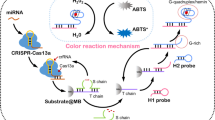
A method is presented that uses photoinduced electron transfer (PET) for the determination of microRNAs (miRNAs) in clinical serum samples and complicated cell samples by using a smartphone. miRNA-21 is adopted as a model analyte. A 3′-phosphorylated DNA probe containing AgNCs is synthesized and hybridized with miRNA-21. Subsequently, the probe is cleaved specifically by duplex-specific nuclease to form 3′-hydroxylated products, then extended by terminal deoxynucleotidyl transferase (TdT) with superlong G for G-quadruplex/hemin units fabrication. In this way, PET occurred between AgNCs and produced G-quadruplex/hemin units, leading to the fluorescence quenching of AgNCs. Notably, the fluorescence images can be captured and translated into digital information by smartphone, resulting in a direct quantitative determination of miRNA. As a result, our strategy for miRNA assay is achieved with a satisfactory detection limit of 1.43 pM. Interestingly, TdT-propelled G-quadruplex/hemin units as multiple electron acceptors promote the sensitivity of miRNA monitoring. Different miRNAs assays are realized by adjusting the complimentary sequences of DNA probe. These qualities not only broaden the practical application of PET-based strategy, but also provide a new insight into the nucleic acid detection.
Graphical abstract

Schematic representation of AgNCs and enzyme-propelled photoinduced electron transfer strategy. It has been successfully applied for detection of miRNA by image analysis software. The method displays portability and accuracy for miRNA determination, meeting the potential for biochemical and clinical applications in resource-limited settings.






Similar content being viewed by others
References
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6:857–866. https://doi.org/10.1038/nrc1997
Ramani V, Qiu R, Shendure J (2015) High-throughput determination of RNA structure by proximity ligation. Nat Biotechnol 33:980–984. https://doi.org/10.1038/nbt.3289
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebet BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838. https://doi.org/10.1038/nature03702
Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T (2007) Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 39:673–677. https://doi.org/10.1038/ng2003
Li S, Xu LG, Ma W, Wu XL, Sun MZ, Kuang H, Wang LB, Kotov NA, Xu CL (2016) Dual-mode ultrasensitive quantification of microRNA in living cells by chiroplasmonic nanopyramids self-assembled from gold and upconversion nanoparticles. J Am Chem Soc 138:306–312. https://doi.org/10.1021/jacs.5b10309
Reddy KB (2015) MicroRNA(miRNA) in cancer. Cancer Cell Int 15:38. https://doi.org/10.1186/s12935-015-0185-1
Chandrasekaran AR, MacIsaac M, Dey P, Levchenko O, Zhou LF, Andres M, Dey BK, Halvorsen K (2019) Cellular microRNA detection with miRacles: microRNA activated. Sci Adv 5:eaau9443. https://doi.org/10.1126/sciadv.aau9443
Kim S, Park JE, Hwang W, Seo J, Lee YK, Hwang JH, Nam JM (2017) Optokinetically encoded nanoprobe-based multiplexing strategy for microRNA profiling. J Am Chem Soc 139:3558–3566. https://doi.org/10.1021/jacs.7b01311
Várallyay É, Burgyán J, Havelda Z (2008) MicroRNA detection by northern blotting using locked nucleic acid probes. Nature 3:190–196. https://doi.org/10.1038/nprot.2007.528
Lee JM, Cho H, Jung Y (2010) Fabrication of a structure-specific RNA binder for array detection of label-free microRNA. Angew Chem Int Edit 49:8662–8665. https://doi.org/10.1002/anie.201004000
Wang C, Han Q, Mo FJ, Chen M, Xiong ZW, Fu YZ (2020) Novel luminescent nanostructured coordination polymer: facile fabrication and application in electrochemiluminescence biosensor for microRNA-141 detection. Anal Chem 92:12145–12151. https://doi.org/10.1021/acs.analchem.0c00130
Xian LM, Xu F, Liu JZ, Xu N, Li HD, Ge HY, Shao K, Fan JL, Xiao GS, Peng XJ (2019) MicroRNA detection with turnover amplification via hybridization-mediated staudinger reduction for pancreatic cancer diagnosis. J Am Chem Soc 141:20490–20497. https://doi.org/10.1021/jacs.9b11272
Ge J, Hu Y, Deng RJ, Li ZH, Zhang KX, Shi ML, Yang D, Cai R, Tan WH (2020) Highly sensitive microRNA detection by coupling nicking-enhanced rolling circle amplification with MoS2 quantum dots. Anal Chem 92:13588–13594. https://doi.org/10.1021/acs.analchem.0c03405
Bai YY, Wu Z, Xu CM, Zhang L, Feng J, Pang DW, Zhang ZL (2020) One-to-many single entity electrochemistry biosensing for ultrasensitive detection of microRNA. Anal Chem 92:853–858. https://doi.org/10.1021/acs.analchem.9b03492
Shamsipur M, Molaabasi F, Hosseinkhani S, Rahmati F (2016) Detection of early stage apoptotic cells based on label-free cytochrome c assay using bioconjugated metal nanoclusters as fluorescent probes. Anal Chem 88:2188–2197. https://doi.org/10.1021/acs.analchem.5b03824
Fan WJ, Qi Y, Qiu LY, He P, Liu CH, Li ZP (2018) Click chemical ligation-initiated on-bead DNA polymerization for the sensitive flow cytometric detection of 3′-terminal 2′-O-methylated plant microRNA. Anal Chem 90:5390–5397. https://doi.org/10.1021/acs.analchem.8b00589
Wu TT, Yang YM, Cao Y, Song YC, Xu LP, Zhang XJ, Wang ST (2018) Bioinspired DNA-inorganic hybrid nanoflowers combined with a personal glucose meter for onsite detection of miRNA. ACS Appl Mater Interfaces 10:42050–42057. https://doi.org/10.1021/acsami.8b15917
Fabri-Faja N, Calvo-Lozano O, Dey P, Terborg RA, Estévez MC, Belushkin A, Yesilköy F, Duempelmann L, Altug H, Pruneri V, Pruneri LM (2019) Early sepsis diagnosis via protein and miRNA biomarkers using a novel point-of-care photonic biosensor. Anal Chim Acta 1077:232–242. https://doi.org/10.1016/j.aca.2019.05.038
Lv WJ, Liu CY, Ma Y, Wang X, Luo JJ, Ye WC (2019) Multi-hydrogen bond assisted SERS detection of adenine based on multifunctional graphene oxide/poly (diallyldimethyl ammonium chloride)/Ag nanocomposites. Talanta 204:372–378. https://doi.org/10.1016/j.talanta.2019.06.012
Zhang LB, Zhu JB, Guo SJ, Li T, Li J, Wang EK (2013) Photoinduced electron transfer of DNA/Ag nanoclusters modulated by G-quadruplex/hemin complex for the construction of versatile biosensors. J Am Chem Soc 135:2403–2406. https://doi.org/10.1021/ja3089857
Chen WY, Lan GY, Chang HT (2011) Use of fluorescent DNA-templated gold/silver nanoclusters for the detection of sulfide ions. Anal Chem 83:9450–9455. https://doi.org/10.1021/ac202162u
Chen ND, Li JY, Feng XZ, Yang YP, Zhu L, Chen XM, Liu X, Li Y, Wang CC, Xia LG (2020) Label-free and self-assembled fluorescent DNA nanopompom for determination of miRNA-21. Microchim Acta 187:432. https://doi.org/10.1007/s00604-020-04377-6
Borghei YS, Hosseini M, Ganjali MR, Ju H (2018) Colorimetric and energy transfer based fluorometric turn-on method for determination of microRNA using silver nanoclusters and gold nanoparticles. Microchim Acta 185:286. https://doi.org/10.1007/s00604-018-2825-3
Borghei YS, Hosseini M, Ganjali MR (2017) Fluorometric determination of microRNA via FRET between silver nanoclusters and CdTe quantum dots. Microchim Acta 184:4713–4721. https://doi.org/10.1007/s00604-017-2512-9
Miao XM, Cheng ZY, Ma HY, Li ZB, Xue N, Wang P (2018) Label-free platform for microRNA detection based on the fluorescence quenching of positively charged gold nanoparticles to silver nanoclusters. Anal Chem 90:1098–1103. https://doi.org/10.1021/acs.analchem.7b01991
Zhang M, Guo SM, Li YR, Zuo P, Ye BC (2012) A label-free fluorescent molecular beacon based on DNA-templated silver nanoclusters for detection of adenosine and adenosine deaminase. Chem Commun 48:5488–5490. https://doi.org/10.1039/c2cc31626a
Guo WW, Yuan JP, Wang EK (2011) Strand exchange reaction modulated fluorescence “off-on” switching of hybridized DNA duplex stabilized silver nanoclusters. j 47:10930–10932. https://doi.org/10.1039/c1cc11921d
Guo WW, Yuan JP, Dong QZ, Wang EK (2009) Highly sequence-dependent formation of fluorescent silver nanoclusters in hybridized DNA duplexes for single nucleotide mutation identification. J Am Chem Soc 132:932–934. https://doi.org/10.1021/ja907075s
Lu SS, Wang S, Zhao JH, Sun J, Yang XR (2017) Fluorescence light-up biosensor for microRNA based on the distance-dependent photoinduced electron transfer. Anal Chem 89:8429–8436. https://doi.org/10.1021/acs.analchem.7b01900
Jin R, Wang FY, Li QY, Yan X, Liu MQ, Chen Y, Zhou WR, Gao H, Sun P, Lu GY (2021) Construction of multienzyme-hydrogel sensor with smartphone detector for on-site monitoring of organophosphorus pesticide. Sens Actuat B-Chem 327:128922. https://doi.org/10.1016/j.snb.2020.128922
Li Y, Zhou LL, Ni W, Luo QY, Zhu CN, Wu YH (2019) Portable and field-ready detection of circulating microRNAs with paper-based bioluminescent sensing and isothermal amplification. Anal Chem 91:14838–14841. https://doi.org/10.1021/acs.analchem.9b04422
Cheng YY, Xie YF, Li CM, Li YF, Huang CZ (2019) Förster resonance energy transfer-based soft nanoballs for specific and amplified detection of microRNAs. Anal Chem 9:11023–11029. https://doi.org/10.1021/acs.analchem.9b01281
Wang W, Kong T, Zhang D, Zhang JN, Cheng GS (2015) Label-free microRNA detection based on fluorescence quenching of gold nanoparticles with a Competitive hybridization. Anal Chem 87:10822–10829. https://doi.org/10.1021/acs.analchem.5b01930
Zhen SJ, Xiao X, Li CH, Huang CZ (2017) An enzyme-free DNA circuit-assisted graphene oxide enhanced fluorescence anisotropy assay for microRNA detection with improved sensitivity and selectivity. Anal Chem 89:8766–8771. https://doi.org/10.1021/acs.analchem.7b00955
Castañeda AD, Brenes NJ, Kondajji A, Crooks RM (2017) Detection of microRNA by electrocatalytic amplification: a general approach for single-particle biosensing. J Am Chem Soc 139:7657–7664. https://doi.org/10.1021/jacs.7b03648
Guo YS, Wang YJ, Yang GX, Xu JJ, Chen HY (2016) MicroRNA-mediated signal amplification coupled with GNP/dendrimers onamass-sensitive biosensor and its applications in intracellular microRNA quantification. Biosens Bioelectron 85:897–902. https://doi.org/10.1016/j.bios.2016.06.013
Leea J, Kimb YK, Leea S, Yoona S, Kima WK (2019) Graphene oxide-based NET strategy for enhanced colorimetric sensing of miRNA. Sens Actuat B-Chem 282:861–867. https://doi.org/10.1016/j.snb.2018.11.149
Yin BC, Liu YQ, Ye BC (2012) One-step, multiplexed fluorescence detection of microRNAs based on duplex-specific nuclease signal amplification. J Am Chem Soc 134:5064–5067. https://doi.org/10.1021/ja300721s
Xi Q, Zhou DM, Kan YY, Ge J, Wu ZK, Yu RQ, Jiang JH (2014) Highly sensitive and selective strategy for microRNA detection based on WS2 nanosheet mediated fluorescence quenching and duplex-specific nuclease signal amplification. Anal Chem 86:1361–1365. https://doi.org/10.1021/ac403944c
Degliangeli F, Kshirsagar P, Brunetti V, Pompa PP, Fiammengo R (2014) Absolute and direct microRNA quantification using DNA-gold nanoparticle probes. J Am Chem Soc 136(6):2264–2267. https://doi.org/10.1021/ja412152x
Xu SH, Nie YY, Jiang LP, Wang J, Xu GY, Wang W, Luo XL (2018) Polydopamine nanosphere/gold nanocluster (Au NC)-based nanoplatform for dual color simultaneous detection of multiple tumor-related microRNAs with DNase-I-assisted target recycling amplification. Anal Chem 90:4039–4045. https://doi.org/10.1021/acs.analchem.7b05253
Li DX, Zhou WJ, Yuan R, Xiang Y (2017) A DNA-fueled and catalytic molecule machine lights up trace under-expressed microRNAs in living cells. Anal Chem 89:9934–9940. https://doi.org/10.1021/acs.analchem.7b02247
Zhang K, Wang K, Zhu X, Gao Y, Xie MH (2014) Rational design of signal-on biosensors by using photoinduced electron transfer between Ag nanoclusters and split G-quadruplex halves-hemin complexes. Chem Commun 50:14221–14224. https://doi.org/10.1039/c4cc06664b
Funding
This work is financially supported by the National Natural Science Foundation of China (Grant Nos. 31800670 and 21603003), the Key Project of Natural Science Foundation of Anhui Provincial Department of Education (Grant No. KJ2019A0583), and the Foundation of State Key Laboratory of High-efficiency Utilization of Coal and Green Chemical Engineering (Grant No. 2018-K34).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOC 11319 kb)
Rights and permissions
About this article
Cite this article
Yang, Y., Liu, S., Cui, X. et al. Sensitive detection of miRNA based on enzyme-propelled multiple photoinduced electron transfer strategy. Microchim Acta 188, 219 (2021). https://doi.org/10.1007/s00604-021-04874-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-021-04874-2