Abstract
Key message
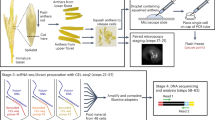
A detailed, step-by-step protocol for isolation of rice gametes for transcriptional profiling, with a general workflow that includes controls for RNA contamination from surrounding cells and tissues is presented.
Abstract
Characterization of the transcriptome and other -omics studies of flowering plant gametes are challenging as a consequence of the small sizes and relative inaccessibility of these cells. Collecting such poorly represented cells is also complicated by potential contamination from surrounding sporophytic, adjacent gametophytic tissues and difficulties in extracting high-quality intact cells. Here we present detailed, step-by-step procedures for collecting intact, unfixed rice (Oryza sativa) egg cells and sperm cells without enzymatic treatments. In addition, we also present a general workflow for assessing sample purity by RT-PCR, using primers specific for marker genes preferentially expressed in surrounding cells and tissues. These protocols should facilitate future studies of genome-scale characterization of gametes in this important model crop.




Similar content being viewed by others
Notes
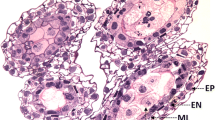
Care should be taken to identify mature florets that are staged correctly. In mature florets, stamens should occupy most of the floret prior to anthesis.
For RT-PCR, we used MyTaq Red Mix (Bioline BIO-25043). All primers listed in Supplementary Table 1 performed using an annealing temperature of 59 °C.
References
Abiko M, Furuta K, Yamauchi Y, Fujita C, Taoka M, Isobe T, Okamoto T (2013) Identification of proteins enriched in rice egg or sperm cells by single-cell proteomics. PLoS ONE. https://doi.org/10.1371/journal.pone.0069578
Anderson SN, Johnson CS, Jones DS, Conrad LJ, Gou X, Russell SD, Sundaresan V (2013) Transcriptomes of isolated Oryza sativa gametes characterized by deep sequencing: evidence for distinct sex-dependent chromatin and epigenetic states before fertilization. Plant J 76(5):729–741. https://doi.org/10.1111/tpj.12336
Anderson SN, Johnson CS, Chesnut J, Jones DS, Khanday I, Woodhouse M, Li C, Conrad LJ, Russell SD, Sundaresan V (2017) The zygotic transition is initiated in unicellular plant zygotes with asymmetric activation of parental genomes. Dev Cell 43(3):349.e4–358.e4. https://doi.org/10.1016/j.devcel.2017.10.005
Cook M, Thilmony R (2012) The OsGEX2 gene promoter confers sperm cell expression in transgenic rice. Plant Mol Biol Rep 30(5):1138–1148. https://doi.org/10.1007/s11105-012-0429-3
Gou X, Wang S, Chen F (2011) Isolation and cytological observation of viable sperm cells of rice. Acta Botanica Sinica 41(6):669–671. https://doi.org/10.1360/zd-2013-43-6-1064
Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa nipponbare reference genome using next generation sequence and optical map data. Rice 6(1):3–10. https://doi.org/10.1186/1939-8433-6-4
Kyozuka J, Shimamoto K, Ag OR (2002) Ectopic expression of OsMADS3, a rice ortholog of AGAMOUS, caused a homeotic transformation of lodicules to stamens in transgenic rice plants. Plant Cell Physiol 43(1):130–135
Lu XC, Gong HQ, Huang ML, Bai SL, He YB, Mao X, Geng Z, Li SG, Wei L, Yuwen JS, Xu ZH, Bai SN (2006) Molecular analysis of early rice stamen development using organ-specific gene expression profiling. Plant Mol Biol 61(6):845–861. https://doi.org/10.1007/s11103-006-0054-3
Ohnishi T, Takanashi H, Mogi M, Takahashi H, Kikuchi S, Yano K, Okamoto T, Fujita M, Kurata N, Tsutsumi N (2011) Distinct gene expression profiles in egg and synergid cells of rice as revealed by cell type-specific microarrays. Plant Physiol 155(2):881–891. https://doi.org/10.1104/pp.110.167502
Okada T, Endo M, Singh MB, Bhalla PL (2005) Analysis of the histone H3 gene family in Arabidopsis and identification of the male-gamete-specific variant AtMGH3. Plant J 44(4):557–568. https://doi.org/10.1111/j.1365-313X.2005.02554.x
Okamoto T (2017) Analysis of proteins enriched in rice gamete. In: Schmidt A (ed) Plant germline development: methods and protocols. Springer, New York, pp 251–263. https://doi.org/10.1007/978-1-4939-7286-9_20
Russell SD, Gou XP, Wong CE, Wang X, Yuan T, Wei XP, Bhalla PL, Singh MB (2012) Genomic profiling of rice sperm cell transcripts reveals conserved and distinct elements in the flowering plant male germ lineage. New Phytol 195(3):560–573. https://doi.org/10.1111/j.1469-8137.2012.04199.x
Russell SD, Jones DS, Anderson S, Wang X, Sundaresan V, Gou X (2017) Isolation of rice sperm cells for transcriptional profiling. In: Schmidt A (ed) Plant germline development: methods and protocols. Springer, New York, pp 211–219. https://doi.org/10.1007/978-1-4939-7286-9_17
Schon MA, Nodine MD (2017) Widespread contamination of Arabidopsis embryo and endosperm transcriptome data sets. Plant Cell 29(4):608–617. https://doi.org/10.1105/tpc.16.00845
Uchiumi T, Uemura I, Okamoto T (2007) Establishment of an in vitro fertilization system in rice (Oryza sativa L.). Planta 226(3):581–589. https://doi.org/10.1007/s00425-007-0506-2
Xiao H, Wang Y, Liu D, Wang W, Li X, Zhao X, Xu J, Zhai W, Zhu L (2003) Functional analysis of the rice AP3 homologue OsMADS16 by RNA interference. Plant Mol Biol 52(5):957–966. https://doi.org/10.1023/A:1025401611354
Xu HP, Tsao TH (1997) Detection and immunolocalization of glycoproteins of the plasma membrane of maize sperm cells. Protoplasma 198(3–4):125–129. https://doi.org/10.1007/BF01287560
Xu H, Weterings K, Vriezen W, Feron R, Xue Y, Derksen J, Mariani C (2002) Isolation and characterization of male-germ-cell transcripts in Nicotiana tabacum. Sex Plant Reprod 14(6):339–346. https://doi.org/10.1007/s00497-002-0128-6
Zhang J, Dong WH, Galli A, Potrykus I (1999) Regeneration of fertile plants from isolated zygotes of rice (Oryza sativa). Plant Cell Rep 19(2):128–132. https://doi.org/10.1007/s002990050722
Zhang YN, Wei DM, He EM, Miao S, Tian HQ, Russell SD (2010) Isolation of male and female gametes of rice. Crop Sci 50(6):2457-2463. https://doi.org/10.2135/cropsci2010.02.0066
Acknowledgements
We thank Imtiyaz Khanday, Jonathan Gent, Sarah Anderson and Daniel Jones for helpful advice for optimizing the experimental methods. We thank Debra Skinner for assistance in artwork. This research was funded by the National Science Foundation (Award No. IOS-1547760) and the USDA Agricultural Experiment Station (Project No. CA-D-XXX-6973-H).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Thomas Dresselhaus.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A contribution to the special issue ‘Cellular Omics Methods in Plant Reproduction Research’.
Scott D. Russell and Venkatesan Sundaresan: Senior authors.
Venkatesan Sundaresan: Lead contact.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, C., Xu, H., Russell, S.D. et al. Step-by-step protocols for rice gamete isolation. Plant Reprod 32, 5–13 (2019). https://doi.org/10.1007/s00497-019-00363-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-019-00363-y