Abstract
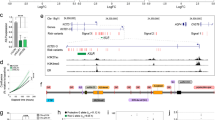
Nonmutational epigenetic reprogramming is a crucial mechanism contributing to the pronounced heterogeneity of prostate cancer (PCa). Among these mechanisms, N6-methyladenosine (m6A)-modified long non-coding RNAs (lncRNAs) have emerged as key players. However, the precise roles of m6A-modified lncRNAs in PCa remain to be elucidated. In this study, methylated RNA immunoprecipitation sequencing (MeRIP-seq) was conducted on primary and metastatic PCa samples, leading to the identification of 21 lncRNAs exhibiting differential methylation and expression patterns. We further established a PCa prognostic signature, named m6A-modified lncRNA score (mLs), based on 9 differential methylated lncRNAs in 4 multicenter cohorts. The high mLs score cohort exhibited a tendency for earlier biochemical recurrence (BCR) compared to the low mLs score cohort. Remarkably, the predictive performance of the mLs score surpassed that of five previously reported lncRNA-based signatures. Functional enrichment analysis underscored a negative correlation between the mLs score and lipid metabolism. Additionally, through MeRIP-qPCR, we pinpointed a hub gene, MIR210HG, which was validated through in vitro and in vivo experiments. These findings collectively illuminate the landscape of m6A-methylated lncRNAs in PCa tissue via MeRIP-seq and harness this information to prognosticate PCa outcomes using the mLs score. Furthermore, our study validates, both experimentally and mechanistically, the facilitative role of MIR210HG in driving PCa progression.








Similar content being viewed by others
Data availability
The datasets utilized in this research has been delineated in the Materials and Methods section. For any further inquiries, please reach out to the corresponding author.
References
Angeles AK, Heckmann D, Flosdorf N et al (2020) The ERG-regulated LINC00920 promotes prostate cancer cell survival via the 14–3-3ε-FOXO pathway. Mol Cancer Res 18:1545–1559. https://doi.org/10.1158/1541-7786.Mcr-20-0021
Barbieri I, Kouzarides T (2020) Role of RNA modifications in cancer. Nat Rev Cancer 20:303–322. https://doi.org/10.1038/s41568-020-0253-2
Carrillo-Reixach J, Torrens L, Simon-Coma M et al (2020) Epigenetic footprint enables molecular risk stratification of hepatoblastoma with clinical implications. J Hepatol 73:328–341. https://doi.org/10.1016/j.jhep.2020.03.025
Chen M, Wei L, Law CT et al (2018) RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology 67:2254–2270. https://doi.org/10.1002/hep.29683
de Winter JC, Gosling SD (2016) Potter J Comparing the Pearson and Spearman correlation coefficients across distributions and sample sizes: a tutorial using simulations and empirical data. Psychol Methods 21:273–90. https://doi.org/10.1037/met0000079
Dominissini D, Moshitch-Moshkovitz S, Schwartz S et al (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485:201–206. https://doi.org/10.1038/nature11112
Feng ZH, Liang YP, Cen JJ et al (2022) m6A-immune-related lncRNA prognostic signature for predicting immune landscape and prognosis of bladder cancer. J Transl Med 20:492. https://doi.org/10.1186/s12967-022-03711-1
Fisher RA (1922) On the interpretation of χ2 from contingency tables, and the calculation of P. J R Stat Soc 85:87–94. https://doi.org/10.2307/2340521
Guo S, Zhang Y, Wang S et al (2021) LncRNA PCA3 promotes antimony-induced lipid metabolic disorder in prostate cancer by targeting MIR-132-3 P/SREBP1 signaling. Toxicol Lett 348:50–58. https://doi.org/10.1016/j.toxlet.2021.05.006
Hanahan D (2022) Hallmarks of cancer: new dimensions. Cancer Discov 12:31–46. https://doi.org/10.1158/2159-8290.Cd-21-1059
Hänzelmann S, Castelo R, Guinney JGSVA (2013) gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7. https://doi.org/10.1186/1471-2105-14-7
Ho KH, Shih CM, Liu AJ et al (2022) Hypoxia-inducible lncRNA MIR210HG interacting with OCT1 is involved in glioblastoma multiforme malignancy. Cancer Sci 113:540–552. https://doi.org/10.1111/cas.15240
Hu X, Peng WX, Zhou H et al (2020) IGF2BP2 regulates DANCR by serving as an N6-methyladenosine reader. Cell Death Differ 27:1782–1794. https://doi.org/10.1038/s41418-019-0461-z
Hu XL, Huang XT, Zhang JN et al (2022) Long noncoding RNA MIR210HG is induced by hypoxia-inducible factor 1α and promotes cervical cancer progression. Am J Cancer Res 12:2783–2797
Hu D, Cao Q, Tong M et al (2022) A novel defined risk signature based on pyroptosis-related genes can predict the prognosis of prostate cancer. BMC Med Genomics 15:24. https://doi.org/10.1186/s12920-022-01172-5
Kato H, Ichinose Y, Ohta M et al (2004) A randomized trial of adjuvant chemotherapy with uracil-tegafur for adenocarcinoma of the lung. N Engl J Med 350:1713–1721. https://doi.org/10.1056/NEJMoa032792
Khaliq M, Manikkam M, Martinez ED et al (2021) Epigenetic modulation reveals differentiation state specificity of oncogene addiction. Nat Commun 12:1536. https://doi.org/10.1038/s41467-021-21784-2
Kundaje A, Meuleman W, Ernst J et al (2015) Integrative analysis of 111 reference human epigenomes. Nature 518:317–30. https://doi.org/10.1038/nature14248
Lang C, Yin C, Lin K et al (2021) m(6) A modification of lncRNA PCAT6 promotes bone metastasis in prostate cancer through IGF2BP2-mediated IGF1R mRNA stabilization. Clin Transl Med 11:e426. https://doi.org/10.1002/ctm2.426
Li R, Zhu J, Zhong W-D et al (2021) PCaDB - a comprehensive and interactive database for transcriptomes from prostate cancer population cohorts. bioRxiv. https://doi.org/10.1101/2021.06.29.449134
Li B, Zhao R, Qiu W et al (2022) The N(6)-methyladenosine-mediated lncRNA WEE2-AS1 promotes glioblastoma progression by stabilizing RPN2. Theranostics 12:6363–6379. https://doi.org/10.7150/thno.74600
Li C, Pan B, Wang X et al (2022) Upregulated LINC01088 facilitates malignant phenotypes and immune escape of colorectal cancer by regulating microRNAs/G3BP1/PD-L1 axis. J Cancer Res Clin Oncol 148:1965–1982. https://doi.org/10.1007/s00432-022-03981-8
Li R, Zhu J, Zhong WD et al (2022) Comprehensive evaluation of machine learning models and gene expression signatures for prostate cancer prognosis using large population cohorts. Cancer Res 82:1832–1843. https://doi.org/10.1158/0008-5472.Can-21-3074
Liberzon A, Birger C, Thorvaldsdóttir H et al (2015) The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst 1:417–425. https://doi.org/10.1016/j.cels.2015.12.004
Liu J, Dou X, Chen C et al (2020) N (6)-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367:580–586. https://doi.org/10.1126/science.aay6018
Liu Z, Gao L, Cheng L et al (2023) The roles of N6-methyladenosine and its target regulatory noncoding RNAs in tumors: classification, mechanisms, and potential therapeutic implications. Exp Mol Med. https://doi.org/10.1038/s12276-023-00944-y
Lu J, Chen J, Lin Z et al (2023) A prognostic signature consisting of N6-methyladenosine modified mRNAs demonstrates clinical potential in prediction of biochemical recurrence and guidance on precision therapy in prostate cancer. Transl Oncol 33:101670. https://doi.org/10.1016/j.tranon.2023.101670
Mann HB, Whitney DR (1947) On a test of whether one of two random variables is stochastically larger than the other. Ann Math Stat 18:50–60
Mottet N, Bellmunt J, Bolla M et al (2017) EAU-ESTRO-SIOG Guidelines on Prostate Cancer. Part 1: screening, diagnosis, and local treatment with curative intent. Eur Urol 71:618–629. https://doi.org/10.1016/j.eururo.2016.08.003
Pei H, Dai Y, Yu Y et al (2023) The tumorigenic effect of lncRNA AFAP1-AS1 is mediated by translated peptide ATMLP under the control of m(6) A methylation. Adv Sci 10:e2300314. https://doi.org/10.1002/advs.202300314
Porman AM, Roberts JT, Duncan ED et al (2022) A single N6-methyladenosine site regulates lncRNA HOTAIR function in breast cancer cells. PLoS Biol 20:e3001885. https://doi.org/10.1371/journal.pbio.3001885
Raj A, van den Bogaard P, Rifkin SA et al (2008) Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods 5:877–9. https://doi.org/10.1038/nmeth.1253
Ritchie ME, Phipson B, Wu D et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47. https://doi.org/10.1093/nar/gkv007
Roundtree IA, Evans ME, Pan T et al (2017) Dynamic RNA modifications in gene expression regulation. Cell 169:1187–1200. https://doi.org/10.1016/j.cell.2017.05.045
Siegel RL, Miller KD, Fuchs HE et al (2022) Cancer statistics, 2022. CA Cancer J Clin 72:7–33. https://doi.org/10.3322/caac.21708
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Tibshirani R (1997) The lasso method for variable selection in the Cox model. Stat Med 16:385–95. https://doi.org/10.1002/(sici)1097-0258(19970228)16:4%3c385::aid-sim380%3e3.0.co;2-3
Wang X, Lu Z, Gomez A et al (2014) N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505:117–20. https://doi.org/10.1038/nature12730
Wang X, Zhao BS, Roundtree IA et al (2015) N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161:1388–99. https://doi.org/10.1016/j.cell.2015.05.014
Wang Y, Li W, Chen X et al (2019) MIR210HG predicts poor prognosis and functions as an oncogenic lncRNA in hepatocellular carcinoma. Biomed Pharmacother 111:1297–1301. https://doi.org/10.1016/j.biopha.2018.12.134
Wang AH, Jin CH, Cui GY et al (2020) MIR210HG promotes cell proliferation and invasion by regulating miR-503-5p/TRAF4 axis in cervical cancer. Aging (Albany NY) 12:3205–3217. https://doi.org/10.18632/aging.102799
Weng H, Huang H, Wu H et al (2018) METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell 22:191–205.e9. https://doi.org/10.1016/j.stem.2017.11.016
Wu Y, Liang Y, Li M et al (2022) Knockdown of long non-coding RNA SNHG8 suppresses the progression of esophageal cancer by regulating miR-1270/BACH1 axis. Bioengineered 13:3384–3394. https://doi.org/10.1080/21655979.2021.2021064
Xiao W, Adhikari S, Dahal U et al (2016) Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Mol Cell 61:507–519. https://doi.org/10.1016/j.molcel.2016.01.012
Yi YC, Chen XY, Zhang J et al (2020) Novel insights into the interplay between m(6)A modification and noncoding RNAs in cancer. Mol Cancer 19:121. https://doi.org/10.1186/s12943-020-01233-2
Yu G, Wang LG, Han Y et al (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16:284–7. https://doi.org/10.1089/omi.2011.0118
Yu L, Sui B, Fan W et al (2021) Exosomes derived from osteogenic tumor activate osteoclast differentiation and concurrently inhibit osteogenesis by transferring COL1A1-targeting miRNA-92a-1-5p. J Extracell Vesicles 10:e12056. https://doi.org/10.1002/jev2.12056
Zhang H, Shi X, Huang T et al (2020) Dynamic landscape and evolution of m6A methylation in human. Nucleic Acids Res 48:6251–6264. https://doi.org/10.1093/nar/gkaa347
Zhong C, Wu K, Wang S et al (2021) Autophagy-related circRNA evaluation reveals hsa_circ_0001747 as a potential favorable prognostic factor for biochemical recurrence in patients with prostate cancer. Cell Death Dis 12:726. https://doi.org/10.1038/s41419-021-04015-w
Zhong C, Long Z, Yang T et al (2023) M6A-modified circRBM33 promotes prostate cancer progression via PDHA1-mediated mitochondrial respiration regulation and presents a potential target for ARSI therapy. Int J Biol Sci 19:1543–1563. https://doi.org/10.7150/ijbs.77133
Zhu KP, Zhang CL, Ma XL et al (2019) Analyzing the interactions of mRNAs and ncRNAs to predict competing endogenous RNA networks in osteosarcoma chemo-resistance. Mol Ther 27:518–530. https://doi.org/10.1016/j.ymthe.2019.01.001
Acknowledgment
We would like to express our gratitude to all the contributors of the public databases and codes utilized in our study for generously providing the data for our analysis.
Funding
This work was supported by grants from the National Natural Science Foundation of China (No.82003271,82103358,82073294,81802555), Education Department of Guangdong Province (2020KTSCX102), Guangdong Special Fund Project of Fundamental and Applied Research (2022A1515010344), The Guangzhou Planned Project of Science and Technology (202102010021, 202102010152), Science and Technology Project of Zhongshan Municipality (2019B1062).
Author information
Authors and Affiliations
Contributions
J.L. and Y.L. conceived of the study. J.L., Y.L. and W.Y. contributed data analysis. Z.C., J.C. and Z.L. collected samples and generated data. J.D. and W.Z. contributed project oversight. Y.H. and Z.L. wrote figure legends. Z.C. and H.L. performed the cell experiments; Q.L. and C.Z. performed the animal experiments. Y.L., J.L., C.C. and W.Y. wrote the paper. All authors participated in writing the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they do not have any conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, Y., Yin, W., Cai, Z. et al. N6-methyladenosine modified lncRNAs signature for stratification of biochemical recurrence in prostate cancer. Hum. Genet. (2023). https://doi.org/10.1007/s00439-023-02603-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00439-023-02603-8