Abstract
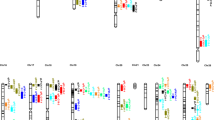
Cotton is the most important fiber crop in the world. Asiatic cotton (Gossypium arboreum, genome A2) is a diploid cotton species producing spinnable fibers and important germplasm for cotton breeding and a significant model for fiber biology. However, the genetic map of Asiatic cotton has been lagging behind tetraploid cottons, as well as other stable crops. This study aimed to construct a high-density SNP genetic map and to map QTLs for important yield and fiber quality traits. Using a recombinant inbred line (RIL) population and genome resequencing technology, we constructed a high-density genetic map that covered 1980.17 cM with an average distance of 0.61 cM between adjacent markers. QTL analysis revealed a total of 297 QTLs for 13 yield and fiber quality traits in three environments, explaining 5.0–37.4% of the phenotypic variance, among which 75 were stably detected in two or three environments. Besides, 47 QTL clusters, comprising 131 QTLs for representative traits, were identified. Our works laid solid foundation for fine mapping and cloning of QTL for yield and fiber quality traits in Asiatic cotton.

Similar content being viewed by others
Availability of data and material
Sequencing data related to this study has been uploaded to NCBI SRA database, which can be accessed through series of SRA numbers PRJNA649393.
Code availability
Not applicable.
References
Badigannavar A, Myers GO (2015) Construction of genetic linkage map and QTL analysis for fiber traits in diploid cotton (Gossypium arboreum x Gossypium herbaceum). J Cotton Sci 19:15–26
Brubaker CL, Paterson AH, Wendel JF (1999) Comparative genetic mapping of allotetraploid cotton and its diploid progenitors. Genome 42:184–203
Chen ZJ, Scheffler BE, Dennis E, Triplett BA, Zhang T, Guo W, Chen X, Stelly DM, Rabinowicz PD, Town CD, Arioli T, Brubaker C, Cantrell RG, Lacape JM, Ulloa M, Chee P, Gingle AR, Haigler CH, Percy R, Saha S, Wilkins T, Wright RJ, Van Deynze A, Zhu Y, Yu S, Abdurakhmonov I, Katageri I, Kumar PA, Mehboob Ur R, Zafar Y, Yu JZ, Kohel RJ, Wendel JF, Paterson AH (2007) Toward sequencing cotton (Gossypium) genomes. Plant Physiol 145:1303–1310
Chen Z, Wang B, Dong X, Liu H, Ren L, Chen J, Hauck A, Song W, Lai J (2014) An ultra-high density bin-map for rapid QTL mapping for tassel and ear architecture in a large F(2) maize population. BMC Genom 15:433
Cingolani P, Platts A, le Wang L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (austin) 6:80–92
Desai A, Chee PW, Rong J, May OL, Paterson AH (2006) Chromosome structural changes in diploid and tetraploid A genomes of Gossypium. Genome 49:336–345
Diouf L, Magwanga RO, Gong WF, He SP, Pan ZE, Jia YH, Kirungu JN, Du XM (2018) QTL mapping of fiber quality and yield-related traits in an intra-specific upland cotton using genotype by sequencing (GBS). Int J Mol Sci 19:441
Du X, Huang G, He S, Yang Z, Sun G, Ma X, Li N, Zhang X, Sun J, Liu M, Jia Y, Pan Z, Gong W, Liu Z, Zhu H, Ma L, Liu F, Yang D, Wang F, Fan W, Gong Q, Peng Z, Wang L, Wang X, Xu S, Shang H, Lu C, Zheng H, Huang S, Lin T, Zhu Y, Li F (2018) Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat Genet 50:796–802
Fan LP, Wang LP, Wang XY, Zhang HY, Zhu YF, Guo JY, Gao WW, Geng HW, Chen QJ, Qu YY (2018) A high-density genetic map of extra-long staple cotton (Gossypium barbadense) constructed using genotyping-by-sequencing based single nucleotide polymorphic markers and identification of fiber traits-related QTL in a recombinant inbred line population. BMC Genom 19:489
Hovav R, Udall JA, Chaudhary B, Hovav E, Flagel L, Hu G, Wendel JF (2008) The evolution of spinnable cotton fiber entailed prolonged development and a novel metabolism. PLOS Genet 4:e25
Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q, Li W, Guo Y, Deng L, Zhu C, Fan D, Lu Y, Weng Q, Liu K, Zhou T, Jing Y, Si L, Dong G, Huang T, Lu T, Feng Q, Qian Q, Li J, Han B (2011) Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet 44:32–39
Huang C, Shen C, Wen T, Gao B, Zhu D, Li X, Ahmed MM, Li D, Lin Z (2018) SSR-based association mapping of fiber quality in upland cotton using an eight-way MAGIC population. Mol Genet Genom 293:793–805
Huang G, Wu Z, Percy RG, Bai M, Li Y, Frelichowski JE, Hu J, Wang K, Yu JZ, Zhu Y (2020) Genome sequence of Gossypium herbaceum and genome updates of Gossypium arboreum and Gossypium hirsutum provide insights into cotton A-genome evolution. Nat Genet 52:516–524
Huang G, Huang JQ, Chen XY, Zhu YX (2021) Recent advances and future perspectives in cotton research. Annu Rev Plant Biol 72:437–462
Kebede H, Burow G, Dani RG, Allen RD (2007) A-genome cotton as a source of genetic variability for Upland cotton (Gossypium hirsutum). Genet Resour Crop Ev 54:885–895
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eugenic 12:172–175
Kulkarni VN, Khadi BM, Maralappanavar MS, Deshapande LA, Narayanan SS (2009) The worldwide gene pools of Gossypium arboreum L. and G. herbaceum L., and their improvement. In: Paterson AH (ed) Genetics and genomics of cotton. Springer US, New York, pp 69–97
Lander E, Kruglyak L (1995) Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 11:241–247
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Genome Project Data Processing S (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079
Li F, Fan G, Lu C, Xiao G, Zou C, Kohel RJ, Ma Z, Shang H, Ma X, Wu J, Liang X, Huang G, Percy RG, Liu K, Yang W, Chen W, Du X, Shi C, Yuan Y, Ye W, Liu X, Zhang X, Liu W, Wei H, Wei S, Huang G, Zhang X, Zhu S, Zhang H, Sun F, Wang X, Liang J, Wang J, He Q, Huang L, Wang J, Cui J, Song G, Wang K, Xu X, Yu JZ, Zhu Y, Yu S (2015) Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotechnol 33:524–530
Li C, Dong Y, Zhao T, Li L, Li C, Yu E, Mei L, Daud MK, He Q, Chen J, Zhu S (2016) Genome-wide SNP linkage mapping and QTL analysis for fiber quality and yield traits in the upland cotton recombinant inbred lines population. Front Plant Sci 7:1356
Liu D, Ma C, Hong W, Huang L, Liu M, Liu H, Zeng H, Deng D, Xin H, Song J, Xu C, Sun X, Hou X, Wang X, Zheng H (2014) Construction and analysis of high-density linkage map using high-throughput sequencing data. PLoS ONE 9:e98855
Liu R, Gong J, Xiao X, Zhang Z, Li J, Liu A, Lu Q, Shang H, Shi Y, Ge Q, Iqbal MS, Deng X, Li S, Pan J, Duan L, Zhang Q, Jiang X, Zou X, Hafeez A, Chen Q, Geng H, Gong W, Yuan Y (2018) GWAS analysis and QTL identification of fiber quality traits and yield components in upland cotton using enriched high-density SNP markers. Front Plant Sci 9:1067
Ma XX, Zhou BL, Lu YH, Guo WZ, Zhang TZ (2008) Simple sequence repeat genetic linkage maps of A-genome diploid cotton (Gossypium arboreum). J Integr Plant Biol 50:491–502
Maqbool A, Abbas W, Rao AQ, Irfan M, Zahur M, Bakhsh A, Riazuddin S, Husnain T (2010) Gossypium arboreum GHSP26 enhances drought tolerance in Gossypium hirsutum. Biotechnol Prog 26:21–25
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Said JI, Knapka JA, Song MZ, Zhang JF (2015a) Cotton QTLdb: a cotton QTL database for QTL analysis, visualization, and comparison between Gossypium hirsutum and G-hirsutum x G-barbadense populations. Mol Genet Genom 290:1615–1625
Said JI, Song MZ, Wang HT, Lin ZX, Zhang XL, Fang DD, Zhang JF (2015b) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum x G. barbadense populations. Mol Genet Genom 290:1003–1025
Shang L, Abduweli A, Wang Y, Hua J, Jenkins J (2016) Genetic analysis and QTL mapping of oil content and seed index using two recombinant inbred lines and two backcross populations in Upland cotton. Plant Breed 135:224–231
Tahir MS, Khan NUI, Sajid-Ur-Rehman S (2011) Development of an Interspecific Hybrid (Triploid) by Crossing Gossypium hirsutum and G. arboreum. Cytologia 76:193–199
Thyssen GN, Jenkins JN, McCarty JC, Zeng L, Campbell BT, Delhom CD, Islam MS, Li P, Jones DC, Condon BD, Fang DD (2019) Whole genome sequencing of a MAGIC population identified genomic loci and candidate genes for major fiber quality traits in upland cotton (Gossypium hirsutum L.). Theoret Appl Genet 132:989–999
Van Ooijen JW (2009) MapQTL® 6.0. Sofware for the mapping of quantitative trait loci in experimental populations of diploid species. Kyazma K.V. Wageningen, Berlin
van Os H, Stam P, Visser RG, van Eck HJ (2005) SMOOTH: a statistical method for successful removal of genotyping errors from high-density genetic linkage data. Theoret Appl Genet 112:187–194
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang H, Huang C, Guo H, Li X, Zhao W, Dai B, Yan Z, Lin Z (2015) QTL mapping for fiber and yield traits in upland cotton under multiple environments. PLoS ONE 10:e0130742
Wang M, Tu L, Yuan D, Zhu SC, Li J, Liu F, Pei L, Wang P, Zhao G, Ye Z, Huang H, Yan F, Ma Y, Zhang L, Liu M, You J, Yang Y, Liu Z, Huang F, Li B, Qiu P, Zhang Q, Zhu L, Jin S, Yang X, Min L, Li G, Chen LL, Zheng H, Lindsey K, Lin Z, Udall JA, Zhang X (2019a) Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nat Genet 51:224–229
Wang W, Sun Y, Yang P, Cai X, Yang L, Ma J, Ou Y, Liu T, Ali I, Liu D, Zhang J, Teng Z, Guo K, Liu D, Liu F, Zhang Z (2019b) A high density SLAF-seq SNP genetic map and QTL for seed size, oil and protein content in upland cotton. BMC Genom 20:599
Wang F, Zhang J, Chen Y, Zhang C, Gong J, Song Z, Zhou J, Wang J, Zhao C, Jiao M, Liu A, Du Z, Yuan Y, Fan S, Zhang J (2020) Identification of candidate genes for key fibre-related QTLs and derivation of favourable alleles in Gossypium hirsutum recombinant inbred lines with G. barbadense introgressions. Plant Biotechnol J 18:707–720
Wendel JF, Brubaker C, Alvarez I, Cronn R, Stewart JM (2009) Evolution and natural history of the cotton genus. In: Paterson AH (ed) Genetics and genomics of cotton. Springer US, New York, pp 3–22
Yu JZ, Ulloa M, Hoffman SM, Kohel RJ, Pepper AE, Fang DD, Percy RG, Burke JJ (2014) Mapping genomic loci for cotton plant architecture, yield components, and fiber properties in an interspecific (Gossypium hirsutum L. × G. barbadense L.) RIL population. Mol Genet Genomics 289:1347–1367
Zhang ZS, Xiao YH, Luo M, Li XB, Luo XY, Hou L, Li DM, Pei Y (2005) Construction of a genetic linkage map and QTL analysis of fiber-related traits in upland cotton (Gossypium hirsutum L.). Euphytica 144:91–99
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu B, Liu C, Wang S, Pan M, Wang Y, Wang D, Ye W, Chang L, Zhang W, Song Q, Kirkbride RC, Chen X, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu X, Zhang H, Wu H, Zhou L, Mei G, Chen S, Tian Y, Xiang D, Li X, Ding J, Zuo Q, Tao L, Liu Y, Li J, Lin Y, Hui Y, Cao Z, Cai C, Zhu X, Jiang Z, Zhou B, Guo W, Li R, Chen ZJ (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33:531–537
Zhang S, Feng L, Xing L, Yang B, Gao X, Zhu X, Zhang T, Zhou B (2016a) New QTLs for lint percentage and boll weight mined in introgression lines from two feral landraces into Gossypium hirsutum acc TM-1. Plant Breeding 135:90–101
Zhang S, Zhu X, Feng L, Gao X, Yang B, Zhang T, Zhou B (2016b) Mapping of fiber quality QTLs reveals useful variation and footprints of cotton domestication using introgression lines. Sci Rep 6:31954
Zhu MS, Liu DL, Liu WG, Li D, Liao YL, Li JH, Fu CY, Fu FH, Huang HJ, Zeng XQ, Ma XZ, Wang F (2017a) QTL mapping using an ultra-high-density SNP map reveals a major locus for grain yield in an elite rice restorer R998. Sci Rep 7:10914
Zhu T, Liang C, Meng Z, Sun G, Meng Z, Guo S, Zhang R (2017b) CottonFGD: an integrated functional genomics database for cotton. BMC Plant Biol 17:101
Zou XY, Gong JW, Duan L, Jiang X, Zhen Z, Fan SM, Ge Q, Liu AY, Gong WK, Li JW, Shi YZ, Wang YL, Fan LQ, Liu RX, Lei K, Zhang Q, Shang HH, Yuan YL (2018) High-density genetic map construction and QTL mapping for fiber strength on Chr24 across multiple environments in a CCRI70 recombinant inbred lines population. Euphytica 214:102
Funding
This study was partially funded by the Genetically Modified Organisms Breeding Major Project of China (2016ZX08005005-001 to Y.H.X.), the National Natural Science Foundation of China (31871539 and U2003209 to Y.H.X.) and Chongqing Science and Technology Commission (cstc2018jscx-msybX0382 to Y.H.X.).
Author information
Authors and Affiliations
Contributions
YX conceived the study, participated in its design and modified the manuscript. YL contributed to data analysis and manuscript writing. YL, TM, LR, JZ, CW, YD, YW, ZZ collected the data from the field; YX and AL constructed RIL population. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Bing Yang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, Y., Mo, T., Ran, L. et al. Genome resequencing-based high-density genetic map and QTL detection for yield and fiber quality traits in diploid Asiatic cotton (Gossypium arboreum). Mol Genet Genomics 297, 199–212 (2022). https://doi.org/10.1007/s00438-021-01848-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01848-0