Abstract
Mastophorus muris (Gmelin, 1790) is a globally distributed parasitic nematode of broad range mammals. The taxonomy within the genus Mastophorus and the cryptic diversity among the genus are controversial among taxonomists. This study provides a detailed morphological description of M. muris from Mus musculus combined with a molecular phylogenetic approach. Moreover, descriptions and molecular data of M. muris from non-Mus rodents and wildcats complement our findings and together provide new insights into their taxonomy. The analysis of M. muris was based on light microscopy and scanning electron microscopy. The morphological description focused on the dentition pattern of the two trilobed pseudolabia. Additionally, we described the position of the vulva, arrangement of caudal pairs of papillae, spicules and measured specimens from both sexes and the eggs. For the molecular phylogenetic approach, we amplified the small subunit ribosomal RNA gene and the internal transcribed spacer, and the cytochrome c oxidase subunit 1. Mastophorus morphotypes based on dentition patterns and phylogenetic clustering indicate a subdivision of the genus in agreement with their host. We recognize two groups without a change to formal taxonomy: One group including those specimens infecting Mus musculus, and the second group including organisms infecting non-Mus rodents. Our genetic and morphological data shed light into the cryptic diversity within the genus Mastopohorus. We identified two host-associated groups of M. muris. The described morphotypes and genotypes of M. muris allow a consistent distinction between host-associated parasites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mastophorus muris (Gmelin, 1790) is an euryxenous nematode belonging to the superfamily Spiruroidea (Spirocercidae: Mastophorinae), with a heteroxenous life cycle (Quentin 1970). Intermediate hosts are arthropods of the orders Orthoptera, Dermaptera, Blattodea and Siphonaptera (Grzybek et al. 2015; Quentin 1970). Different species of small rodents (Chitwood 1938; Skrjabin 1961; Rojas and Digiani 2003; Grzybek et al. 2015; Julius et al. 2018; Neupane et al. 2020) including Mus musculus (Chitwood 1938; Skrjabin 1961; Kataranovski et al. 2008; Baird et al. 2012) are described as definitive hosts but felines (Skrjabin 1961; Torres et al. 1998), canines (Chitwood 1938; Skrjabin 1961) and even marsupials (Skrjabin 1961; Smales 1995; Smith and Kinsella 2011) have also been reported as hosts.
Within the genus Mastophorus, a subdivision into varieties associated with certain hosts has been controversially discussed, and different morphological features have been proposed for this classification (Chitwood 1938; Wertheim 1962; Rojas and Digiani 2003). A major contribution to the classification of Mastophorus was provided by Chitwood (1938), who suggested a subdivision of the genus into two varieties based on a single morphological characteristic and host preference: (1) M. muris var. muris (large denticles, Rattus norvegicus and Felis catus) wherein Mus specimens (intermediate in teeth length) were included for simplicity and (2) M. muris var. ascaroides (smaller denticles, Geomyoidae, Cricetidae and Canis latrans) (Chitwood 1938).
The genus Mastophorus has a complex taxonomic relationship with the closely related genus Protospirura (Chitwood 1938; Read and Millemann 1953; Skrjabin 1961; Wertheim 1962; Smales 1995; Rojas and Digiani 2003). The two genera of the superfamily Spiruroidea are morphologically distinguishable based on two traits: the nature of the dentition on the pseudolabia and the proximal position of the vulva with regard to the middle of the body (Chitwood 1938). The shape of the pharynx, egg measurements and spicule characteristics have been suggested as additional differentiation characters (Chitwood 1938; Wertheim 1962) as well as ontogenetic characteristics (Quentin 1970). Nevertheless, despite the discrepancies in the classification of Mastophorus and Protospirura (Rojas and Digiani 2003; Smales et al. 2009), there is a cryptic morphological diversity within the genus Mastophorus that has not been fully explored.
Here, we compare Mastophorus muris specimens from different hosts (Mus musculus, Felis silvestris silvestris, Myodes glareolus and Apodemus flavicollis) to examine the differentiation of the genus in varieties according to their host range. We provide general morphological descriptions and molecular data of specimens from different hosts. To differentiate the specimen from our study, we focused on the detailed description of the dentition pattern besides phylogenetic analyses.
Material and methods
Sample collection
A total of 567 house mice (Mus musculus) collected in Brandenburg (Germany), during annual field trips in autumn 2016 to 2018, were used for the present study (capture permit No. 2347/35/2014). Mice were dissected for inspection of helminths in the body cavity and gastrointestinal tract. Specimens from each host were individually collected and preserved in 10% Neutral-Buffered Formalin solution for detailed morphological description and in 70% (v/v) ethanol for DNA extraction and stored at room temperature until further analysis. Feces were collected and stored in a 2.5% (w/v) solution of potassium dichromate (K2Cr2O7) at 4 °C until further microscopic observations of parasite eggs.
Four M. muris specimens from non-Mus rodents were collected in Berlin in 2010, two from the bank vole (Myodes glareolus) and two from the yellow-necked mouse (Apodemus flavicollis) (one female and one male from each host) (Maaz et al. 2018), stored in 70% (v/v) ethanol and used for morphological description and DNA extraction. Morphological characteristics and extracted DNA from M. muris specimens collected from the wildcat (Felis silvestris silvestris, N = 2) were integrated in further analyses.
Voucher specimens for M. muris from Mus were deposited in the Natural History Museum in Berlin, Germany in the department “Vermes” under specimens numbers E.7635 – E.7639.
Morphological analysis
Morphological description of specimens (N = 125, 78 females and 47 males) was performed following taxonomic keys of parasitic nematodes (Hall 1916; Skrjabin 1961; Chabaud 1975; Sutton 1989). Morphometric data of the length, width and vulva position were recorded with an Olympus SZ61 stereo microscope. To visualize the structure of the spicules, male specimens fixed in ethanol were treated with a potassium hydroxide solution 10% (w/v) for 3 days at room temperature. Eggs were collected in a flotation of feces with a saturated salt solution (Jarquín-Díaz et al. 2019). Micrographs of eggs (N = 30, 400 × magnification) and spicules (N = 6, 100 × magnification) were taken with an Axioplan Carl-Zeiss light microscope and measured using Adobe Photoshop CC v 14.2.1.
Specimens (N = 16, eight females and four males from Mus, two females and two males from non-Mus) for scanning electron microscopy (SEM) were first fixed in 2% (v/v) paraformaldehyde and 2.5% (v/v) glutaraldehyde in phosphate buffer (pH = 7.4) and then treated with 2% (v/v) osmium tetroxide. They were rinsed in distilled water, dehydrated in an alcohol series, critical point dried in carbon dioxide (BAL-TEC CPD030 Critical Point Dryer) and finally gold-coated (BAL-TEC SCD005 Sputter Coater). The samples were examined using an SEM LEO 1430 (Zeiss) and the associated SmartSEM V06.00 operating software, and the resulting scanning electron micrographs were post-processed using Adobe Photoshop CC v 14.2.1.
Data analysis
Parasite prevalence, abundance and intensity as defined by Lafferty et al. (1997) were determined. All calculations were performed in R (R Core Team 2008). For prevalence, 95% confidence intervals were calculated using Sterne’s exact method (Rózsa et al. 2000), using the package “epiR” (Stevenson et al. 2018).
DNA extraction
The morphologically less informative part of a single worm specimen was mechanically disrupted with a micro pestle in 20 µL of nuclease-free water (the other part was saved for morphological assessment). Genomic DNA was extracted using the innuPREP DNA Mini Kit (Analytik Jena AG, Jena, Germany), following the protocol of the manufacturer for tissue samples and rodent tails. Adjustments were made within the lysis step (30 µL proteinase K and 1-h incubation time) and the elution step (adding 50 µL elution buffer with one repetition). Modifications to lysis conditions were applied (30 µL of protein kinase and 1 h at 50 °C incubation). The DNA was eluted twice in a final volume of 50 µL.
PCR amplification
Previously published primer pairs specific for nematodes were used, which target partial sequences of the nuclear genome: 18S rDNA (18S) (Floyd et al. 2005), the internal transcribed spacer (ITS) region (including ITS1, 5.8S and ITS2) (Gasser and Hoste 1995) and partial sequence of the mitochondrial cytochrome c oxidase subunit 1 (CO1) gene (Bowles et al. 1993; Casiraghi et al. 2001; Blaxter 2004). In addition, a primer pair aiming to complete the 18S region was designed, based on the sequence of M. muris from wildcat (MG818763) and the already amplified regions using Geneious R6 v. 6.1.8 (https://www.geneious.com) (Kearse et al. 2012) (Supplementary Table 1). PCR was performed using ThermoScientific DreamTaq DNA Polymerase (Thermo Fisher Scientific Inc.) as detailed in Supplementary Table 2. Amplified PCR products with the expected size were purified with the SAP-Exo Kit (Jena Bioscience GmbH, Jena, Germany), following the specifications of the manufacturer and sequenced in both directions by LGC (LGC Genomics GmbH, Berlin, Germany).
Consensus sequences for each gene and specimen were generated by assembly of forward and reverse sequencing reads, and further alignment of overlapping regions between amplicons was generated with different primer pairs by assembling reads with reference sequences (18S MG818763 and CO1 AJ537512) in Geneious. The nearly complete 18S region (~ 1670 bp), sequences with ~ 850 bp for the CO1 and ~ 1000 bp for ITS for Mastophorus specimens from Mus and Apodemus (none for ITS region) were submitted to NCBI GenBank with the accession numbers: 18S [MN086286–MN086291], CO1 [MK867474–MK867480] and ITS [MK829001–MK829007]. Genetic data of M. muris from Myodes could not be obtained.
Phylogenetic analyses
Datasets for each gene were generated individually, including all available sequences in the GenBank from Mastophorus and closely related organisms only from the superfamily Spiruroidea. Close related sequences MT512662 (Protospirura sp.—CO1), KT894811, KT894812 (Protospirura numidica—18S), JQ771745 and JQ771746 (Spiruridae sp.—18S) were excluded from any phylogenetic analysis due to lack or questionable morphological identification and taxonomic assignment. Relevant available information from those sequences regarding host and geographical origin, length and authors are listed in Supplementary Table 3. The individual sequence datasets for 18S, CO1 and ITS were aligned using the profile-to-profile method implemented within the R package DECIPHER v.2.22 (Wright 2015, 2020). The CO1 dataset was aligned using the codon-based algorithm. For all datasets, missing data was indicated in the sequences as “?”. The R package Phangorn v. 2.11.1 (Schliep 2011; Schliep et al. 2016) was used to determine the substitution model with the best fit for each alignment (appropriated sequence evolution models for each dataset—18S: TPM2 + I, ITS: HKY + G, CO1: TIM3 + G). Phylogenetic estimation was done by maximum likelihood (ML) implemented in Phangorn v. 2.11.1 (Schliep 2011; Schliep et al. 2017) with 1000 bootstrap replicates and Bayesian inference (BI) implemented in MrBayes v. 3.2.7 (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003) using two heated and two cold chains sampled every 100 generations for a total of 1,000,000 generations with an average standard deviation of split frequencies below 0.01 and using a relative burn in of 25% for diagnostic. Phylogenetic inference accounted for missing data.
For all genetic analyses, sequences from Dirofilaria were used as an outgroup for rooting (Supplementary Table 3). Visualization and editing of the phylogenetic trees were conducted in Figtree v.1.4.4 (http://tree.bio.ed.ac.uk/software/figtree/) and Inkscape v. 0.92 (https://inkscape.org).
Results
Occurrence of Mastophorus muris in house mice
A total of 207 M. muris were collected from 21 of 567 investigated mice, corresponding to a prevalence of 3.7% (95% confidence interval (CI): 2.35–5.62). Infected mice were from 14 different localities in North-Eastern Germany. In all cases, worms were located in the stomach of the host. The maximum intensity was 46 M. muris in one host, mean intensity was 9.86 (95% CI: 6.14–16.27) and the abundance was 0.37 (95% CI: 0.19–0.75).
Morphological descriptions of M. muris from M. musculus
Morphological observations and measurements for our Mastophorus specimens from SEM and light microscopy are summarized in Table 1. The body surface of all specimens shows a transversal striation, which is attenuated towards the anterior end, showing a circular mouth opening surrounded by two trilobed pseudolabia (Fig. 1A, D). Each pseudolabium is composed of one lateral and two submedian lobes. The lateral lobe is large, square-shaped (Fig. 1B, E) and framed by two smaller, slender and more tapered submedian lobes (Fig. 1C, F). Four cephalic papillae are located at the base of the pseudolabia, one pair per labium (Fig. 1A, D). At the distal margin of each lobe, “denticle-like” structures of variable size and irregular shape are visible (Fig. 1B–F). Larger denticles protrude at both edges and in the middle of each lobe (Fig. 1B–F). The denticles are located with different membranes.
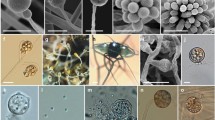
Scanning electron micrographs of Mastophorus muris specimens from Mus musculus. A En face view of the mouth opening, schematized in (D) showing two trilobed pseudolabia each consisting of one lateral (ll) and two submedian (sml) lobes, with two cephalic papillae (cep) located at its base. Note different shapes of lobes: lateral lobes are square-shaped (B, E), framed by two smaller, slender submedian lobes (C, F). Scheme (D) illustrates the visible dentition (marked in B, C, E and F) at the distal margin of each lobe. Dentition consists of a large central tooth (1), smaller median denticles (2) and a large tooth on each edge (3). Variations of the dentition of the pseudolabia are shown from two specimens (specimen AA0256 in B and C, AA0351 in E and F; see Supplementary Table 4 for list of samples and further details). A separation in two membranes is visible at the toothed distal margin of the lateral lobe (B). cep—cephalic papilla, ll—lateral lobe, sml—submedian lobe
At the distal margin of the lateral lobe, the separation into an outer membrane (Fig. 1B) that bears the large denticles at the edges (3) and an inner membrane with the large central tooth (1) and a variable number of smaller denticles (2) is visible (Fig. 1B). The number of smaller denticles varies between lobes and from specimen to specimen (Fig. 1B, C, E, F, Supplementary Table 4). For the Mastophorus specimens from Mus, a general dentition pattern can be specified with: 1–(2 + n)–1–(2 + n)–1.
Females (N = 78) are 9.53–39.16 (26.43 ± 7.01) mm long and 0.30–1.78 (1.13 ± 0.34) mm wide. The vulva (Fig. 2A) is a transverse fissure located anterior to the middle of the body, in a position around 34.10–42.38% (N = 10) of the total body length. The posterior end of female specimens is rounded, and the cloaca can be observed (Fig. 2B). Mastophorus muris eggs (N = 30, fecal flotations from different hosts) are oval and thick-shelled (Fig. 3A), with a length of 0.054–0.064 (0.058 ± 0.002) mm and a width of 0.033–0.036 (0.034 ± 0.001) mm and contain a first-stage larva.
Scanning electron micrographs of female (A–B) and male (C–D) Mastophorus muris specimens from Mus musculus. Ventral view shows the vulva located anterior to the middle of the body (A). Ventro-lateral view of the female tail showing cloaca and a rounded tip (B). Lateral view of the coiled male tail showing a wide caudal ala (indicated with an arrow) (C). At the ventral view of the tail longitudinal striations, cuticular modifications (marked with a circle), the cloaca, the phasmidial orifices, distal sessile caudal papillae and six pairs of pre-/postcloacal pedunculate papillae are present (D). v—vulva, c—cloaca, prcp—precloacal papillae, pocp—postcloacal papillae, Ph—phasmidial orifices, sessile caudal papillae—scp
Mature egg (A) and the posterior end of a male specimen (B) of Mastophorus muris from Mus musculus, light microscopy. A Thick-shelled Mastophorus muris egg shows a first-stage larva detected in the floated feces. B Lateral view of the coiled tail showing spicules of different length and width (alkaline treatment)
Males (N = 47) are 10.10–27.96 (18.61 ± 3.24) mm long and 0.38–0.94 (0.69 ± 0.13) mm wide. Their posterior end is longer and more coiled compared to female specimens, and has a wide caudal ala (Fig. 2C, indicated with an arrow). The posterior surface is heavily ornamented ventrally with longitudinal striations and cuticular modifications (Fig. 2D). At the posterior end, in total six pairs of pedunculate caudal papillae are present: four pre-cloacal pairs and two post-cloacal pairs (Fig. 2D). At the anterior lip of the cloacal aperture, an unpaired median papilla was observed (Fig. 2D). Additionally, a sessile papillae and phasmidial orifices are observed (Fig. 2D). We observed two unequal spicules become visible (Fig. 3B), the larger right spicule is 0.910–1.519 (1.217 ± 0.151) mm long and 0.016–0.030 (0.024 ± 0.005) mm wide, and the smaller left spicule is 0.639–1.161 (0.974 ± 0.144) mm long and 0.012–0.026 (0.020 ± 0.004) mm wide.
Morphological description of M. muris from non-Mus rodents
One female and one male specimen from A. flavicollis and M. glareolus were analyzed, using light microscopy and SEM. General structures of the body surface, the apical and posterior region were found to be indistinguishable between specimens from different rodent hosts (Fig. 4). Measurements for body size and vulva position are consistent with the previous descriptions of M. muris (Table 1), including our own description of M. muris from M. musculus.
Scanning electron micrographs of Mastophorus muris specimens from Apodemus flavicollis (A–C) and from Myodes glareolus (D–F). Face view of the mouth opening, which is surrounded by two trilobed pseudolabia each with one lateral (ll) and two submedian (sml) lobes and with four cephalic papillae (cep) located on their base (A and D). Visible dentition is marked with (1) for a large median tooth and with (2) for smaller denticles, variable in numbers (E). A lateral view of the coiled posterior end of male specimens shows the cloaca (c) and six pairs of pre-/postcloacal papillae (prcp/pocp) (C and F). The extended spicules (sp) are visible in the lateral-ventral view of the tail of one Myodes specimen (F). c—cloaca, prcp—precloacal papillae, pocp—postcloacal papillae, sp—spicule
Dentition pattern could be observed in specimens from Apodemus and Myodes hosts (Fig. 4A–D). No large denticles were detected at the edge of the lobes (Fig. 4E), and only the smaller submedian lobes show a large central tooth (1) framed by two or three smaller denticles (2) on each side (dentition pattern 3–1–3). The central lobe of the specimen from Myodes shows a dentition with seven to nine denticles of unequal medium size (Fig. 4E).
Morphological description of M. muris from F. silvestris
From wildcat carcasses, two female M. muris were isolated, one from the intestine and one from the pulmonary vessel of the lung. The Mastophorus from wildcats had a circular mouth opening and non-compressed pharynx (not shown) and presented the following traits under a light microscope (Supplementary Fig. S1): (1) vulva position at the anterior part of the body (31.93%); (2) dentition with a central big tooth and a variable uneven number of smaller denticles on the trilobed pseudolabia; (3) narrow oval eggs (N = 6, 0.053 × 0.030 mm). Morphological traits and measurements were in agreement with previously published data (Table 1).
Dentition of M. muris in comparison to previous reports
Dentition patterns are variable between specimens (Supplementary Table 4). Nevertheless, we were able to generalize the following dentition pattern for specimens from Mus hosts per lobe (Fig. 5): a large central tooth (1), large denticles on the edges of each lobe (3) and a variable number of smaller denticles in between (2). This can be expressed as the formula 1–(2 + n)–1–(2 + n)–1 (the dash separating large denticles and smaller denticles in brackets). In contrast, non-Mus specimens (from Myodes, Rattus and Felis) (Wertheim 1962) exhibit a large central tooth (1), a variable number of smaller denticles in between (2) and no large denticles at the edges of each lobe. For M. muris from Graomys (Fig. 5, Rojas and Digiani 2003) a fixed pattern was reported, always having three smaller denticles between large single denticles on one lobe (1–3–1–3–1).
Illustration of the variants of dentition patterns of Mastophorus muris obtained from different hosts. The schema illustrates the dentition consisting of a large central tooth (1), various numbers of smaller median denticles (2) and a large denticles on each edge (3) if applicable (dashed lines represent variability). In the line below, the dentition formula for each host is shown, either for all three lobes per pseudolabia [][][] or one representative lobe []. The subdivision of the genus Mastophorus by Chitwood is shown in gray and associated with our assignment in Mastophorus muris associated with Mus (pink) and Mastophorus muris associated with non-Mus hosts (purple) for specimens from Felis, Myodes and Rattus. * Previous published data for Rattus and Graomys
Genetic differences between M. muris from different hosts
To assess the phylogenetic pattern of M. muris from house mice to previously investigated specimens, we inferred phylogenetic trees, using the most commonly reported genetic markers for these nematodes.
A phylogenetic tree for 18S sequences was based on 16 sequences (1692–1748 bp), including M. muris and other nematodes (Spirurina) (Fig. 6A). Sequences from specimens from Mus (N = 6) formed a well-supported clade, separating them from M. muris specimens, isolated from A. flavicollis (N = 1), F. silvestris silvestris (N = 1) and Rattus norvegicus (N = 1). Gongylonema sequences were recovered in a clade with the sister group being a sequence deposited as Protospirura sp. to GenBank (accession number: KY462830.1). The latter sequence obtained from a specimen isolated from Mastomys coucha in South Africa is nearly identical (99%) to a partial (631 bp) 18S sequence of P. muricola (KP760162) from a gorilla of Central African Republic, but has a sequence identity of only 95% to P. numidica (KT894812, KT894811) not included for our analysis due to its controversial morphological assignment (Costa et al. 2018).
Phylogenetic tree based on 18S (A), CO1 (B) and ITS (C) sequences. Analyses include reference sequences (marked with an (*) asterisk) from Spiruroidea and as outgroup Dirofilaria spp. isolates. Mastophorus sequences are found in two distinct clades, one with isolates from Mus (pink) and a second clade with isolates from wildcat (black), rat (blue), Apodemus (green) and Sigmodon (brown) colored in purple. Bayesian posterior probabilities followed by bootstrap values are displayed on the branches and the substitution rate per site in the bottom scale bar
A phylogenetic tree based on CO1 sequences (Fig. 6B) was inferred from 24 sequences (369–858 bp), 10 Mastophorus sequences from different hosts (six Mus, one Apodemus, three Sigmodon, one Felis and one from a wild rat), 11 further sequences of representatives of the superfamily Spiruroidea and 3 Dirofilaria sp. sequences. In agreement with the 18S phylogenetic tree, M. muris from house mice formed a distinct clade with high support and sequence identities of 99.6–100% (795 bp). Partial CO1 sequences of M. muris isolates from Apodemus and Felis are 99.6% (801 bp) identical, thus the same Mastophorus species. We observed sequence identities of only around 87% (795 bp) between specimens from Mus and non-Mus hosts (Felis, Apodemus, Rattus and Sigmodon).
The phylogenetic tree for ITS sequences (Fig. 6C) was based on 17 sequences (490–1351 bp): five sequences of the superfamily Spiruroidea, one sequence from M. muris from wildcat and our six sequences from Mus were included.
All markers support a close relationship of Mastophorus sequences from Mus musculus in one clade, contrasted with a second clade consisting of all other available Mastophorus sequences from other hosts. A genetic distinction between two Mastophorus groups according to the hosts they were isolated from is overall moderately to well supported.
Discussion
We provide a detailed morphological description of M. muris focused on the dentition pattern of the pseudolabia as a character allowing the distinction of host-associated varieties. We compare this novel morphological distinction with published descriptions and link it with genetic data following the principles of integrative taxonomy (Blaxter 2004; Dayrat 2005; De Queiroz 2007). Genetic data in the present study confirm the phylogenetic placement of M. muris varieties as sister group to Protospirura sequences available in databases.
A distinction based on morphology between M. muris specimens from different hosts was only apparent in the dentition pattern on the lobes of the pseudolabia. We had additionally evaluated the size of the worms, the size of spicules, the lobe substructures of the pseudolabiae, the position of the vulva in females and the arrangement of papillae on the tail of males. The suitability of the dentition pattern as a distinguishing morphological character had been proposed in previous reports (Chitwood 1938; Wertheim 1962; Rojas and Digiani 2003); as a consequence, other morphological characters also suggested in nematodes systematics such as distances to nerve ring, excretory pore and deirids from the cephalic end, the lengths of the pharynx, muscular and glandular esophagus, and tail length etc. were not considered in the present study, which might represent a limitation in our work. However, in contrast to Chitwood (1938) who based subdivision of the genus on the size of denticles, we propose the composition of dentition (pattern of large and small denticles per lobe) as a distinguishing trait.
According to the classification by Chitwood (1938), only one morphological character (size of denticles) allows a subdivision in variants of the genus Mastophorus. Within this classification, specimens from Mus (with denticles of intermediate size) are subsumed with those described from Myodes, Rattus and Felis into the group M. muris var. muris (with large denticles) (Chitwood 1938). Specimens from Graomys (with smaller denticles) are categorized to M. muris var. ascaroides (Chitwood 1938). Here, we suggest a subdivision based rather on the composition of the dentition (i.e. based on the presence/absence of the central large or large denticles on the edge of lobes). These characters allow a consistent distinction of two variants: one occurring in Mus hosts and the second in non-Mus (Felis, Myodes and Rattus) hosts. The classification solely based on the dentition pattern we propose here would render specimen from Graomys hosts, classified as M. muris var. ascaroides (Chitwood 1938; Rojas and Digiani 2003), indistinguishable from our M. muris from Mus hosts. However, the presence of additional unpaired papilla on the posterior end of male specimens from Graomys distinguishes these from specimens from Mus.
We generalized the dentition for M. muris from Mus musculus to 1–(2 + n)–1–(2 + n)–1 per lobe of the pseudolabia and grouped these specimens in M. muris associated with Mus. All Mastophorus specimens from non-Mus hosts do not possess large denticles at the edges of pseudolabial lobes. Therefore, we assigned these specimens to M. muris associated with non-Mus hosts. Specimens of Mastophorus from Graomys (Rojas and Digiani 2003) represent an exception, because dentition assigns them to M. muris associated with Mus, but the number of caudal papillae allows a differentiation in this case. We suggest additional research to clarify whether specimens from Graomys justify instituting a new Mastophorus variant for them.
In conclusion, we found that the dentition pattern is the most reliable morphological character to distinguish host-associated variants of M. muris. We support a subdivision of Mastophorus into two variants: M. muris associated with Mus and M. muris associated with non-Mus hosts (hosts: Apodemus, Felis and Rattus). Previous descriptions of M. muris contain numerous traits with high morphological and morphometric variability (Chitwood 1938; Wertheim 1962; Rojas and Digiani 2003). This is a challenge for the potential subdivision of the genus as it is possible that we overlooked variability in the dentition pattern for specimens from non-Mus hosts due to our shallow sampling of few specimens. We argue, however, that the variants observed for worms from Myodes and Rattus hosts fall clearly outside of the variability observed in specimens from Mus hosts. The latter were sampled densely in an area overlapping the sampling for the other rodents. Mastophorus from the same host but different geographical regions showed low variability based on their partial CO1-sequences which confirmed host association. For example, the Mastophorus from Apodemus and Felis (Mastophorus probably from a preyed Apodemus) which were collected from geographical regions approx. 600 km apart (Tegel, Berlin and Usingen, Hesse in Germany) showed high identity of their partial CO1 sequences. The same applies to the CO1 sequences of Mastophorus isolates from Sigmodon which were collected approx. 400 km apart (Piedmont region of Georgia and Costal Plains region of Georgia; Thompson et al. 2019) and the isolates from Oxymycterus which were collected approx. 1000 km apart (Ilha Grande, Rio Janeiro and Luizote, Minas Gerais in Brazil; de Barros 2015). Nonetheless, we recommend further investigations covering more of the host spectrum and denser sampling for multiple hosts from different geographical regions to validate host association within the genus Mastophorus. The observed morphological and genetic variations distinguish isolates corresponding to different host usage that might justify separation into different species, if not genera. However, comparing species pairs, whether our results influence or motivate studies to advocate the change in taxonomy, goes beyond the scope of our work.
In addition to the morphological evidence, our study considers the information provided by the phylogeny with different marker genes, while the CO1 gene has been shown to be more informative because of the higher substitution rate (Blouin 2002) which results in a higher resolution of the CO1 tree. Comparing species pairs, Blouin reported genetic divergence of around 10% (range 6.9–13.0) for the CO1 gene (Blouin 2002). Thus, the M. muris specimens from Mus and non-Mus may be considered different species. In addition, ITS sequences distinguish between closely related parasitic nematode species and were consistent with both 18S and CO1 analyses; our M. muris isolates from house mice cluster in one clade in a sister group relation to the Mastophorus sequence from non-Mus hosts. Overall, our genetic analyses based on the three marker genes (18S, CO1 and ITS) support the subdivision of the genus Mastophorus in the two proposed variants with moderate to good support.
Considering the differentiation of members from the genera Mastophorus and Protospirura, our phylogenetic analyses suggest the separation as stated by previous morphological descriptions (Quentin et al. 1968; Quentin 1969). The phylogenetic tree based on ITS and CO1 supports the respective monophyly of the genus Mastophorus. Thus, based on the novel available genetic data, the relationship between the genera Mastophorus and Protospirura within the order Spirurida could be less controversial. The cosmopolitan Protospirura muricola is clearly separated from Mastophorus spp. based on the phylogeny provided here. The sequences from P. muricola built a separate group with CO1 sequence identities of 84.99–85.62% (472 bp) to Mastophorus from Mus. Misidentifications, erroneously assigned sequences like those from P. numidica sequences obtained from specimens collected from Oxymycterus in Brazil (de Barros 2015) assigned to the genus Prostospirura without describing details or illustrating their specimens, while belonging to the genus Mastophorus (Costa et al. 2018, 2022), are challenging to conclude and might promote confusion in the classification. While further studies should focus on data generation, validation (by associated taxonomic annotation) and phylogenetic analysis of reference sequences to clarify the confusion of Mastophorus with Protospirura, our work provides a deeper insight into the morphological and phylogenetic diversity of the genus Mastophorus.
Data availability
Sequences obtained and used for the analysis are available at NCBI GenBank with the accession numbers: 18S [MN086286–MN086291], CO1 [MK867474–MK867480] and ITS [MK829001–MK829007]. Voucher specimens for M. muris from house mouse identified in this study were deposited in the Natural History Museum in Berlin, Germany in the department “Vermes” under specimens numbers E.7635–E.7639.
References
Baird SJE, Ribas A, Macholán M, Albrecht T, Piálek J, Goüy de Bellocq J (2012) Where are the wormy mice? A reexamination of hybrid parasitism in the European house mouse hybrid zone. Evolution 66:2757–2772. https://doi.org/10.1111/j.1558-5646.2012.01633.x
Blaxter ML (2004) The promise of a DNA taxonomy. Phil Trans R Soc B 359:669–679. https://doi.org/10.1098/rstb.2003.1447
Blouin MS (2002) Molecular prospecting for cryptic species of nematodes: mitochondrial DNA versus internal transcribed spacer. Int J Parasitol 32:527–531. https://doi.org/10.1016/S0020-7519(01)00357-5
Bowles J, Hope M, Tiu WU, Liu X, McManus DP (1993) Nuclear and mitochondrial genetic markers highly conserved between Chinese and Philippine Schistosoma japonicum. Acta Trop 55:217–229. https://doi.org/10.1016/0001-706X(93)90079-Q
Casiraghi M, Anderson TJC, Bandi C, Bazzocchi C, Genchi C (2001) A phylogenetic analysis of filarial nematodes: comparison with the phylogeny of Wolbachia endosymbionts. Parasitology 122:93–103. https://doi.org/10.1017/S0031182000007149
Chabaud AG (1975) Keys to genera of the order Spirurida. Part 2. Spiruroidea, Habronematoidea and Acuarioidea. In: Anderson RC, Chabaud AG, Willmott S (eds) Keys to the nematode parasites of vertebrates, No 3 Archival, vol 2009. CABI, Wallingford, pp 361–390
Chitwood BG (1938) The status of Protospirura vs. Mastophorus with a consideration of the species of these genera. Livro jubilar do Professor Lauro Travassos, 115–118
Costa NA, Simões RO, Vilela RV, Souza JGR, Cardoso ST, Leiner NO, Gentile R, Maldonado AJ (2018) Morphological and genetic characterization of Pterygodermatites (Paucipectines) zygodontomis (Nematoda: Rictulariidae) from Necromys lasiurus (Rodentia: Sigmodontinae) from Uberlândia, Brazil. J Helminthol 92:618–629. https://doi.org/10.1017/S0022149X17000736
Costa NA, dos Santos CT, da Costa-Neto SF, Alvarez MR, Junior AM, Gentile R (2022) Helminths of sigmodontine rodents in an agroforestry mosaic in the Brazilian Atlantic Forest: patterns and processes of the metacommunity structure. Int J Parasitol Parasites Wildl 18:82–91. https://doi.org/10.1016/j.ijppaw.2022.04.008
Dayrat B (2005) Towards integrative taxonomy. Biol J Linn Soc 85:407–417. https://doi.org/10.1111/j.1095-8312.2005.00503.x
de Barros JSL (2015) Morphological taxonomy and molecular phylogeny of Physaloptera (Nematoda: Spirurida). Dissertation, Instituto Oswaldo Cruz
De Queiroz K (2007) Species concepts and species delimitation. Syst Biol 56:879–886. https://doi.org/10.1080/10635150701701083
Floyd RM, Rogers AD, Lambshead PJD, Smith CR (2005) Nematode-specific PCR primers for the 18S small subunit rRNA gene. Mol Ecol Notes 5:611–612. https://doi.org/10.1111/j.1471-8286.2005.01009.x
Gasser RB, Hoste H (1995) Genetic markers for closely-related parasitic nematodes. Mol and Cell Probes 9:315–319. https://doi.org/10.1016/S0890-8508(95)91588-5
Grzybek M, Bajer A, Behnke-Borowczyk J, Al-Sarraf M, Behnke JM (2015) Female host sex-biased parasitism with the rodent stomach nematode Mastophorus muris in wild bank voles (Myodes glareolus). Parasitol Res 114:523–533. https://doi.org/10.1007/s00436-014-4214-0
Hall MC (1916) Nematode parasites of mammals of the order Rodentia, Lagomorpha, and Hyracoidea. No. 2131. Proc US Natl Mus 50:1–258. https://doi.org/10.5479/si.00963801.50-2131.1
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755. https://doi.org/10.1093/bioinformatics/17.8.754
Jarquín-Díaz VH, Balard A, Jost J, Kraft J, Dikmen MN, Kvičerová J, Heitlinger E (2019) Detection and quantification of house mouse Eimeria at the species level—challenges and solutions for the assessment of coccidia in wildlife. Int J Parasitol Parasites Wildl 10:29–40. https://doi.org/10.1016/j.ijppaw.2019.07.004
Julius RS, Schwan EV, Chimimba CT (2018) Helminth composition and prevalence of indigenous and invasive synanthropic murid rodents in urban areas of Gauteng Province, South Africa. J Helminthol 92:445–454. https://doi.org/10.1017/S0022149X17000761
Kataranovski DS, Vukicevic-Radio OD, Kataranovski MV, Radovic DL, Mirkov II (2008) Helminth fauna of Mus musculus Linnaeus, 1758 from the suburban area of Belgrade, Serbia. Arch Biol Sci 60:609–617. https://doi.org/10.2298/ABS0804609K
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Lafferty KD, Shostak AW, Bush AO, Lotz JM (1997) Parasitology meets ecology on its own terms: Margolis et al. revisited. J Parasitol 575–583. https://doi.org/10.2307/3284227
Maaz D, Krücken J, Blümke J, Richter D, McKay-Demeler J, Matuschka F-R, Hartmann S, von Samson-Himmelstjerna G (2018) Factors associated with diversity, quantity and zoonotic potential of ectoparasites on urban mice and voles. PLoS ONE 13:e0199385. https://doi.org/10.1371/journal.pone.0199385
Neupane B, Miller AL, Evans AL, Olsson GE, Höglund J (2020) Seasonal variation of Mastophorus muris (Nematoda: Spirurida) in the water vole Arvicola amphibius from southern Sweden. J Helminthol 94. https://doi.org/10.1017/S0022149X18000937
Quentin JC (1969) Cycle biologique de Protospirura muricola Gedoelst, 1916, Nematoda spiruridae. Ann Parasitol Hum Comp 44:485–503. https://doi.org/10.1051/parasite/1969444485
Quentin JC (1970) Morphogénèse larvaire du Spiruride Mastophorus muris (Gmelin, 1790). Ann Parasitol Hum Comp 45:839–855. https://doi.org/10.1051/parasite/1970456839
Quentin JC, Karimi Y, Rodriguez de Almeida C (1968) Protospirura numidica criceticola, n. subsp. parasite de rongeurs cricetidae du Brésil: cycle evolutif. Ann Parasitol Hum Comp 43:583–596. https://doi.org/10.1051/parasite/1968435583
R Core Team (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
Read CP, Millemann RE (1953) Helminth parasites in kangaroo rats. Univ Calif Publ Zool 59:61–80. https://doi.org/10.5555/19530801868
Rojas MDC, Digiani MC (2003) First record of Mastophorus muris (Gmelin, 1790) (Nematoda: Spiruroidea) from a wild host in South America. Parasite 10:375–378. https://doi.org/10.1051/parasite/2003104375
Ronquist F, Huelsenbeck J (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Rózsa L, Reiczigel J, Majoros G (2000) Quantifying parasites in samples of hosts. J Parasitol 86:228–232. https://doi.org/10.1645/0022-3395(2000)086[0228:QPISOH]2.0.CO;2
Schliep KP (2011) phangorn: phylogenetic analysis in R. Bioinformatics 27:592–593. https://doi.org/10.1093/bioinformatics/btq706
Schliep K, Potts AA, Morrison DA, Grimm GW (2016) Intertwining phylogenetic trees and networks. PeerJ Preprints 4:e2054v1. https://doi.org/10.7287/peerj.preprints.2054v1
Schliep ME, Alonzo CN, Morris MA (2017) Beyond RCTs: innovations in research design and methods to advance implementation science. Evid Based Commun Assess Interv 11:82–98
Skrjabin KI (1961) Key to parasitic nematodes. Israel Program for Scientific Transactions: Jerusalem. pp 497
Smales LR (1995) Mastophorus muris (Nematoda:Spriocercodae) from the musky rat-kangaroo, Hypsiprymodom moschatus. Trans R Soc S Aust 119:95–96
Smales LR, Harris PD, Behnke JM (2009) A redescription of Protospirura muricola Gedoelst, 1916 (Nematoda: Spiruridae), a parasite of murid rodents. Syst Parasitol 72:15. https://doi.org/10.1007/s11230-008-9147-5
Smith JA, Kinsella JM (2011) Gastric Spiruridiasis caused by Mastophorus muris in a captive population of striped possums (Dactylopsila trivirgata). J Zoo Wildl Med 42:357–359
Stevenson M, Stevenson MM, BiasedUrn I (2018) Package ‘epiR’. Tools for the analysis of epidemiological data R package version 0.9–62. https://mirror.ibcp.fr/pub/CRAN/web/packages/epiR/epiR.pdf
Sutton CA (1989) Contribution to the knowledge of Argentina’s parasitological fauna XVII. Spirurida (Nematoda) from Neotropical Cricetidae: Physaloptera calnuensis n. sp. and Protospirura numidica criceticola Quentin, Karimi and Rodriguez de Almeida. Bull Mus Nation Hist Nat Paris 4 º ser 11 Section A 1:61–67
Thompson AT, Cleveland CA, Koser TM, Wyckoff ST, Yabsley MJ (2019) The occurrence of Physaloptera hispida and a Mastophorus sp. in pulmonary vessels of hispid cotton rats (Sigmodon hispidus) from Georgia, U.S.A. J Parasitol 105:718–723. https://doi.org/10.1645/18-176
Torres J, García-Perea R, Gisbert J, Feliu C (1998) Helminth fauna of the Iberian lynx, Lynx pardinus. J Helminthol 72:221–226. https://doi.org/10.1017/s0022149x00016473
Wertheim G (1962) A Study of Mastophorus muris (Gmelin, 1790) (Nematoda: Spiruridae). Trans Am Micros Soc 81:274–279. https://doi.org/10.2307/3224051
Wright ES (2015) DECIPHER: harnessing local sequence context to improve protein multiple sequence alignment. BMC Bioinforma 16:1–14. https://doi.org/10.1186/s12859-015-0749-z
Wright ES (2020) RNAconTest: comparing tools for noncoding RNA multiple sequence alignment based on structural consistency. RNA 26:531–540. https://doi.org/10.1261/rna.073015.119
Acknowledgements
We thank Jaroslav Piálek and his team (Institute of Vertebrate Biology, AS CR, Brno, Department of Population Biology in Studenec) for help with sample collection. We thank Alice Balard (Research Group Ecology and Evolution of Molecular Parasite-Host Interactions) for her helpful discussion, comments and additional support.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by the German Foundation of Scientific Research (DFG) [grant number: 285969495/HE 7320/2–1 to EH] and the German Academic Exchange Service (DAAD) [scholarship to VHJD] and the Research Training Group 2046 “Parasite Infections: From Experimental Models to Natural Systems” [associated student VHJD].
Author information
Authors and Affiliations
Contributions
JJ, EH and VHJD designed the study. JJ, JH, LD, DM, EH, and VHJD collected the samples. JJ, TS and PM performed laboratory and microscopy work. JJ and VHJD performed the analysis. JJ wrote the original draft manuscript. JH and VHJD wrote the final version with contributions and feedback from all the authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
House mice (Mus musculus) for the collection of helminths used in present study were captured under the permit no. 2347/35/2014.
Consent to participate
Not applicable.
Consent for publication
All the information derived from the captured mice is allowed to be published and available for the scientific community and general public.
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Hiroshi Sato
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
†Jörg Hirzmann is deceased
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jost, J., Hirzmann, J., Ďureje, Ľ. et al. Dentition patterns and molecular diversity of Mastophorus muris (Gmelin, 1790) (Nematoda: Spiruroidea) support a host-associated subdivision. Parasitol Res 123, 237 (2024). https://doi.org/10.1007/s00436-024-08259-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00436-024-08259-1