Abstract
With the explosion in knowledge about the molecular landscape of lymphoid malignancies and the increasing availability of high throughput techniques, molecular diagnostics in hematopathology has moved from isolated marker studies to a more comprehensive approach, integrating results of multiple genes analyzed with a variety of techniques on the DNA and RNA level. Although diagnosis of lymphoma still relies on the careful integration of clinical, morphological, phenotypic, and, if necessary molecular features, and only few entities are defined strictly by genetic features, genetic profiling has contributed profoundly to our current understanding of lymphomas and shaped the two current lymphoma classifications, the International Consensus Classification and the fifth edition of the WHO classification of lymphoid malignancies. In this review, the current state of the art of molecular diagnostics in lymphoproliferations is summarized, including clonality analysis, mutational studies, and gene expression profiling, with a focus on practical applications for diagnosis and prognostication. With consideration for differences in accessibility of high throughput techniques and cost limitations, we tried to distinguish between diagnostically relevant and in part disease-defining molecular features and optional, more extensive genetic profiling, which is usually restricted to clinical studies, patients with relapsed or refractory disease or specific therapeutic decisions. Although molecular diagnostics in lymphomas currently is primarily done for diagnosis and subclassification, prognostic stratification and predictive markers will gain importance in the near future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hematopathology has always been at the forefront in using molecular studies as tools for scientific progress and diagnostic advances. From the very beginning, molecular findings such as the identification of recurrent translocations or detection of clonality have shaped both our understanding of lymphoma and their classification [39].
With the relatively recent introduction of massively parallel sequencing, there has been an explosion in the knowledge about the genetic landscape of lymphoid proliferations [20]. This also had an important impact on the two new classifications for lymphoma, the International Consensus Classification (ICC) and the WHO HAEM5 classification [2, 12], as well as on the diagnostic practice. Molecular studies have four main applications in lymphoma pathology: (1) diagnostic, for confirmation of a neoplastic lymphoproliferation and for disease subclassification; (2) prognostic, for patient stratification and as aid in therapy decisions; (3) predictive, for the identification of targets of therapy; and (4) for the detection of measurable (formerly minimal) residual disease (MRD). Genetic alterations including chromosomal translocations, copy number variations and mutations, gene expression profiles, and potentially epigenetic alterations can be used for diagnostic purposes. In addition, immunoglobulin (IG) and T-cell receptor (TR) gene rearrangements provide unique targets for detecting clonality in a lymphoproliferation. In this review, we summarize current molecular diagnostics in lymphoma, with a focus on clinical relevance. Acute lymphoblastic leukemia/lymphoblastic lymphoma, MRD detection, and detailed methodological aspects are beyond the scope of this review.
Clonality analysis
Clonality analysis is an important technique in the diagnosis of lymphoid proliferations, which allows distinction between the immunogenetic variability in reactive lymphoid proliferations and the lack thereof in lymphoid neoplasia. The rationale of clonality analysis is based on the highly diverse spectrum of B and T cells which is generated to allow an adaptive immune response to the diverse array of antigens the individual may encounter. This diversity is generated by rearrangement and the junctional diversity of IG and TR genes. In a lymphoid proliferation, this diversity can be studied by multiplex PCR amplification of IG and TR genes. For conventional clonality analysis, this is followed by fragment size analysis of the resulting PCR products [46, 109]. In lymphoma, the neoplastic cells all originate from the same transformed cell and therefore have the same IG or TR rearrangement. Accordingly, a large proportion of the PCR products in a lymphoma sample will have the same size resulting in one or two clonal peaks in fragment size analysis. This contrasts with reactive lymphoid tissues, which consist of a diverse population of cells with a large diversity in IG and TR genes, generating a polyclonal Gaussian curve. To achieve a high sensitivity for the detection of clonality, multiple targets are tested (i.e., IGH and IGK; TRG and TRB) which provide complementary information.
In recent years, it has become possible to analyze the IG and TR products with next-generation sequencing (NGS), allowing a much more detailed analysis of the IG and TR rearrangements present in a sample [10, 90]. Within the EuroClonality-NGS working group, an assay was designed with novel primers that generate small amplicons, making it more suitable for formalin-fixed paraffin-embedded (FFPE) tissue [90]. The IG clonality analysis makes use of the complementarity of the different targets. PCR reactions are performed in three tubes (IGH VJ; IGH DJ; IGK VJ and IGKV/Intron-Kde), followed by library preparation for sequencing.
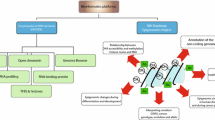
To analyze the sequencing data from NGS-based clonality analysis, a bio-informatic analysis pipeline is used to identify clonotypes in the raw sequencing data [11, 18]. A clonotype is defined by the involved genes and the amino acid sequence of the junction between these genes. The distribution of the different clonotypes can then be viewed in tabular format or with different types of visualizations (Fig. 1).
A Overview of conventional and NGS-based clonality analysis. From DNA isolated from a lymphoproliferation suspicious for neoplasia, IG and/or TR genes are amplified in a PCR reaction. For conventional clonality analysis, the PCR product is analyzed by fragment size analysis. For NGS-based clonality analysis, the PCR product is sequenced with a next-generation sequencing platform after which bio-informatic analysis is performed. B Example of an NGS-based clonality analysis result for IGH DJ with a polyclonal pattern, output from ARResT/Interrogate. The amino acid length of the junction is indicated on the x-axis, and the percentage of reads is indicated on the y-axis. The 50 most prominent clonotypes are indicated with different colors. Less frequent clonotypes in the background are shown in gray
Before interpreting, it is important to verify the quality of the results from NGS-based clonality analysis. DNA quality, DNA input, the lymphoid component, the number of cells suspicious for malignancy, the number of reads, and the number of clonotypes detected in the sample are important parameters. In particular, a pattern in which a limited number of clonotypes is present in a high percentage with a very limited background suggests a low template and should be signed out as non-evaluable. This low template pattern can be caused by poor DNA quality, few B-cells, or the presence of a prominent clone which cannot be amplified due to somatic hypermutation leading to primer mismatch.
Similar to conventional clonality analysis, NGS-based clonality analysis can be used to distinguish reactive lymphoid proliferations from neoplasia. Studies with different assays have shown that NGS-based IG clonality analysis is at least as sensitive as conventional BIOMED-2 clonality analysis [6, 30, 71]. In conventional clonality analysis, small clones are often inapparent in the polyclonal background, especially if the size of the clone is near the center of the Gaussian curve. With NGS, small clones can be detected within the polyclonal background. In a technical feasibility study of the EuroClonality-amplicon-based NGS protocol, a lymphoma diluted in a polyclonal sample could still be detected as clonal with a limit of detection of 2.5%[90]. Sensitivity can be increased if the ratio between the dominant clonotype and the background, and replicate analyses are included in the assessment of the assay. If the purpose of the analysis is to search for the presence of an index clonotype, the sensitivity is even higher (Fig. 2).
Assessment of clonal relationship between a mantle cell lymphoma in the lymph node (A) and the bone marrow (B). In these results from the IGK VJ target, a clear clone is detected in the lymph node in 98% of reads with a ratio between the #1 clonotype and the average of clonotypes #3–#7 of 363. The same clone is detected in the bone marrow in 10% of reads with a ratio of 1.8. Especially when looking for a known clonotype, NGS-based clonality analysis has a very high sensitivity
This high sensitivity is very helpful in the study of lymphomas with few tumor cells in a prominent reactive background such as classic Hodgkin lymphoma (cHL) or T-cell/histiocyte-rich large B-cell lymphoma [106, 107]. Finally, the development of NGS-based clonality analysis has the potential to improve the diagnostic accuracy of notoriously difficult lymphoproliferations including skin lymphomas, early T-cell lymphomas, and lymphoproliferations in the context of immune deficiency and immune dysregulation. However, further studies are needed to determine the true potential and practical application of NGS-based clonality analysis in these diseases.
NGS-based clonality analysis has the potential to make interpretation of the results more objective as it provides quantitative and qualitative (i.e., actual sequence) data on the clonotypes that are present. Quantitative criteria for interpretation have been proposed which are based on a combination of the percentage of reads attributed to the most prominent clonotype and the ratio between the most prominent clonotype and the background [6, 30, 71]. When using a quantitative approach to NGS-based clonality interpretation, it is important to keep in mind that many variables are involved; results should always be interpreted in the histological and clinical context and taking into consideration results from other molecular analyses. Also, it must be emphasized that clonality does not equal malignancy; clonal results can be obtained in reactive conditions and lymphomas can generate a polyclonal result due to quality issues, few tumor cells, or an inability to detect the B-cell clone due to somatic hypermutation [47].
The sequences of the clonotypes from NGS-based clonality analysis allow for accurate clonal comparison. In patients with multiple synchronous or metachronous hematological malignancies, prognosis or treatment might be different depending on whether the lesions are clonally related or not [86]. Importantly, hematological malignancies that originate from the same clone may differ in morphology and immunophenotype which may even include a switch in lineage, sometimes due to so-called transdifferentiation [32]. In conventional clonality analysis, comparison based on clonal peak sizes can be inaccurate due to small sequence changes caused by mutations and technical artifacts [50, 70]. To conclude that two lymphoproliferations are clonally related generally requires identical peak sizes in at least two targets [34]. With NGS-based clonality analysis, the exact sequences can be compared which allows clonal comparison from a single target. In addition, NGS-based clonality analysis is very helpful to solve difficult cases with divergent clonal evolution. Concurrently or subsequently occurring lymphoid proliferations might have a shared ancestor clone as evidenced by shared IG/TR rearrangements for some targets, but they could at the same time have different IG/TR rearrangements on the other allele or in another target [7, 43]. Another application of NGS-based IG sequencing is the determination of the mutational status of the rearranged IG heavy chain gene (see below).
In conclusion, NGS-based clonality analysis opens up new possibilities for the diagnosis of lymphoid proliferations, but requires integration of results in the clinical, histological, and molecular context and requires knowledge about the potential and the technical and immunobiological pitfalls of IG/TR rearrangements. Recently, practical guidelines on the interpretation of NGS-based clonality results were described by the EuroClonality-NGS working group [108].
Molecular studies in mature B-cell lymphomas
The differential diagnosis of small B-cell lymphomas is determined by the clinical presentation (leukemic, nodal, extranodal), morphology, and the immunophenotype. These features also guide the selection of molecular studies, including the detection of translocations and mutations, and their interpretation. Although many cases do not require these tests, mutational profiling is increasingly used for more precise subtyping and prognostic purposes (Table 1). Of note, even hallmark mutations such as MYD88L265P or BRAFV600E for lymphoplasmacytic lymphoma and hairy cell leukemia, respectively, are not completely specific, but rarely may be found in other small B-cell lymphomas.
Chronic lymphocytic leukemia/small lymphocytic lymphoma (CLL/SLL)
Although the diagnosis of CLL/SLL is straightforward in most cases, genetic alterations are of great importance for prognostic stratification and influence therapy decisions. The presence or absence of somatic hypermutation in the rearranged IG heavy chain gene, taking a <98% identity with the germline sequence as cutoff, is of prognostic and therapeutic relevance [1]. In addition, some subsets of CLL with stereotyped B-cell receptors and CLL expressing IG lambda V3-21 with the R110 point mutation show distinct prognostic features [1, 66]. Although the cytogenetic and mutational landscape of CLL is highly diverse and shows differences between IGH-mutated and unmutated cases, alterations of TP53, usually bi-allelic (i.e., mutation and del17p) are the single most important prognostic and predictive parameter and need to be tested before initiation of therapy [64]. Of importance, even cases with small subclonal mutations (variant allele frequency (VAF) <10%) show a poorer outcome and response to therapy [85]. Mutational analysis of BTK, PLCG2, and BCL2 is recommended for CLL under certain targeted therapies (ibrutinib and venetoclax) [64].
Lymphoplasmacytic lymphoma (LPL)
LPL, in its classic form presenting as Waldenström’s macroglobulinemia, shows the canonical MYD88L265P in 95 to 97% of cases, and CXCR4 is mutated in 30 to 40% [91, 104]. IgM monoclonal gammopathy of undetermined significance (MGUS) and non-IgM LPL show these mutations at lower frequency. Especially for IgM MGUS and LPL with low tumor burden, sensitive techniques need to be employed for detection [22]. The presence or absence of these two mutations has a significant impact on clinical features and therapy response and should be determined in all LPL cases. Whereas the IgM-expressing multiple myeloma is universally negative, rare cases of splenic [74], nodal, and extranodal marginal cell lymphomas in the ocular adnexal region show MYD88 mutations [76], as well as 2–3% of CLL, the latter cases associated with favorable biology [64].
Marginal zone B-cell lymphomas
Due to the lack of a specific immunophenotype or a defining or characteristic genetic alteration, diagnosis of marginal zone lymphomas (MZL) relies on a combination of clinical, morphological, and phenotypic features, but mutational profiling and translocation detection may help to separate them from other small B cell lymphomas. A subset of MALT lymphomas shows recurrent translocations frequently involving the MALT1 locus on 18q21; their frequency, as well as those of some recurrent mutations is strongly dependent on the primary organ manifestation [110]. In small B-cell lymphomas with nodal or bone marrow presentation, KLF2, NOTCH1/2, PTPRD, BIRC, TRAF3, CARD11, and IRF8 mutations favor a diagnosis of MZL. Of note, nodal MZL rarely may show MYD88 or BRAF mutations [49, 76, 96].
Follicular lymphoma and variants
Conventional follicular lymphoma (FL) is characterized by the presence of a t(14;18)(q32;q21) translocation in about 85–90% of cases. The mutational profile is characterized by frequent alterations of epigenetic regulators, transcription factors, and others, whereas transformation is associated with a different set of genes, including TP53 mutations and MYC translocations[44, 49]. Although risk stratification models incorporating mutations, such as the m7-FLIPI [75], have been proposed, they are strongly dependent on the type of therapy. Therefore, genetic profiling currently is not of clinical or diagnostic relevance in typical FL, except for the detection of EZH2 mutations before treatment with the inhibitor tazemetostat [62]. However, it is useful for discerning FL variants, including the newly described BCL2 rearrangement negative, CD23+ follicle center lymphoma [12, 67], which shows a high frequency of STAT6 mutations, cutaneous follicle center lymphoma and pediatric type FL, which shows frequent alterations of TNSFRSF14, MAP2K1, and IRF8, distinct from adult FL [92].
Mantle cell lymphoma
Mantle cell lymphoma (MCL) is characterized by the presence of the t(11;14) translocation, usually easily demonstrable by the surrogate marker cyclin D1 overexpression. MCL can be divided into the conventional subtype, derived from naïve B cells with unmutated IGHV genes, usually SOX11+, whereas the leukemic non-nodal type shows somatic hypermutated IGHV genes, lacks SOX11, and shows less genomic complexity [65]. Clinically relevant for both subtypes are TP53 mutations, associated with a more aggressive clinical course, higher proliferation rate, and frequent blastoid morphology. The rare cyclin D1− MCL show rearrangements of CCND2 or CCND3, which should be demonstrated by FISH or other techniques, since immunohistochemistry is unreliable in this setting [57].
Other small B-cell lymphomas
Hairy cell leukemia is virtually universally characterized by the BRAFV600E hotspot mutation, with the exception of cases using the IGHV4-34 family, which may show MAP2K1 mutations and are similar to hairy cell leukemia variant [102].
Plasma cell neoplasms
Multiple myeloma (MM) and its precursor lesion, non-IgM MGUS, show great genetic heterogeneity, with 45–50% showing trisomies of odd-numbered chromosomes, and 40–45% showing recurrent IGH translocations with a variety of partners, which are currently detected by interphase FISH [20]. Due to the prognostic impact of genetic alterations, the ICC has formally divided MM into cytogenetic subgroups [12]. Mutational profiling currently is not of diagnostic relevance outside of clinical studies, but an absence of MYD88 mutations helps to exclude LPL [27].
Diffuse large B-cell lymphoma and other large B-cell lymphomas
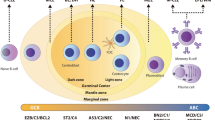
Diffuse large B-cell lymphoma (DLBCL) is both the most common and most diverse lymphoma occurring in adults, and a number of parameters, including but not restricted to clinical site and pathologic features, are used for subclassification. One of the earliest, and at the time most paradigm-shifting, approaches to applying molecular tools to the characterization of DLBCLs, particularly those that belong to the largest category of DLBCLs, not otherwise specified (NOS), occurred almost a quarter-century ago with the identification of two major forms based upon their putative cell-of-origin (COO) [4]. This original study employed gene expression profiling (GEP) using DNA microarrays to identify and divide most of these DLBCL, NOS cases into those of either germinal center B-cell (GCB) or activated B-cell (ABC) origin. This bifurcation also had prognostic connotations, in that the latter predicted for worse outcomes [113]. At a mechanistic level, GCB-DLBCLs tend to harbor mutations in chromatin-modifying genes while ABC-DLBCLs are more likely to display mutations that result in activation of the NFKB and BCR pathways [40, 115]. Notwithstanding the availability of a commercial RNA-based assay (Lymph2Cx) [116] that has yet to be widely adopted, translation of the COO strategy into routine clinical use has been difficult. This has led to the development of a number of surrogate immunohistochemical (IHC) algorithms that, unfortunately, do not perfectly reflect the GEP-based classification [35, 59]. While the binary use of COO, using IHC algorithms, has been retained in contemporary classification schemes and is used in diagnostic practice, it has become clear that this seemingly simple separation may not be sufficiently robust to capture the complexity of the molecular underpinnings of DLBCL to be of independent clinical and therapeutic value [19, 72].
This complexity has more recently been unraveled using a compendium of genomic approaches by different investigators that have independently largely converged into somewhat overlapping classifications identifying up to seven major subtypes of DLBCL [13, 45, 93] (Fig. 3). The complexity and comprehensive genetic testing required to unravel these different subtypes is currently not readily adoptable by routine diagnostic laboratories, but publicly available online tools such as LymphGen may be used to facilitate classification [87, 114]. Of note, the LymphGen tool does not subtype 30-40% of cases of DLBCL into any of the major groups, in contrast to other proposed molecular subclassifications of DLBCL which assign all evaluable cases to a specific category. Many of the key or defining features in the various clusters are already available for testing in some or even most routine settings, since they can be ascertained by a combination of FISH (for BCL2 and BCL6 translocations, and TP53 loss) and targeted mutational analysis (evaluating MYD88, CD79B, EZH2, CREBBP, KMT2D, NOTCH1, NOTCH2, TP53, SGK1, TET2, and SOCS1, among others), although it is unlikely that such focused approaches fully capture the complexity of these subtypes. Nevertheless, a simplified 20-gene classifier has been described and has the potential to approximate the LymphGen approach to classify four main subgroups [60]. Conspicuously, and perhaps surprisingly, (almost entirely) absent as a major variable in the stratification of the different genomic categories of DLBCL is testing for a MYC translocation that has long been a key component of evaluation of this group of lymphomas. However, its presence does help refine one of the subgroups, and it remains an important initial genetic abnormality to evaluate in all aggressive B-cell lymphomas, particularly as a necessary first step in identifying the genetically-defined subtype of high-grade B-cell lymphoma (HGBL) of double-hit lymphoma (DHL), as discussed below. Provocatively, many of these major genomic subtypes of DLBCL share features evident in other, and often indolent, B-cell lymphomas such as marginal zone lymphoma, follicular lymphoma, and small lymphocytic lymphoma/chronic lymphocytic leukemia, as well as other defined DLBCLs, including those arising in extranodal immune-privileged sites or the mediastinum.
Molecular subtypes of DLBCL, NOS incorporating gene expression profiles and recurrent genetic alterations (modified according to [20]). The relationship between COO and the probabilistic assignments to genetics-based subgroups are shown. The size of the subgroup circles approximates the proportions of patients in each group. Tumors assigned with high confidence to ≥2 subgroups are assigned to the composite group, while ∼37% of tumors are not assigned to any subgroup with sufficient confidence (other). The hallmark genetic features are frequent within the respective subgroup but are not required for that assignment. Although capture of the total complexity would require a workup beyond current diagnostic standards, sequencing of a limited gene panel together with immunophenotyping and FISH for recurrent translocations can help to (tentatively) assign a significant subset of DLBCL, NOS to one of the molecular subgroups
Another genetically defined LBCL entity is associated with an 11q aberration which in most cases harbors the combination of a proximal gain at 11q23.2-11q23.3 and distal loss at 11q24-qter, although some only show one of these features [88]. This lymphoma was previously classified as a Burkitt-like lymphoma but that designation has subsequently been revised since these cases, though morphologically and immunophenotypically sharing features with Burkitt lymphoma (BL), actually lack mutations that are typically evident in BL, as discussed below [31]. Commercially available 11q FISH probes are imperfect in identifying this defining aberration, and copy number array or other genomic analyses may be necessary to make this diagnosis. Notably, while classified as a LBCL by the ICC, WHO-HAEM5 has designated this as a form of HGBL [2, 12]. The rationale of the ICC for classifying the cases with 11q aberration as LBCL, rather than using the morphologically inspired term HGBL is based on their clinically less aggressive course and favorable outcome after therapy [31]. LBCLs with IRF4 rearrangements are another subtype that requires genetic testing for its diagnosis, typically by FISH although they also harbor mutations of IRF4 that may further support the diagnosis [80]. This entity is handled differently by ICC, which categorizes such cases within follicular lymphoma, given its overlapping features and typically indolent behavior, as compared with WHO-HAEM5, which groups this together with other LBCLs.
High-grade B-cell lymphoma
One of the two originally designated entities of high-grade B-cell lymphoma (HGBL) was one with MYC and BCL2 and/or BCL6 translocations, which was also termed double-hit (DH) or triple-hit (TH) lymphoma. However, this has subsequently been split into two distinct categories (1) HGBL with MYC and BCL2 rearrangements, which account for 80–90% of such cases; and (2) HGBL with MYC and BCL6 rearrangements, which account for 10–20% of such cases, with substantial evidence allowing the former to be retained as a bona fide entity, while the latter seems to be more heterogeneous and has been relegated, by the ICC but not WHO, to a provisional entity [41]. Virtually all of the MYC+BCL2 HGBLs are of GCB origin, whereas only around 50% of the MYC+BCL6 HGBLs are. The mutational landscape of the two DHLs is also quite distinct. Since there are no pathologic features (including morphologic, COO, or proliferative index) that can reliably identify DHLs, all DLBCLs should be screened for these defining translocations. MYC break-apart probes (BAP) alone are inadequate for detecting all MYC translocations, and different commercially available probes vary in sensitivity [63]. The addition of dual-color dual fusion MYC::IGH FISH probes may increase the yield by detecting translocations missed by the MYC BAP [42]. While the MYC partner in Burkitt lymphoma (BL, see below) is essentially always an IG gene, this is not the case here, and non-IG genes are involved in many (at least 40%) cases, including BCL6, IRF4, and PAX5 [16]. The suggestion that DHLs with MYC translocations involving IG have a worse prognosis appears to be unresolved [58, 84]. While DH MYC+BCL2 HGBLs are defined by these translocations that are imperfectly identified by cytogenetics or FISH, they can also be identified, perhaps even more reliably, by transcriptional profiling and have variably been termed molecular high grade (MHG) or DHITSig wherein cryptic translocations, missed by cytogenetics or FISH, are evident, occurring in 20% of cases [24, 37, 94]. It remains to be determined when and whether such testing can be applied in the routine diagnostic setting. Such lymphomas that almost always are of GCB origin may also, together with Burkitt lymphoma (see below), fall under the umbrella term of dark-zone lymphomas or DZSig lymphomas, with these cases refining the DLBCL-COO classification by identifying patients within this category with an inferior prognosis [3]. HGBL that is not a DHL or THL, termed HGBL, NOS, while enriched in MYC translocations (seen in ~45%) do not have a consistent unifying genetic profile, and remains a problematic diagnosis that is heavily reliant on often subjectively determined morphologic features.
Burkitt lymphoma
The genetic hallmark of BL is a balanced translocation involving MYC and one of the three immunoglobulin genes (IGH, IGK, or IGL, in approximately 80%, 15%, and 5% of cases, respectively). The translocations occur in the germinal center in mature B-cells, either at the time of somatic hypermutation (SHM) or class switch recombination (CSR). In sporadic BL (see below), this event happens most often at the time of CSR (~75%), interestingly with a proclivity to involve IGHA, rarely seen in other MYC-translocated lymphomas [54]. While traditionally epidemiologically stratified into three types — sporadic, endemic, and immunodeficiency-associated BL — a perhaps more mechanistic (though imperfect) and somewhat overlapping approach might be to separate cases based upon the involvement (or not) of EBV. Thus, EBV-positive cases may have fewer, or at least different, additional driver mutations [33, 51]. BL lacking EBV tends to harbor canonical mutations affecting ID3, TCF3, and CCND3 that are also more common in pediatric than in adult cases [82], while EBV-positive BL tends to be enriched in mutations affecting DDX3X, GNA13, and FOX01 [101]. The mutations evident in EBV-negative cases are quite unique to BL, while different mutations in other GC-derived aggressive B-cell lymphomas such as those affecting genes encoding chromatin modifiers are not seen, which together might be used to bolster the diagnosis of BL in routine diagnosis [33]. BL that dichotomizes based upon EBV status and mutational profile also differ in their timing of the MYC translocation, occurring during SHM in EBV-positive BL and more typically during CSR in EBV-negative BL. Testing for TP53 mutations is also of value in BL, providing important prognostic information for risk stratification [68]. Cases previously reported as TdT+ BL are no longer classified as BL and rather should be designated as B-lymphoblastic lymphoma/leukemia with MYC rearrangements. They differ from bona fide BL at three levels: (1) the MYC::IGH translocation occurs during VDJ recombination in the bone marrow; (2) they harbor RAS mutations, and lack those prevalent in BL; and (3) they do not express a functional B-cell receptor [111]. An approach to the initial FISH-based genetic workup of aggressive B-cell lymphomas, including DLBCL-NOS, HGBL, and BL is portrayed in Fig. 4, and current genetic tests are summarized in Table 2.
An approach to the use of FISH in aggressive B-cell lymphomas. This algorithm incorporating morphology, some immunohistochemical features, and detection of recurrent rearrangements allows subtyping of most aggressive B-cell lymphomas. Lymphomas with a Burkitt-like, blastoid, or large-cell morphology that have MYC with BCL2 and/or BCL6 rearrangements are categorized as HGBL, with either MYC and BCL2 (with or without BCL6 R) or MYC and BCL6 R. HGBL, NOS lacks these double hits and may have either Burkitt-like or blastoid morphology. TdT expression, but not CD34, may be seen in HGBL. Burkitt lymphomas typically have a MYC R without BCL2 or BCL6 and a mature GC B-cell phenotype. B-lymphoblastic lymphomas are blastoid-appearing, usually TdT+ and frequently but not always CD34+. Cases with a large B-cell morphology that lack a DH, generally, are categorized as DLBCL, NOS, or as one of the more specific types of large B-cell lymphoma. The possibility of a large B-cell lymphomas with 11q aberrations should be considered particularly with a Burkitt-like morphologic appearance but with coarse apoptotic material in the tingible body macrophages and lack of a MYC R. Some cases, however, will more closely resemble a DLBCL. DLBCL, diffuse large B-cell lymphoma; R, rearrangement; HGBL: high-grade B-cell lymphoma; DH, double hit (modified according to [41])
Classic Hodgkin lymphoma
Although the genetic landscape of classic Hodgkin lymphoma (CHL) and also nodular lymphocyte-predominant Hodgkin/B-cell lymphoma has been elucidated in the recent years, mutational profiling currently plays no significant role in practical diagnosis [112]. Of note, detection of B-cell clonality can be achieved in a significant subset of cases with modern protocols, and therefore, clonality studies are not suited to differentiate between Hodgkin lymphoma and other B-cell lymphomas with CHL-like morphology. Conversely, the detection of T-cell clonality and mutations characteristic for TFH lymphoma (see below) may aid in differentiating T-cell lymphomas with Reed-Sternberg cells from CHL [103].
Peripheral T-cell lymphomas
Peripheral T-cell lymphoma (PTCL) represents a heterogenous group of lymphoid malignancies, difficult to diagnose that constitutes approximately 10% of all lymphomas. PTCLs are divided according to their clinical presentation and site of involvement in leukemic, nodal, extranodal, and cutaneous [99]. The current ICC and WHO HAEM5 lymphoma classifications, recognize 33 and 36 diseases entities, respectively [2, 12]. The classification of PTCL in both lymphoma classifications is based on morphology and immunophenotype; however, this might change in the future since characteristic genetic alterations are increasingly recognized in PTCL. In contrast to B-cell lymphomas, clonality analysis is often needed to confirm a neoplastic T-cell proliferation with the caveat that finding a TR clonal rearrangement, per se, is not synonymous of a T-cell neoplasm.
Genetic alterations with potential relevance for the diagnosis of nodal T-cell lymphomas
A predominant genetic alteration has been identified in only a minority of nodal T-cell lymphomas, such as ALK or DUSP22 rearrangement (R) in anaplastic large T-cell lymphoma (ALCL). Although most nodal T-cell lymphomas have diverse molecular alterations, there are commonly affected cellular processes and signaling pathways [26]. In general, T-cell lymphomas are characterized by alterations in the TR and JAK/STAT signaling pathways, as well as mutations in epigenetic regulators [20]. These genetic alterations might have diagnostic, prognostic, or predictive impact in different entities (Table 3).
Anaplastic large cell lymphoma
Systemic ALCL is divided in two groups based on the presence or absence of ALK-R on chromosome 2p23. The most frequent partner gene is NPM1 on chromosome 5q35.1, followed by a variety of other fusion partners that encode oncogenic ALK fusion proteins. The demonstration of ALK-rearrangement or protein by immunohistochemistry is diagnostic of the disease and has allowed the recognition of several histologic variants, including the common variant (60% of the cases), the lymphohistiocytic variant (10%), the small-cell variant (5-10%), the Hodgkin-like variant (3%) and cases with more than one pattern [25]. Constitutive ALK activation promotes tumorigenesis through the activation of multiple signaling pathways such as PLCγ, PI3K-AKT1, JAK-STAT, mTOR, and MEK-ERK [15, 78]. Most ALCL have clonally rearranged TR genes despite the failure to express TR molecules on the cell surface [9]. The mutational landscape includes recurrent mutations of TP53, epigenetic modifiers, and genes in the TR pathway [53]. More recently, it has been shown that ALK-positive ALCL has recurrent missense mutations in the extracellular domain of NOTCH1 (12%; distinct from the truncating mutations of the PEST domain in B-cell lymphomas), a potential therapeutic target [48].
ALK-negative ALCL, by definition, lacks ALK rearrangement, and although it is considered a single disease entity, molecular data indicates the existence of several distinct subgroups [73]. Similar to ALK+ALCL, activation of the JAK-STAT signaling pathway seems to play an important role and is in many cases secondary to STAT3, JAK1, and JAK3 mutations, but also through rearrangements involving ROS1, TYK2 of FRK. Two recurrent chromosomal translocations with characteristic clinicopathological features are recognized. ALK-negative ALCL with DUSP22-R (20–30%), has monomorphic cytology, absence of cytotoxic molecules and PD-L1 expression, lacks JAK-STAT pathway activation, has a characteristic gene expression signature and recurrent MSC mutations [55, 56]. The 2022 ICC considers ALK-negative ALCL with DUSP22-R a genetic subtype of ALK-negative ALCL with usually favorable outcome [12, 26]. FISH analysis for DUSP22 is recommended to identify this genetic subgroup (Fig. 5). The second recurrent translocation involves TP63 (5–8%) which encodes a p63 fusion protein and is associated with aggressive clinical behavior and poor outcome [73]. Truncated ERBB4 overexpression is observed in 24% of ALK-negative ALCL and represents a potential therapeutic target [89].
Anaplastic large T-cell lymphoma with DUSP22 rearrangement. A H&E stain shows a lymph node with diffuse infiltration of large anaplastic cells (original magnification 100×). Insert: FISH demonstrates a DUSP22 break with 1 colocalization signal (yellow arrow) and 1 split signal (green and red arrows) consistent with gene rearrangement. FISH analysis was performed using LSI Dual color break-apart probe from Vysis abbot Molecular. B Higher magnification shows the monotonous, anaplastic cytology of the tumor cells. C Giemsa stain highlights the blastic chromatin with multiple nucleoli and abundant basophilic cytoplasm (B, C original magnification 400×). D, E The tumor cells show a strong characteristic CD30 stain (D). The cells are positive for CD3 (E) but remain negative for perforin (F), TIA1 and granzyme B (not shown), and CD5 (G) (original magnification 400×)
Follicular helper T-cell lymphoma
TFH lymphoma has distinctive clinical, morphological, immunophenotypic, and genetic features. Angioimmunoblastic T-cell lymphoma (AITL) is the prototype and best-characterized subtype [26]. The mutational landscape of TFH lymphoma includes genes involved in epigenetic pathways such as TET2 (up to 90%), IDH2 (20–45%), and DNMT3A (20–30%), mutations in the small GTPase RHOA (50–70%) and mutations in the TR signaling pathway including PLCG1, CD28, FYN, and VAV1 [105]. Recurrent translocations have also been identified including ICOS::CD28, ITK::SYK, and VAV1 fusions [23]. These genetic alterations are seen across the morphological spectrum of TFH lymphoma supporting the unifying concept of one disease entity. However, IDH2 mutations are exclusively identified in AITL with characteristic morphology (medium to large cells with clear cytoplasm) and immunophenotype (strong CD10 and CXCL13 expression) (Fig. 6) [97], whereas ITK::SYK fusions are mostly identified in the follicular type [98]. TET2 and DNMT3A are the most frequent mutations observed in clonal hematopoiesis (CH) [29], suggesting that CH plays an important role in the pathogenesis of TFH [52]. CH also explains the association of myeloid neoplasms with TFH lymphoma, particularly after cytotoxic therapy [52]. TFH lymphoma is probably the only T-cell lymphoma where mutational analysis is currently recommended to support the diagnosis [12]. In difficult cases, the presence of RHOA and/or IDH2 mutations supports the diagnosis of TFH lymphoma. In the interpretation of molecular findings, it is important to recognize the low allelic frequency often observed in ROHA and IDH2 mutations (VAF <5%) and to avoid misinterpretation of TET2 and DNMT3A as tumor-specific mutations [97].
Angioimmunoblastic T-cell lymphoma with IDH2R172K c.515G>C mutation. A H&E stain shows a lymph node with proliferation of large cells with abundant clear cytoplasm. (original magnification, 200x). Insert: the case is stained with the specific antibody against the IDH2R172K mutation. This specific variant is observed in 25–30% of all cases with IDH2R172 mutation. The paranuclear cytoplasmic positivity is characteristic of the antibody (original magnification, 400×). The variant allelic frequency of this case was 30% by NGS. B Higher magnification to show the cytology of the clear cells (original magnification, 400×). C–F The clear cells are strongly positive for CD10 (C), PD1 (D), ICOS (E), and CXCL13 (F). (original magnification, 400×)
Peripheral T-cell lymphoma, not otherwise specified
PTCL, NOS represents a heterogenous group of lymphomas with variable morphology, immunophenotype, cytogenetics and molecular features. The diagnosis of PTCL, NOS often requires the confirmation through TR clonality analysis, supported by mutational analysis [100]. Recently, 2 molecular subgroups were identified through GEP analysis, namely PTCL-TBX21 and PTCL-GATA3 [38]. PTCL-TBX21 is characterized by low genomic complexity, higher frequency of epigenetic modifier genes (TET1, TET3, DNMT3A), and more favorable outcome. In contrast, PTCL-GATA3 is associated with high genomic complexity, gains in STAT3 and MYC, frequent mutations in TP53 and PRDM1, and poor outcome [36]. A novel immunohistochemical algorithm has been proposed to reproduce these two molecular groups using 4 antibodies; TBX21, CXCR3, GATA3, and CCR4 [5]. This algorithm needs validation but likely will be integrated in routine diagnosis in the future. Cytotoxic PTCLs are included within PTCL, NOS and molecularly cluster mainly in the PTCL-TBX21 group.
Primary nodal EBV-positive T- or NK-cell lymphoma is now separated from PTCL, NOS and is considered a provisional entity in the ICC [12, 79]. By definition, it is a systemic disease without nasopharyngeal involvement and affecting mainly elderly or immunocompromised patients. The most common mutated genes are TET2, PIK3CD, DDX3X, and STAT3. A characteristic finding is recurrent losses of 14q11.2 where the TRA/D genes are located, supporting the T cell derivation of this lymphoma [69].
Genetic alterations with potential relevance for the diagnosis of extranodal and leukemic/disseminated T- and NK-cell lymphomas
Some neoplasms derived from mature T and NK cells, mostly from innate immune cells, affect predominantly extranodal sites [21]. These include extranodal NK/T-cell lymphoma (ENKTCL), nasal type, the group of gastrointestinal lymphomas, hepatosplenic T-cell lymphomas (HSTCL), and the rare group of γδ T-cell lymphomas and subcutaneous panniculitis-like T-cell lymphomas (SPTCL). Other neoplasms involve predominantly the peripheral blood and bone marrow such as T-cell large granular lymphocytic leukemia (T-LGLL), T-cell prolymphocytic leukemia (T-PLL), adult T-cell lymphoma/leukemia (ATLL) and aggressive NK-cell leukemia (ANKL). The diagnosis of these diseases is based on their characteristic clinicopathologic features. Besides TR clonality assessment (with the exception of ENKTCL if derived from NK-cells and ANKL, which lack clonal TR rearrangements), molecular analysis is not required for the diagnosis. Nevertheless, extranodal T- and NK-cell lymphomas share the activation of the JAK/STAT pathway, a potential therapeutic target [20]. In difficult differential diagnosis, the mutational pattern, although not specific, might favor one disease over another (Table 4).
Enteropathy-associated T-cell lymphoma (EATL) is characterized by frequent mutations in the JAK/STAT pathway, most commonly involving JAK1 and STAT3 and rarely JAK3, STAT5B, TYK2, or SOCS1. KMT2D and TET2 are also frequently mutated in EATL [17]. In contrast, monomorphic epitheliotropic intestinal T-cell lymphoma (MEITL) is characterized by mutations or deletions of SETD2 (>90%), resulting in reduced or absent of expression of lysine 36 in histone H3 (H3K36me3). Other characteristic co-occurrent mutations are STAT5B (<60%) and JAK3 (35–50%). The absence of H3K36me3 expression by immunohistochemistry is an excellent surrogate marker for SETD2 inactivation [81].
ENKTCL and ANKL share a similar mutational profile, frequently affecting the JAK/STAT pathway (STAT3, STAT5B, JAK3), epigenetic regulators (BCOR, KMT3D, ARID1A, EP300), tumor suppressor genes (TP53, MGA) and the RNA helicase DDX3X [21, 61, 79].
The differential diagnosis of HSTCL, T-LGLL, and γδ T-cell lymphoproliferations is not always straightforward. Characteristic genetic alterations of HSTCL are the presence of isochromosome 7q (80%) and trisomy 8 (50%) regardless of the αβ/γδ derivation [77]. Inactivating mutations in SETD2 (70%), and in other chromatin-modifying genes (INO80, TET3, SMARCA2) are frequently identified (62%). Other mutations frequently observed are STAT5B (31%), STAT3 and PIK3CD and rare cases also carry TP53, URB5, and IDH2 (5–10%) mutations [20]. In contrast, T-LGLL carry STAT3 mutations in up to 50% of the cases and co-occur frequently with KMT2D and SETD1B mutations [14]. STAT5B mutations are mainly associated with the indolent CD4+ T-LGLL and a subgroup of CD8+ aggressive T-LGLL [8].
T-PLL is characterized by 14q chromosomal inversions with breakpoints in the long arm at q11 and q32 seen in 80% of the cases. These alterations involve the TCL1 family genes and less often MTCP1 gene, which is homologous to TCL1A/B at Xq28. Recurrent mutations in the JAK/STAT pathway are observed up to 75% of the cases. Monoallelic deletions and/or mutations of ATM are common [20, 99]. Demonstration of TCL1 chromosomal alteration by FISH or TCL1 expression in T-cells by immunohistochemistry is diagnostic of T-PLL.
Germline mutations of the HAVCR2 gene have been demonstrated in patients with SPTCL [28]. Two hot spot mutations have been identified, one predominates in East Asia (HAVCR2Y82C) and the other in Europeans (HAVCR2I97M). HAVCR2 encodes for TIM3, an inhibitory receptor expressed on interferon-γ producing T-cells involved in regulating inflammation. Patients with HAVCR2 mutation are younger (<30 years), often associated with hemophagocytic lymphohistiocytosis (HLH) and worse prognosis [95]. The investigation of HAVCR2 mutations may allow risk stratification and provide a target for therapy.
Conclusions and outlook
As evidenced by the increasing emphasis on molecular features in contemporary lymphoma classifications, molecular diagnostics is gaining in importance for everyday decision-making both for the pathologist and the treating clinician [2, 12, 20]. Although many lymphomas still can be classified reliably with the morphology and immunophenotyping and in part investigation of single markers, we will see an increasing use of multigene panels or whole exome sequencing for detailed genetic profiling and probably a replacement of different techniques such as PCR and FISH by a comprehensive NGS-based approach. Newer techniques including whole genome and single-cell sequencing and methylation and chromatin profiling likely will also enter clinical practice for specific applications [21]. Detection of cell-free tumor DNA in the peripheral blood, so-called liquid biopsy shows promise for the follow-up of patients with aggressive lymphomas under treatment, although sensitivity and standardization issues and potential pitfalls such as clonal evolution and clonal hematopoiesis require more work before broad clinical application [83].
References
Agathangelidis A, Chatzidimitriou A, Chatzikonstantinou T, Tresoldi C, Davis Z, Giudicelli V, Kossida S, Belessi C, Rosenquist R, Ghia P, Langerak AW, Davi F, Stamatopoulos K, on behalf of ERIC, the European Research Initiative on CLL (2022) Immunoglobulin gene sequence analysis in chronic lymphocytic leukemia: the 2022 update of the recommendations by ERIC, the European Research Initiative on CLL. Leukemia 36:1961–1968. https://doi.org/10.1038/s41375-022-01604-2
Alaggio R, Amador C, Anagnostopoulos I, Attygalle AD, Araujo IBO, Berti E, Bhagat G, Borges AM, Boyer D, Calaminici M, Chadburn A, Chan JKC, Cheuk W, Chng WJ, Choi JK, Chuang SS, Coupland SE, Czader M, Dave SS et al (2022) The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 36:1720–1748. https://doi.org/10.1038/s41375-022-01620-2
Alduaij W, Collinge B, Ben-Neriah S, Jiang A, Hilton LK, Boyle M, Meissner B, Chong L, Miyata-Takata T, Slack GW, Farinha P, Craig JW, Lytle A, Savage KJ, Villa D, Gerrie AS, Freeman CL, Gascoyne RD, Connors JM et al (2023) Molecular determinants of clinical outcomes in a real-world diffuse large B-cell lymphoma population. Blood 141:2493–2507. https://doi.org/10.1182/blood.2022018248
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson J Jr, Lu L, Lewis DB, Tibshirani R, Sherlock G et al (2000) Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403:503–511. https://doi.org/10.1038/35000501
Amador C, Greiner TC, Heavican TB, Smith LM, Galvis KT, Lone W, Bouska A, D'Amore F, Pedersen MB, Pileri S, Agostinelli C, Feldman AL, Rosenwald A, Ott G, Mottok A, Savage KJ, de Leval L, Gaulard P, Lim ST et al (2019) Reproducing the molecular subclassification of peripheral T-cell lymphoma-NOS by immunohistochemistry. Blood 134:2159–2170. https://doi.org/10.1182/blood.2019000779
Arcila ME, Yu W, Syed M, Kim H, Maciag L, Yao J, Ho C, Petrova K, Moung C, Salazar P, Rijo I, Baldi T, Zehir A, Landgren O, Park J, Roshal M, Dogan A, Nafa K (2019) Establishment of immunoglobulin heavy (IGH) chain clonality testing by next-generation sequencing for routine characterization of B-cell and plasma cell neoplasms. J Mol Diagn 21:330–342. https://doi.org/10.1016/j.jmoldx.2018.10.008
Beishuizen A, Verhoeven MA, van Wering ER, Hahlen K, Hooijkaas H, van Dongen JJ (1994) Analysis of Ig and T-cell receptor genes in 40 childhood acute lymphoblastic leukemias at diagnosis and subsequent relapse: implications for the detection of minimal residual disease by polymerase chain reaction analysis. Blood 83:2238–2247
Bhattacharya D, Teramo A, Gasparini VR, Huuhtanen J, Kim D, Theodoropoulos J, Schiavoni G, Barila G, Vicenzetto C, Calabretto G, Facco M, Kawakami T, Nakazawa H, Falini B, Tiacci E, Ishida F, Semenzato G, Kelkka T, Zambello R, Mustjoki S (2022) Identification of novel STAT5B mutations and characterization of TCRbeta signatures in CD4+ T-cell large granular lymphocyte leukemia Blood. Cancer J 12:31. https://doi.org/10.1038/s41408-022-00630-8
Bonzheim I, Geissinger E, Roth S, Zettl A, Marx A, Rosenwald A, Muller-Hermelink HK, Rudiger T (2004) Anaplastic large cell lymphomas lack the expression of T-cell receptor molecules or molecules of proximal T-cell receptor signaling. Blood 104:3358–3360. https://doi.org/10.1182/blood-2004-03-1037
Bruggemann M, Kotrova M, Knecht H, Bartram J, Boudjogrha M, Bystry V, Fazio G, Fronkova E, Giraud M, Grioni A, Hancock J, Herrmann D, Jimenez C, Krejci A, Moppett J, Reigl T, Salson M, Scheijen B, Schwarz M et al (2019) Standardized next-generation sequencing of immunoglobulin and T-cell receptor gene recombinations for MRD marker identification in acute lymphoblastic leukaemia; a EuroClonality-NGS validation study. Leukemia 33:2241–2253. https://doi.org/10.1038/s41375-019-0496-7
Bystry V, Reigl T, Krejci A, Demko M, Hanakova B, Grioni A, Knecht H, Schlitt M, Dreger P, Sellner L, Herrmann D, Pingeon M, Boudjoghra M, Rijntjes J, Pott C, Langerak AW, Groenen P, Davi F, Bruggemann M et al (2017) ARResT/Interrogate: an interactive immunoprofiler for IG/TR NGS data. Bioinformatics 33:435–437. https://doi.org/10.1093/bioinformatics/btw634
Campo E, Jaffe ES, Cook JR, Quintanilla-Martinez L, Swerdlow SH, Anderson KC, Brousset P, Cerroni L, de Leval L, Dirnhofer S, Dogan A, Feldman AL, Fend F, Friedberg JW, Gaulard P, Ghia P, Horwitz SM, King RL, Salles G et al (2022) The International Consensus Classification of Mature Lymphoid Neoplasms: a report from the Clinical Advisory Committee. Blood 140:1229–1253. https://doi.org/10.1182/blood.2022015851
Chapuy B, Stewart C, Dunford AJ, Kim J, Kamburov A, Redd RA, Lawrence MS, Roemer MGM, Li AJ, Ziepert M, Staiger AM, Wala JA, Ducar MD, Leshchiner I, Rheinbay E, Taylor-Weiner A, Coughlin CA, Hess JM, Pedamallu CS et al (2018) Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med 24:679–690. https://doi.org/10.1038/s41591-018-0016-8
Cheon H, Xing JC, Moosic KB, Ung J, Chan VW, Chung DS, Toro MF, Elghawy O, Wang JS, Hamele CE, Hardison RC, Olson TL, Tan SF, Feith DJ, Ratan A, Loughran TP (2022) Genomic landscape of TCRalphabeta and TCRgammadelta T-large granular lymphocyte leukemia. Blood 139:3058–3072. https://doi.org/10.1182/blood.2021013164
Chiarle R, Simmons WJ, Cai H, Dhall G, Zamo A, Raz R, Karras JG, Levy DE, Inghirami G (2005) Stat3 is required for ALK-mediated lymphomagenesis and provides a possible therapeutic target. Nat Med 11:623–629. https://doi.org/10.1038/nm1249
Chong LC, Ben-Neriah S, Slack GW, Freeman C, Ennishi D, Mottok A, Collinge B, Abrisqueta P, Farinha P, Boyle M, Meissner B, Kridel R, Gerrie AS, Villa D, Savage KJ, Sehn LH, Siebert R, Morin RD, Gascoyne RD et al (2018) High-resolution architecture and partner genes of MYC rearrangements in lymphoma with DLBCL morphology. Blood Adv 2:2755–2765. https://doi.org/10.1182/bloodadvances.2018023572
Cording S, Lhermitte L, Malamut G, Berrabah S, Trinquand A, Guegan N, Villarese P, Kaltenbach S, Meresse B, Khater S, Dussiot M, Bras M, Cheminant M, Tesson B, Bole-Feysot C, Bruneau J, Molina TJ, Sibon D, Macintyre E et al (2022) Oncogenetic landscape of lymphomagenesis in coeliac disease. Gut 71:497–508. https://doi.org/10.1136/gutjnl-2020-322935
Darzentas N (2022) ARResT/Interrogate Immunoprofiling Platform: Concepts, Workflows, and Insights. Methods Mol Biol 2453:571–584. https://doi.org/10.1007/978-1-0716-2115-8_26
Davies A, Cummin TE, Barrans S, Maishman T, Mamot C, Novak U, Caddy J, Stanton L, Kazmi-Stokes S, McMillan A, Fields P, Pocock C, Collins GP, Stephens R, Cucco F, Clipson A, Sha C, Tooze R, Care MA et al (2019) Gene-expression profiling of bortezomib added to standard chemoimmunotherapy for diffuse large B-cell lymphoma (REMoDL-B): an open-label, randomised, phase 3 trial. Lancet Oncol 20:649–662. https://doi.org/10.1016/S1470-2045(18)30935-5
de Leval L, Alizadeh AA, Bergsagel PL, Campo E, Davies A, Dogan A, Fitzgibbon J, Horwitz SM, Melnick AM, Morice WG, Morin RD, Nadel B, Pileri SA, Rosenquist R, Rossi D, Salaverria I, Steidl C, Treon SP, Zelenetz AD et al (2022) Genomic profiling for clinical decision making in lymphoid neoplasms. Blood 140:2193–2227. https://doi.org/10.1182/blood.2022015854
de Leval L, Feldman AL, Pileri S, Nakamura S, Gaulard P (2023) Extranodal T- and NK-cell lymphomas. Virchows Arch 482:245–264. https://doi.org/10.1007/s00428-022-03434-0
Dogliotti I, Jimenez C, Varettoni M, Talaulikar D, Bagratuni T, Ferrante M, Perez J, Drandi D, Puig N, Gilestro M, Garcia-Alvarez M, Owen R, Jurczak W, Tedeschi A, Leblond V, Kastritis E, Kersten MJ, D'Sa S, Kascak M et al (2023) Diagnostics in Waldenstrom's macroglobulinemia: a consensus statement of the European Consortium for Waldenstrom's Macroglobulinemia. Leukemia 37:388–395. https://doi.org/10.1038/s41375-022-01762-3
Drieux F, Ruminy P, Sater V, Marchand V, Fataccioli V, Lanic MD, Viennot M, Viailly PJ, Sako N, Robe C, Dupuy A, Vallois D, Veresezan L, Poullot E, Picquenot JM, Bossard C, Parrens M, Lemonnier F, Jardin F et al (2021) Detection of gene fusion transcripts in peripheral T-cell lymphoma using a multiplexed targeted sequencing assay. J Mol Diagn 23:929–940. https://doi.org/10.1016/j.jmoldx.2021.04.013
Ennishi D, Jiang A, Boyle M, Collinge B, Grande BM, Ben-Neriah S, Rushton C, Tang J, Thomas N, Slack GW, Farinha P, Takata K, Miyata-Takata T, Craig J, Mottok A, Meissner B, Saberi S, Bashashati A, Villa D et al (2019) Double-hit gene expression signature defines a distinct subgroup of germinal center B-cell-like diffuse large B-cell lymphoma. J Clin Oncol 37:190–201. https://doi.org/10.1200/JCO.18.01583
Falini B, Lamant-Rochaix L, Campo E, Jaffe ES, Gascoyne RD, Stein H, Müller-Hermelink HK, Kinney MC (2017) Anaplastic large cell lymphoma, ALK-positive. In: Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri S, Stein H, Thiele J (eds) WHO classification of tumours of haematopoetic and lymphoid tissues. IARC, Lyon, France, pp 413–418
Feldman AL, Laurent C, Narbaitz M, Nakamura S, Chan WC, de Leval L, Gaulard P (2023) Classification and diagnostic evaluation of nodal T- and NK-cell lymphomas. Virchows Arch 482:265–279. https://doi.org/10.1007/s00428-022-03412-6
Fend F, Dogan A, Cook JR (2023) Plasma cell neoplasms and related entities-evolution in diagnosis and classification. Virchows Arch 482:163–177. https://doi.org/10.1007/s00428-022-03431-3
Gayden T, Sepulveda FE, Khuong-Quang DA, Pratt J, Valera ET, Garrigue A, Kelso S, Sicheri F, Mikael LG, Hamel N, Bajic A, Dali R, Deshmukh S, Dervovic D, Schramek D, Guerin F, Taipale M, Nikbakht H, Majewski J et al (2018) Germline HAVCR2 mutations altering TIM-3 characterize subcutaneous panniculitis-like T cell lymphomas with hemophagocytic lymphohistiocytic syndrome. Nat Genet 50:1650–1657. https://doi.org/10.1038/s41588-018-0251-4
Genovese G, Kahler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, Chambert K, Mick E, Neale BM, Fromer M, Purcell SM, Svantesson O, Landen M, Hoglund M, Lehmann S, Gabriel SB, Moran JL, Lander ES, Sullivan PF et al (2014) Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med 371:2477–2487. https://doi.org/10.1056/NEJMoa1409405
Glenn ST, Galbo PM Jr, Luce JD, Miles KM, Singh PK, Glynias MJ, Morrison C (2023) Development and implementation of an automated and highly accurate reporting process for NGS-based clonality testing. Oncotarget 14:450–461. https://doi.org/10.18632/oncotarget.28429
Gonzalez-Farre B, Ramis-Zaldivar JE, Salmeron-Villalobos J, Balague O, Celis V, Verdu-Amoros J, Nadeu F, Sabado C, Ferrandez A, Garrido M, Garcia-Bragado F, de la Maya MD, Vagace JM, Panizo CM, Astigarraga I, Andres M, Jaffe ES, Campo E, Salaverria I (2019) Burkitt-like lymphoma with 11q aberration: a germinal center-derived lymphoma genetically unrelated to Burkitt lymphoma. Haematologica 104:1822–1829. https://doi.org/10.3324/haematol.2018.207928
Goodlad JR, Xiao W, Amador C, Cook JR, Happ L, Thakkar D, Dave S, Dogan A, Duffield A, Nejati R, Ott G, Wasik M, Czader M (2023) Phenotypic and genotypic infidelity in B-lineage neoplasms, including transdifferentiation following targeted therapy: Report from the 2021 SH/EAHP Workshop. Am J Clin Pathol 159:538–553. https://doi.org/10.1093/ajcp/aqad035
Grande BM, Gerhard DS, Jiang A, Griner NB, Abramson JS, Alexander TB, Allen H, Ayers LW, Bethony JM, Bhatia K, Bowen J, Casper C, Choi JK, Culibrk L, Davidsen TM, Dyer MA, Gastier-Foster JM, Gesuwan P, Greiner TC et al (2019) Genome-wide discovery of somatic coding and noncoding mutations in pediatric endemic and sporadic Burkitt lymphoma. Blood 133:1313–1324. https://doi.org/10.1182/blood-2018-09-871418
Groenen P, van den Brand M, Kroeze LI, Amir AL, Hebeda KM (2023) Read the clonotype: Next-generation sequencing-based lymphocyte clonality analysis and perspectives for application in pathology Front. Oncol 13:1107171. https://doi.org/10.3389/fonc.2023.1107171
Hans CP, Weisenburger DD, Greiner TC, Gascoyne RD, Delabie J, Ott G, Muller-Hermelink HK, Campo E, Braziel RM, Jaffe ES, Pan Z, Farinha P, Smith LM, Falini B, Banham AH, Rosenwald A, Staudt LM, Connors JM, Armitage JO, Chan WC (2004) Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 103:275–282. https://doi.org/10.1182/blood-2003-05-1545
Heavican TB, Bouska A, Yu J, Lone W, Amador C, Gong Q, Zhang W, Li Y, Dave BJ, Nairismagi ML, Greiner TC, Vose J, Weisenburger DD, Lachel C, Wang C, Fu K, Stevens JM, Lim ST, Ong CK et al (2019) Genetic drivers of oncogenic pathways in molecular subgroups of peripheral T-cell lymphoma. Blood 133:1664–1676. https://doi.org/10.1182/blood-2018-09-872549
Hilton LK, Tang J, Ben-Neriah S, Alcaide M, Jiang A, Grande BM, Rushton CK, Boyle M, Meissner B, Scott DW, Morin RD (2019) The double-hit signature identifies double-hit diffuse large B-cell lymphoma with genetic events cryptic to FISH. Blood 134:1528–1532. https://doi.org/10.1182/blood.2019002600
Iqbal J, Wright G, Wang C, Rosenwald A, Gascoyne RD, Weisenburger DD, Greiner TC, Smith L, Guo S, Wilcox RA, Teh BT, Lim ST, Tan SY, Rimsza LM, Jaffe ES, Campo E, Martinez A, Delabie J, Braziel RM et al (2014) Gene expression signatures delineate biological and prognostic subgroups in peripheral T-cell lymphoma. Blood 123:2915–2923. https://doi.org/10.1182/blood-2013-11-536359
Jaffe ES, Harris NL, Stein H, Isaacson PG (2008) Classification of lymphoid neoplasms: the microscope as a tool for disease discovery. Blood 112:4384–4399. https://doi.org/10.1182/blood-2008-07-077982
Jiang Y, Dominguez PM, Melnick AM (2016) The many layers of epigenetic dysfunction in B-cell lymphomas. Curr Opin Hematol 23:377–384. https://doi.org/10.1097/MOH.0000000000000249
King RL, Hsi ED, Chan WC, Piris MA, Cook JR, Scott DW, Swerdlow SH (2023) Diagnostic approaches and future directions in Burkitt lymphoma and high-grade B-cell lymphoma. Virchows Arch 482:193–205. https://doi.org/10.1007/s00428-022-03404-6
King RL, McPhail ED, Meyer RG, Vasmatzis G, Pearce K, Smadbeck JB, Ketterling RP, Smoley SA, Greipp PT, Hoppman NL, Peterson JF, Baughn LB (2019) False-negative rates for MYC fluorescence in situ hybridization probes in B-cell neoplasms. Haematologica 104:e248–e251. https://doi.org/10.3324/haematol.2018.207290
Kroeze LI, Scheijen B, Hebeda KM, Rijntjes J, Luijks J, Evers D, Hobo W, Groenen P, van den Brand M (2022) PAX5 P80R-mutated B-cell acute lymphoblastic leukemia with transformation to histiocytic sarcoma: clonal evolution assessment using NGS-based immunoglobulin clonality and mutation analysis. Virchows Arch. https://doi.org/10.1007/s00428-022-03428-y
Lackraj T, Goswami R, Kridel R (2018) Pathogenesis of follicular lymphoma. Best Pract Res Clin Haematol 31:2–14. https://doi.org/10.1016/j.beha.2017.10.006
Lacy SE, Barrans SL, Beer PA, Painter D, Smith AG, Roman E, Cooke SL, Ruiz C, Glover P, Van Hoppe SJL, Webster N, Campbell PJ, Tooze RM, Patmore R, Burton C, Crouch S, Hodson DJ (2020) Targeted sequencing in DLBCL, molecular subtypes, and outcomes: a Haematological Malignancy Research Network report. Blood 135:1759–1771. https://doi.org/10.1182/blood.2019003535
Langerak AW, Groenen PJ, Bruggemann M, Beldjord K, Bellan C, Bonello L, Boone E, Carter GI, Catherwood M, Davi F, Delfau-Larue MH, Diss T, Evans PA, Gameiro P, Garcia Sanz R, Gonzalez D, Grand D, Hakansson A, Hummel M et al (2012) EuroClonality/BIOMED-2 guidelines for interpretation and reporting of Ig/TCR clonality testing in suspected lymphoproliferations. Leukemia 26:2159–2171. https://doi.org/10.1038/leu.2012.246
Langerak AW, Molina TJ, Lavender FL, Pearson D, Flohr T, Sambade C, Schuuring E, Al Saati T, van Dongen JJ, van Krieken JH (2007) Polymerase chain reaction-based clonality testing in tissue samples with reactive lymphoproliferations: usefulness and pitfalls. A report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 21:222–229. https://doi.org/10.1038/sj.leu.2404482
Larose H, Prokoph N, Matthews JD, Schlederer M, Hogler S, Alsulami AF, Ducray SP, Nuglozeh E, FMS F, Elmouna A, Ceccon M, Mologni L, Gambacorti-Passerini C, Hoefler G, Lobello C, Pospisilova S, Janikova A, Woessmann W, Damm-Welk C et al (2021) Whole Exome Sequencing reveals NOTCH1 mutations in anaplastic large cell lymphoma and points to Notch both as a key pathway and a potential therapeutic target. Haematologica 106:1693–1704. https://doi.org/10.3324/haematol.2019.238766
Laurent C, Cook JR, Yoshino T, Quintanilla-Martinez L, Jaffe ES (2023) Follicular lymphoma and marginal zone lymphoma: how many diseases? Virchows Arch 482:149–162. https://doi.org/10.1007/s00428-022-03432-2
Lee SE, Kang SY, Yoo HY, Kim SJ, Kim WS, Ko YH (2016) Clonal relationships in recurrent B-cell lymphomas. Oncotarget 7:12359–12371. https://doi.org/10.18632/oncotarget.7132
Leoncini L (2022) Epstein-Barr virus positivity as a defining pathogenetic feature of Burkitt lymphoma subtypes. Br J Haematol 196:468–470. https://doi.org/10.1111/bjh.17922
Lewis NE, Petrova-Drus K, Huet S, Epstein-Peterson ZD, Gao Q, Sigler AE, Baik J, Ozkaya N, Moskowitz AJ, Kumar A, Horwitz SM, Zhang Y, Arcila ME, Levine RL, Roshal M, Dogan A, Xiao W (2020) Clonal hematopoiesis in angioimmunoblastic T-cell lymphoma with divergent evolution to myeloid neoplasms. Blood Adv 4:2261–2271. https://doi.org/10.1182/bloodadvances.2020001636
Lobello C, Tichy B, Bystry V, Radova L, Filip D, Mraz M, Montes-Mojarro IA, Prokoph N, Larose H, Liang HC, Sharma GG, Mologni L, Belada D, Kamaradova K, Fend F, Gambacorti-Passerini C, Merkel O, Turner SD, Janikova A, Pospisilova S (2021) STAT3 and TP53 mutations associate with poor prognosis in anaplastic large cell lymphoma. Leukemia 35:1500–1505. https://doi.org/10.1038/s41375-020-01093-1
Lopez C, Kleinheinz K, Aukema SM, Rohde M, Bernhart SH, Hubschmann D, Wagener R, Toprak UH, Raimondi F, Kreuz M, Waszak SM, Huang Z, Sieverling L, Paramasivam N, Seufert J, Sungalee S, Russell RB, Bausinger J, Kretzmer H et al (2019) Genomic and transcriptomic changes complement each other in the pathogenesis of sporadic Burkitt lymphoma. Nat Commun 10:1459. https://doi.org/10.1038/s41467-019-08578-3
Luchtel RA, Dasari S, Oishi N, Pedersen MB, Hu G, Rech KL, Ketterling RP, Sidhu J, Wang X, Katoh R, Dogan A, Kip NS, Cunningham JM, Sun Z, Baheti S, Porcher JC, Said JW, Jiang L, Hamilton-Dutoit SJ et al (2018) Molecular profiling reveals immunogenic cues in anaplastic large cell lymphomas with DUSP22 rearrangements. Blood 132:1386–1398. https://doi.org/10.1182/blood-2018-03-838524
Luchtel RA, Zimmermann MT, Hu G, Dasari S, Jiang M, Oishi N, Jacobs HK, Zeng Y, Hundal T, Rech KL, Ketterling RP, Lee JH, Eckloff BW, Yan H, Gaonkar KS, Tian S, Ye Z, Kadin ME, Sidhu J et al (2019) Recurrent MSC (E116K) mutations in ALK-negative anaplastic large cell lymphoma. Blood 133:2776–2789. https://doi.org/10.1182/blood.2019000626
Martin-Garcia D, Navarro A, Valdes-Mas R, Clot G, Gutierrez-Abril J, Prieto M, Ribera-Cortada I, Woroniecka R, Rymkiewicz G, Bens S, de Leval L, Rosenwald A, Ferry JA, Hsi ED, Fu K, Delabie J, Weisenburger D, de Jong D, Climent F, O'Connor SJ, Swerdlow SH, Torrents D, Beltran S, Espinet B, Gonzalez-Farre B, Veloza L, Costa D, Matutes E, Siebert R, Ott G, Quintanilla-Martinez L, Jaffe ES, Lopez-Otin C, Salaverria I, Puente XS, Campo E, Bea S (2019) CCND2 and CCND3 hijack immunoglobulin light-chain enhancers in cyclin D1(-) mantle cell lymphoma Blood 133:940-9951. doi: https://doi.org/10.1182/blood-2018-07-862151
McPhail ED, Maurer MJ, Macon WR, Feldman AL, Kurtin PJ, Ketterling RP, Vaidya R, Cerhan JR, Ansell SM, Porrata LF, Nowakowski GS, Witzig TE, Habermann TM (2018) Inferior survival in high-grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements is not associated with MYC/IG gene rearrangements. Haematologica 103:1899–1907. https://doi.org/10.3324/haematol.2018.190157
Meyer PN, Fu K, Greiner TC, Smith LM, Delabie J, Gascoyne RD, Ott G, Rosenwald A, Braziel RM, Campo E, Vose JM, Lenz G, Staudt LM, Chan WC, Weisenburger DD (2011) Immunohistochemical methods for predicting cell of origin and survival in patients with diffuse large B-cell lymphoma treated with rituximab. J Clin Oncol 29:200–207. https://doi.org/10.1200/JCO.2010.30.0368
Mishina T, Oshima-Hasegawa N, Tsukamoto S, Fukuyo M, Kageyama H, Muto T, Mimura N, Rahmutulla B, Nagai Y, Kayamori K, Hino Y, Mitsukawa S, Takeda Y, Ohwada C, Takeuchi M, Tsujimura H, Iseki T, Nakaseko C, Ikeda JI et al (2021) Genetic subtype classification using a simplified algorithm and mutational characteristics of diffuse large B-cell lymphoma in a Japanese cohort. Br J Haematol 195:731–742. https://doi.org/10.1111/bjh.17765
Montes-Mojarro IA, Chen BJ, Ramirez-Ibarguen AF, Quezada-Fiallos CM, Perez-Baez WB, Duenas D, Casavilca-Zambrano S, Ortiz-Mayor M, Rojas-Bilbao E, Garcia-Rivello H, Metrebian MF, Narbaitz M, Barrionuevo C, Lome-Maldonado C, Bonzheim I, Fend F, Steinhilber J, Quintanilla-Martinez L (2020) Mutational profile and EBV strains of extranodal NK/T-cell lymphoma, nasal type in Latin America. Mod Pathol 33:781–791. https://doi.org/10.1038/s41379-019-0415-5
Morschhauser F, Tilly H, Chaidos A, McKay P, Phillips T, Assouline S, Batlevi CL, Campbell P, Ribrag V, Damaj GL, Dickinson M, Jurczak W, Kazmierczak M, Opat S, Radford J, Schmitt A, Yang J, Whalen J, Agarwal S et al (2020) Tazemetostat for patients with relapsed or refractory follicular lymphoma: an open-label, single-arm, multicentre, phase 2 trial. Lancet Oncol 21:1433–1442. https://doi.org/10.1016/S1470-2045(20)30441-1
Munoz-Marmol AM, Sanz C, Tapia G, Marginet R, Ariza A, Mate JL (2013) MYC status determination in aggressive B-cell lymphoma: the impact of FISH probe selection. Histopathology 63:418–424. https://doi.org/10.1111/his.12178
Nadeu F, Diaz-Navarro A, Delgado J, Puente XS, Campo E (2020) Genomic and Epigenomic Alterations in Chronic Lymphocytic Leukemia. Annual Review of Pathology: Mechanisms of Disease 15:149–177. https://doi.org/10.1146/annurev-pathmechdis-012419-032810
Nadeu F, Martin-Garcia D, Clot G, Diaz-Navarro A, Duran-Ferrer M, Navarro A, Vilarrasa-Blasi R, Kulis M, Royo R, Gutierrez-Abril J, Valdes-Mas R, Lopez C, Chapaprieta V, Puiggros M, Castellano G, Costa D, Aymerich M, Jares P, Espinet B et al (2020) Genomic and epigenomic insights into the origin, pathogenesis, and clinical behavior of mantle cell lymphoma subtypes. Blood 136:1419–1432. https://doi.org/10.1182/blood.2020005289
Nadeu F, Royo R, Clot G, Duran-Ferrer M, Navarro A, Martin S, Lu J, Zenz T, Baumann T, Jares P, Puente XS, Martin-Subero JI, Delgado J, Campo E (2021) IGLV3-21R110 identifies an aggressive biological subtype of chronic lymphocytic leukemia with intermediate epigenetics. Blood 137:2935–2946. https://doi.org/10.1182/blood.2020008311
Nann D, Ramis-Zaldivar JE, Muller I, Gonzalez-Farre B, Schmidt J, Egan C, Salmeron-Villalobos J, Clot G, Mattern S, Otto F, Mankel B, Colomer D, Balague O, Szablewski V, Lome-Maldonado C, Leoncini L, Dojcinov S, Chott A, Copie-Bergman C et al (2020) Follicular lymphoma t(14;18)-negative is genetically a heterogeneous disease. Blood Adv 4:5652–5665. https://doi.org/10.1182/bloodadvances.2020002944
Newman AM, Zaka M, Zhou P, Blain AE, Erhorn A, Barnard A, Crossland RE, Wilkinson S, Enshaei A, De Zordi J, Harding F, Taj M, Wood KM, Televantou D, Turner SD, Burke GAA, Harrison CJ, Bomken S, Bacon CM, Rand V (2022) Genomic abnormalities of TP53 define distinct risk groups of paediatric B-cell non-Hodgkin lymphoma. Leukemia 36:781–789. https://doi.org/10.1038/s41375-021-01444-6
Ng SB, Chung TH, Kato S, Nakamura S, Takahashi E, Ko YH, Khoury JD, Yin CC, Soong R, Jeyasekharan AD, Hoppe MM, Selvarajan V, Tan SY, Lim ST, Ong CK, Nairismagi ML, Maheshwari P, Choo SN, Fan S et al (2018) Epstein-Barr virus-associated primary nodal T/NK-cell lymphoma shows a distinct molecular signature and copy number changes. Haematologica 103:278–287. https://doi.org/10.3324/haematol.2017.180430
Nishiuchi R, Yoshino T, Teramoto N, Sakuma I, Hayashi K, Nakamura S, Seino Y, Akagi T (1996) Clonal analysis by polymerase chain reaction of B-cell lymphoma with late relapse: a report of five cases. Cancer 77:757–762
Nollet F, Vanhouteghem K, Vermeire S, Maelbrancke E, Emmerechts J, Devos H, Cauwelier B (2019) Evaluation of next-generation sequencing-based clonality analysis of T-cell receptor gamma gene rearrangements based on a new interpretation algorithm. Int J Lab Hematol 41:242–249. https://doi.org/10.1111/ijlh.12954
Nowakowski GS, Chiappella A, Gascoyne RD, Scott DW, Zhang Q, Jurczak W, Ozcan M, Hong X, Zhu J, Jin J, Belada D, Bergua JM, Piazza F, Mocikova H, Molinari AL, Yoon DH, Cavallo F, Tani M, Yamamoto K et al (2021) ROBUST: A Phase III Study of Lenalidomide Plus R-CHOP Versus Placebo Plus R-CHOP in Previously Untreated Patients With ABC-Type Diffuse Large B-Cell Lymphoma. J Clin Oncol 39:1317–1328. https://doi.org/10.1200/JCO.20.01366
Parrilla Castellar ER, Jaffe ES, Said JW, Swerdlow SH, Ketterling RP, Knudson RA, Sidhu JS, Hsi ED, Karikehalli S, Jiang L, Vasmatzis G, Gibson SE, Ondrejka S, Nicolae A, Grogg KL, Allmer C, Ristow KM, Wilson WH, Macon WR, Law ME, Cerhan JR, Habermann TM, Ansell SM, Dogan A, Maurer MJ, Feldman AL (2014) ALK-negative anaplastic large cell lymphoma is a genetically heterogeneous disease with widely disparate clinical outcomes Blood 124:1473-1480. doi: https://doi.org/10.1182/blood-2014-04-571091
Parry M, Rose-Zerilli MJ, Ljungstrom V, Gibson J, Wang J, Walewska R, Parker H, Parker A, Davis Z, Gardiner A, McIver-Brown N, Kalpadakis C, Xochelli A, Anagnostopoulos A, Fazi C, de Castro DG, Dearden C, Pratt G, Rosenquist R et al (2015) Genetics and Prognostication in Splenic Marginal Zone Lymphoma: Revelations from Deep Sequencing. Clin Cancer Res 21:4174–4183. https://doi.org/10.1158/1078-0432.CCR-14-2759
Pastore A, Jurinovic V, Kridel R, Hoster E, Staiger AM, Szczepanowski M, Pott C, Kopp N, Murakami M, Horn H, Leich E, Moccia AA, Mottok A, Sunkavalli A, Van Hummelen P, Ducar M, Ennishi D, Shulha HP, Hother C et al (2015) Integration of gene mutations in risk prognostication for patients receiving first-line immunochemotherapy for follicular lymphoma: a retrospective analysis of a prospective clinical trial and validation in a population-based registry. Lancet Oncol 16:1111–1122. https://doi.org/10.1016/S1470-2045(15)00169-2
Pillonel V, Juskevicius D, Ng CKY, Bodmer A, Zettl A, Jucker D, Dirnhofer S, Tzankov A (2018) High-throughput sequencing of nodal marginal zone lymphomas identifies recurrent BRAF mutations. Leukemia 32:2412–2426. https://doi.org/10.1038/s41375-018-0082-4
Pro B, Allen P, Behdad A (2020) Hepatosplenic T-cell lymphoma: a rare but challenging entity. Blood 136:2018–2026. https://doi.org/10.1182/blood.2019004118
Quintanilla-Martinez L, Pittaluga S, Miething C, Klier M, Rudelius M, Davies-Hill T, Anastasov N, Martinez A, Vivero A, Duyster J, Jaffe ES, Fend F, Raffeld M (2006) NPM-ALK-dependent expression of the transcription factor CCAAT/enhancer binding protein beta in ALK-positive anaplastic large cell lymphoma. Blood 108:2029–2036. https://doi.org/10.1182/blood-2005-10-014258
Quintanilla-Martinez L, Swerdlow SH, Tousseyn T, Barrionuevo C, Nakamura S, Jaffe ES (2023) New concepts in EBV-associated B, T, and NK cell lymphoproliferative disorders. Virchows Arch 482:227–244. https://doi.org/10.1007/s00428-022-03414-4
Ramis-Zaldivar JE, Gonzalez-Farre B, Balague O, Celis V, Nadeu F, Salmeron-Villalobos J, Andres M, Martin-Guerrero I, Garrido-Pontnou M, Gaafar A, Sunol M, Barcena C, Garcia-Bragado F, Andion M, Azorin D, Astigarraga I, Sagaseta de Ilurdoz M, Sabado C, Gallego S et al (2020) Distinct molecular profile of IRF4-rearranged large B-cell lymphoma. Blood 135:274–286. https://doi.org/10.1182/blood.2019002699
Roberti A, Dobay MP, Bisig B, Vallois D, Boechat C, Lanitis E, Bouchindhomme B, Parrens MC, Bossard C, Quintanilla-Martinez L, Missiaglia E, Gaulard P, de Leval L (2016) Type II enteropathy-associated T-cell lymphoma features a unique genomic profile with highly recurrent SETD2 alterations. Nat Commun 7:12602. https://doi.org/10.1038/ncomms12602
Rohde M, Bonn BR, Zimmermann M, Lange J, Moricke A, Klapper W, Oschlies I, Szczepanowski M, Nagel I, Schrappe M, Project M-M-S, Project IM-S, Loeffler M, Siebert R, Reiter A, Burkhardt B (2017) Relevance of ID3-TCF3-CCND3 pathway mutations in pediatric aggressive B-cell lymphoma treated according to the non-Hodgkin Lymphoma Berlin-Frankfurt-Munster protocols. Haematologica 102:1091–1098. https://doi.org/10.3324/haematol.2016.156885
Roschewski M, Rossi D, Kurtz DM, Alizadeh AA, Wilson WH (2022) Circulating Tumor DNA in lymphoma: principles and future directions. Blood Cancer Discov 3:5–15. https://doi.org/10.1158/2643-3230.BCD-21-0029
Rosenwald A, Bens S, Advani R, Barrans S, Copie-Bergman C, Elsensohn MH, Natkunam Y, Calaminici M, Sander B, Baia M, Smith A, Painter D, Pham L, Zhao S, Ziepert M, Jordanova ES, Molina TJ, Kersten MJ, Kimby E et al (2019) Prognostic Significance of MYC Rearrangement and Translocation Partner in Diffuse Large B-Cell Lymphoma: A Study by the Lunenburg Lymphoma Biomarker Consortium. J Clin Oncol 37:3359–3368. https://doi.org/10.1200/JCO.19.00743
Rossi D, Khiabanian H, Spina V, Ciardullo C, Bruscaggin A, Fama R, Rasi S, Monti S, Deambrogi C, De Paoli L, Wang J, Gattei V, Guarini A, Foa R, Rabadan R, Gaidano G (2014) Clinical impact of small TP53 mutated subclones in chronic lymphocytic leukemia. Blood 123:2139–2147. https://doi.org/10.1182/blood-2013-11-539726
Rossi D, Spina V, Gaidano G (2018) Biology and treatment of Richter syndrome. Blood 131:2761–2772. https://doi.org/10.1182/blood-2018-01-791376
Runge HFP, Lacy S, Barrans S, Beer PA, Painter D, Smith A, Roman E, Burton C, Crouch S, Tooze R, Hodson DJ (2021) Application of the LymphGen classification tool to 928 clinically and genetically-characterised cases of diffuse large B cell lymphoma (DLBCL). Br J Haematol 192:216–220. https://doi.org/10.1111/bjh.17132
Salaverria I, Martin-Guerrero I, Wagener R, Kreuz M, Kohler CW, Richter J, Pienkowska-Grela B, Adam P, Burkhardt B, Claviez A, Damm-Welk C, Drexler HG, Hummel M, Jaffe ES, Kuppers R, Lefebvre C, Lisfeld J, Loffler M, Macleod RA et al (2014) A recurrent 11q aberration pattern characterizes a subset of MYC-negative high-grade B-cell lymphomas resembling Burkitt lymphoma. Blood 123:1187–1198. https://doi.org/10.1182/blood-2013-06-507996
Scarfo I, Pellegrino E, Mereu E, Kwee I, Agnelli L, Bergaggio E, Garaffo G, Vitale N, Caputo M, Machiorlatti R, Circosta P, Abate F, Barreca A, Novero D, Mathew S, Rinaldi A, Tiacci E, Serra S, Deaglio S et al (2016) Identification of a new subclass of ALK-negative ALCL expressing aberrant levels of ERBB4 transcripts. Blood 127:221–232. https://doi.org/10.1182/blood-2014-12-614503
Scheijen B, Meijers RWJ, Rijntjes J, van der Klift MY, Mobs M, Steinhilber J, Reigl T, van den Brand M, Kotrova M, Ritter JM, Catherwood MA, Stamatopoulos K, Bruggemann M, Davi F, Darzentas N, Pott C, Fend F, Hummel M, Langerak AW et al (2019) Next-generation sequencing of immunoglobulin gene rearrangements for clonality assessment: a technical feasibility study by EuroClonality-NGS. Leukemia 33:2227–2240. https://doi.org/10.1038/s41375-019-0508-7
Schmidt J, Federmann B, Schindler N, Steinhilber J, Bonzheim I, Fend F, Quintanilla-Martinez L (2015) MYD88 L265P and CXCR4 mutations in lymphoplasmacytic lymphoma identify cases with high disease activity. Br J Haematol. https://doi.org/10.1111/bjh.13361
Schmidt J, Ramis-Zaldivar JE, Nadeu F, Gonzalez-Farre B, Navarro A, Egan C, Montes-Mojarro IA, Marafioti T, Cabecadas J, van der Walt J, Dojcinov S, Rosenwald A, Ott G, Bonzheim I, Fend F, Campo E, Jaffe ES, Salaverria I, Quintanilla-Martinez L (2017) Mutations of MAP2K1 are frequent in pediatric-type follicular lymphoma and result in ERK pathway activation. Blood 130:323–327. https://doi.org/10.1182/blood-2017-03-776278
Schmitz R, Wright GW, Huang DW, Johnson CA, Phelan JD, Wang JQ, Roulland S, Kasbekar M, Young RM, Shaffer AL, Hodson DJ, Xiao W, Yu X, Yang Y, Zhao H, Xu W, Liu X, Zhou B, Du W et al (2018) Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N Engl J Med 378:1396–1407. https://doi.org/10.1056/NEJMoa1801445
Sha C, Barrans S, Cucco F, Bentley MA, Care MA, Cummin T, Kennedy H, Thompson JS, Uddin R, Worrillow L, Chalkley R, van Hoppe M, Ahmed S, Maishman T, Caddy J, Schuh A, Mamot C, Burton C, Tooze R et al (2019) Molecular High-Grade B-Cell Lymphoma: Defining a Poor-Risk Group That Requires Different Approaches to Therapy. J Clin Oncol 37:202–212. https://doi.org/10.1200/JCO.18.01314
Sonigo G, Battistella M, Beylot-Barry M, Ingen-Housz-Oro S, Franck N, Barete S, Boulinguez S, Dereure O, Bonnet N, Socie G, Brice P, Boccara O, Bodemer C, Adamski H, D'Incan M, Ortonne N, Fraitag S, Brunet-Possenti F, Dalle S et al (2020) HAVCR2 mutations are associated with severe hemophagocytic syndrome in subcutaneous panniculitis-like T-cell lymphoma. Blood 135:1058–1061. https://doi.org/10.1182/blood.2019003811
Spina V, Khiabanian H, Messina M, Monti S, Cascione L, Bruscaggin A, Spaccarotella E, Holmes AB, Arcaini L, Lucioni M, Tabbo F, Zairis S, Diop F, Cerri M, Chiaretti S, Marasca R, Ponzoni M, Deaglio S, Ramponi A et al (2016) The genetics of nodal marginal zone lymphoma. Blood 128:1362–1373. https://doi.org/10.1182/blood-2016-02-696757
Steinhilber J, Mederake M, Bonzheim I, Serinsoz-Linke E, Muller I, Fallier-Becker P, Lemonnier F, Gaulard P, Fend F, Quintanilla-Martinez L (2019) The pathological features of angioimmunoblastic T-cell lymphomas with IDH2(R172) mutations. Mod Pathol 32:1123–1134. https://doi.org/10.1038/s41379-019-0254-4
Streubel B, Vinatzer U, Willheim M, Raderer M, Chott A (2006) Novel t(5;9)(q33;q22) fuses ITK to SYK in unspecified peripheral T-cell lymphoma. Leukemia 20:313–318. https://doi.org/10.1038/sj.leu.2404045
Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri S, Stein H, Thiele J (2017) WHO Classification of Tumours of Haematopoetic and Lymphoid Tissues. IARC
Syrykh C, Gorez P, Pericart S, Grand D, Escudie F, Cabarrou B, Oberic L, Ysebaert L, Lamant L, Laurent C, Evrard SM, Brousset P (2021) Molecular diagnosis of T-cell lymphoma: a correlative study of PCR-based T-cell clonality assessment and targeted NGS. Blood Adv 5:4590–4593. https://doi.org/10.1182/bloodadvances.2021005249
Thomas N, Dreval K, Gerhard DS, Hilton LK, Abramson JS, Ambinder RF, Barta S, Bartlett NL, Bethony J, Bhatia K, Bowen J, Bryan AC, Cesarman E, Casper C, Chadburn A, Cruz M, Dittmer DP, Dyer MA, Farinha P et al (2023) Genetic subgroups inform on pathobiology in adult and pediatric Burkitt lymphoma. Blood 141:904–916. https://doi.org/10.1182/blood.2022016534
Tiacci E, Pettirossi V, Schiavoni G, Falini B (2017) Genomics of Hairy Cell Leukemia. J Clin Oncol 35:1002–1010. https://doi.org/10.1200/JCO.2016.71.1556
Tousseyn TA, King RL, Fend F, Feldman AL, Brousset P, Jaffe ES (2023) Evolution in the definition and diagnosis of the Hodgkin lymphomas and related entities. Virchows Arch 482:207–226. https://doi.org/10.1007/s00428-022-03427-z
Treon SP, Xu L, Guerrera ML, Jimenez C, Hunter ZR, Liu X, Demos M, Gustine J, Chan G, Munshi M, Tsakmaklis N, Chen JG, Kofides A, Sklavenitis-Pistofidis R, Bustoros M, Keezer A, Meid K, Patterson CJ, Sacco A et al (2020) Genomic Landscape of Waldenstrom Macroglobulinemia and Its Impact on Treatment Strategies. J Clin Oncol 38:1198–1208. https://doi.org/10.1200/JCO.19.02314
Vallois D, Dobay MP, Morin RD, Lemonnier F, Missiaglia E, Juilland M, Iwaszkiewicz J, Fataccioli V, Bisig B, Roberti A, Grewal J, Bruneau J, Fabiani B, Martin A, Bonnet C, Michielin O, Jais JP, Figeac M, Bernard OA et al (2016) Activating mutations in genes related to TCR signaling in angioimmunoblastic and other follicular helper T-cell-derived lymphomas. Blood 128:1490–1502. https://doi.org/10.1182/blood-2016-02-698977
van Bladel DAG, Stevens WBC, van den Brand M, Kroeze LI, Groenen P, van Krieken J, Hebeda KM, Scheijen B (2022) Novel Approaches in Molecular Characterization of Classical Hodgkin Lymphoma. Cancers (Basel) 14. https://doi.org/10.3390/cancers14133222
van Bladel DAG, van den Brand M, Rijntjes J, Pamidimarri Naga S, Haacke D, Luijks J, Hebeda KM, van Krieken J, Groenen P, Scheijen B (2022) Clonality assessment and detection of clonal diversity in classic Hodgkin lymphoma by next-generation sequencing of immunoglobulin gene rearrangements. Mod Pathol 35:757–766. https://doi.org/10.1038/s41379-021-00983-8
van den Brand M, Mobs M, Otto F, Kroeze LI, Castro DG, Stamatopoulos K, Davi F, Bravetti C, Kolijn PM, Vlachonika E, Stewart JP, Pott C, Hummel M, Darzentas N, Langerak AW, Fend F, Groenen PJ (2023) EuroClonality-NGS recommendations for evaluation of B cell clonality analysis by next-generation sequencing - a structured approach with the DEPART algorithm. J Mol Diagn. https://doi.org/10.1016/j.jmoldx.2023.06.011
van Dongen JJ, Langerak AW, Bruggemann M, Evans PA, Hummel M, Lavender FL, Delabesse E, Davi F, Schuuring E, Garcia-Sanz R, van Krieken JH, Droese J, Gonzalez D, Bastard C, White HE, Spaargaren M, Gonzalez M, Parreira A, Smith JL et al (2003) Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17:2257–2317. https://doi.org/10.1038/sj.leu.2403202
Vela V, Juskevicius D, Dirnhofer S, Menter T, Tzankov A (2022) Mutational landscape of marginal zone B-cell lymphomas of various origin: organotypic alterations and diagnostic potential for assignment of organ origin. Virchows Arch 480:403–413. https://doi.org/10.1007/s00428-021-03186-3
Wagener R, Lopez C, Kleinheinz K, Bausinger J, Aukema SM, Nagel I, Toprak UH, Seufert J, Altmuller J, Thiele H, Schneider C, Kolarova J, Park J, Hubschmann D, Murga Penas EM, Drexler HG, Attarbaschi A, Hovland R, Kjeldsen E et al (2018) IG-MYC (+) neoplasms with precursor B-cell phenotype are molecularly distinct from Burkitt lymphomas. Blood 132:2280–2285. https://doi.org/10.1182/blood-2018-03-842088
Weniger MA, Kuppers R (2021) Molecular biology of Hodgkin lymphoma. Leukemia 35:968–981. https://doi.org/10.1038/s41375-021-01204-6
Wright G, Tan B, Rosenwald A, Hurt EH, Wiestner A, Staudt LM (2003) A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc Natl Acad Sci U S A 100:9991–9996. https://doi.org/10.1073/pnas.1732008100
Wright GW, Huang DW, Phelan JD, Coulibaly ZA, Roulland S, Young RM, Wang JQ, Schmitz R, Morin RD, Tang J, Jiang A, Bagaev A, Plotnikova O, Kotlov N, Johnson CA, Wilson WH, Scott DW, Staudt LM (2020) A probabilistic classification tool for genetic subtypes of diffuse large b cell lymphoma with therapeutic implications. Cancer Cell 37(551-568):e514. https://doi.org/10.1016/j.ccell.2020.03.015
Young RM, Staudt LM (2013) Targeting pathological B cell receptor signalling in lymphoid malignancies. Nat Rev Drug Discov 12:229–243. https://doi.org/10.1038/nrd3937
Zamo A, Gerhard-Hartmann E, Ott G, Anagnostopoulos I, Scott DW, Rosenwald A, Rauert-Wunderlich H (2022) Routine application of the Lymph2Cx assay for the subclassification of aggressive B-cell lymphoma: report of a prospective real-world series. Virchows Arch 481:935–943. https://doi.org/10.1007/s00428-022-03420-6
Funding
Open Access funding enabled and organized by Projekt DEAL. Does not apply
Author information
Authors and Affiliations
Contributions
FF, MvdB, PG, LQM, and FF have all contributed to the writing of the manuscript and preparation of illustrations and have reviewed and corrected the final version of the manuscript. FF was responsible for the overall assembly of the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Compliance with ethical standards
The authors confirm that the work complies with ethical standards.
Conflict of interest
The authors state that there is no conflict of interest relevant to this manuscript.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fend, F., van den Brand, M., Groenen, P.J. et al. Diagnostic and prognostic molecular pathology of lymphoid malignancies. Virchows Arch 484, 195–214 (2024). https://doi.org/10.1007/s00428-023-03644-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-023-03644-0