Abstract
Main conclusion
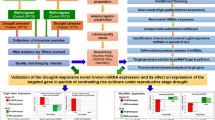
Our work provides a survey of mature miRNAs, their target genes and primary precursors identified by in-silico approach in leaf transcriptomes of five selected Hypericum species.
Abstract
MiRNAs are small non-coding RNA molecules found in animals, terrestrial plants, several algae and molds. As their role lies in the post-transcriptional gene silencing, these tiny molecules regulate many biological processes. Phyto-miRNAs are considered the important regulators of secondary metabolism in medicinal plants. The genus Hypericum comprises many producers of bioactive compounds, mainly unique naphtodianthrones with a great therapeutic potential. The main goal of our work was to identify genetically conserved miRNAs, characterize their primary precursors and target sequences in the leaf transcriptomes of five Hypericum species using in-silico approach. We found 20 sequences of potential Hypericum pri-miRNAs, and predicted and computationally validated their secondary structures. The mature miRNAs were identified by target genes screening analysis. Whereas predicted miRNA profiles differed in less genetically conserved families, the highly conserved miRNAs were found in almost all studied species. Moreover, we detected several novel highly likely miRNA–mRNA interactions, such as mir1171 with predicted regulatory role in the biosynthesis of melatonin in plants. Our work contributes to the knowledge of Hypericum miRNAome and miRNA–mRNA interactions.



Similar content being viewed by others
References
Axtell MJ, Meyers BC (2018) Revisiting criteria for plant microRNA Annotation in the era of big data. Plant Cell 30:272. https://doi.org/10.1105/tpc.17.00851
Bartel DP (2018) Metazoan microRNAs. Cell 173:20–51. https://doi.org/10.1016/j.cell.2018.03.006
Borges F, Martienssen RA (2015) The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol 16:727–741. https://doi.org/10.1038/nrm4085
Bulgakov VP, Avramenko TV (2015) New opportunities for the regulation of secondary metabolism in plants: focus on microRNAs. Biotechnol Lett 37:1719–1727. https://doi.org/10.1007/s10529-015-1863-8
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Cavalieri D, Rizzetto L, Tocci N et al (2016) Plant microRNAs as novel immunomodulatory agents. Sci Rep 6:25761
Clokie SJH, Lau P, Kim HH et al (2012) MicroRNAs in the pineal gland: miR-483 regulates melatonin synthesis by targeting arylalkylamine N-acetyltransferase. J Biol Chem 287:25312–25324. https://doi.org/10.1074/jbc.M112.356733
Dai X, Zhuang Z, Zhao PX (2018) psRNATarget: a plant small RNA target analysis server (2017 release). Nucleic Acids Res 46:W49–W54. https://doi.org/10.1093/nar/gky316
Djami-Tchatchou AT, Sanan-Mishra N, Ntushelo K, Dubery IA (2017) Functional roles of microRNAs in agronomically important plants—potential as targets for crop improvement and protection. Front Plant Sci 8:378. https://doi.org/10.3389/fpls.2017.00378
Ellis J, Dodds P, Pryor T (2000) Structure, function and evolution of plant disease resistance genes. Curr Opin Plant Biol 3:278–284
Ergun S (2019) Cross-kingdom gene regulation via miRNAs of Hypericum perforatum (St. John’s wort) flower dietetically absorbed: an in silico approach to define potential biomarkers for prostate cancer. Comput Biol Chem 80:16–22. https://doi.org/10.1016/j.compbiolchem.2019.02.010
Fu L, Niu B, Zhu Z et al (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Galla G, Volpato M, Sharbel TF, Barcaccia G (2013) Computational identification of conserved microRNAs and their putative targets in the Hypericum perforatum L. flower transcriptome. Plant Reprod 26:209–229. https://doi.org/10.1007/s00497-013-0227-6
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50:151–158. https://doi.org/10.1016/0014-4827(68)90403-5
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652. https://doi.org/10.1038/nbt.1883
Gupta OP, Karkute SG, Banerjee S et al (2017) Contemporary understanding of miRNA-based regulation of secondary metabolites biosynthesis in plants. Front Plant Sci 8:374. https://doi.org/10.3389/fpls.2017.00374
Jeyaraj A, Liu S, Zhang X et al (2017) Genome-wide identification of microRNAs responsive to Ectropis oblique feeding in tea plant (Camellia sinensis L.). Sci Rep 7:13634. https://doi.org/10.1038/s41598-017-13692-7
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAS and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218
Kalvari I, Argasinska J, Quinones-Olvera N et al (2018) Rfam 13.0: shifting to a genome-centric resource for non-coding RNA families. Nucleic Acids Res 46:D335–D342. https://doi.org/10.1093/nar/gkx1038
Kang K, Kong K, Park S et al (2011) Molecular cloning of a plant N-acetylserotonin methyltransferase and its expression characteristics in rice. J Pineal Res 50:304–309. https://doi.org/10.1111/j.1600-079X.2010.00841.x
Kang Y-J, Yang D-C, Kong L et al (2017) CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res 45:W12–W16. https://doi.org/10.1093/nar/gkx428
Kim H, Kim SW, Seok KH et al (2018) Hypericin-assisted photodynamic therapy against anaplastic thyroid cancer. Photodiagnosis Photodyn Ther 24:15–21. https://doi.org/10.1016/j.pdpdt.2018.08.008
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39:D152–157. https://doi.org/10.1093/nar/gkq1027
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc Natl Acad Sci USA 101:12753–12758. https://doi.org/10.1073/pnas.0403115101
Lelandais-Briere C, Sorin C, Declerck M et al (2010) Small RNA diversity in plants and its impact in development. Curr Genomics 11:14–23. https://doi.org/10.2174/138920210790217918
Li B, Fillmore N, Bai Y et al (2014) Evaluation of de novo transcriptome assemblies from RNA-Seq data. Genome Biol 15:553. https://doi.org/10.1186/s13059-014-0553-5
Luo R, Liu B, Xie Y et al (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18. https://doi.org/10.1186/2047-217X-1-18
Margis R, Fusaro AF, Smith NA et al (2006) The evolution and diversification of Dicers in plants. FEBS Lett 580:2442–2450. https://doi.org/10.1016/j.febslet.2006.03.072
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Murch SJ, Simmons CB, Saxena PK (1997) Melatonin in feverfew and other medicinal plants. Lancet 350:1598–1599. https://doi.org/10.1016/S0140-6736(05)64014-7
Robson NKB (2016) And then came molecular phylogenetics—reactions to a monographic study of Hypericum (Hypericaceae). Phytotaxa 255(3):181 198. https://doi.org/10.11646/phytotaxa.255.3.1
Sabzehzari M, Naghavi MR (2019) Phyto-miRNA: a molecule with beneficial abilities for plant biotechnology. Gene 683:28–34. https://doi.org/10.1016/j.gene.2018.09.054
Samad AFA, Sajad M, Nazaruddin N et al (2017) MicroRNA and transcription factor: key players in plant regulatory network. Front Plant Sci 8:565. https://doi.org/10.3389/fpls.2017.00565
Song X, Li Y, Cao X, Qi Y (2019) MicroRNAs and their regulatory roles in plant–environment interactions. Annu Rev Plant Biol 70:489–525. https://doi.org/10.1146/annurev-arplant-050718-100334
Soták M, Czeranková O, Klein D et al (2016) Comparative transcriptome reconstruction of four Hypericum species focused on hypericin biosynthesis. Front Plant Sci 7:1039. https://doi.org/10.3389/fpls.2016.01039
Tan D-X, Zheng X, Kong J et al (2014) Fundamental issues related to the origin of melatonin and melatonin isomers during evolution: relation to their biological functions. Int J Mol Sci 15:15858–15890. https://doi.org/10.3390/ijms150915858
Wang J, Mei J, Ren G (2019) Plant microRNAs: biogenesis, homeostasis, and degradation. Front Plant Sci 10:360. https://doi.org/10.3389/fpls.2019.00360
Xu L, Zhang X, Cheng W et al (2019) Hypericin-photodynamic therapy inhibits the growth of adult T-cell leukemia cells through induction of apoptosis and suppression of viral transcription. Retrovirology 16:5. https://doi.org/10.1186/s12977-019-0467-0
Yang JH, Han SJ, Yoon EK, Lee WS (2006) Evidence of an auxin signal pathway, microRNA167-ARF8-GH3, and its response to exogenous auxin in cultured rice cells. Nucleic Acids Res 34:1892–1899. https://doi.org/10.1093/nar/gkl118
Zhang J-W, Long Y, Xue M et al (2017) Identification of microRNAs in response to drought in common wild rice (Oryza rufipogon Griff.) shoots and roots. PLoS ONE 12:e0170330. https://doi.org/10.1371/journal.pone.0170330
Zhang K, Gao S, Guo J et al (2018) Hypericin-photodynamic therapy inhibits proliferation and induces apoptosis in human rheumatoid arthritis fibroblast-like synoviocytes cell line MH7A. Iran J Basic Med Sci 21:130–137. https://doi.org/10.22038/IJBMS.2018.23871.5991
Zhao D, Yu Y, Shen Y et al (2019) Melatonin synthesis and function: Evolutionary history in animals and plants. Front Endocrinol (Lausanne) 10:249. https://doi.org/10.3389/fendo.2019.00249
Zinati Z, Shamloo-Dashtpagerdi R, Behpouri A (2016) In silico identification of miRNAs and their target genes and analysis of gene co-expression network in saffron (Crocus sativus L.) stigma. Mol Biol Res Commun 5:233–246
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415. https://doi.org/10.1093/nar/gkg595
Acknowledgements
The work was supported by the Slovak Research and Development Agency under Grant number APVV-18-0125 and the Scientific Grant Agency of Slovak Republic under Grant number VEGA 1/0013/19.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Petijová, L., Jurčacková, Z. & Čellárová, E. Computational screening of miRNAs and their targets in leaves of Hypericum spp. by transcriptome-mining: a pilot study. Planta 251, 49 (2020). https://doi.org/10.1007/s00425-020-03342-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03342-0