Abstract
Background
Amyotrophic lateral sclerosis (ALS) is a late-onset neurodegenerative disorder. Mitochondrial dysfunction is involved in the complex pathophysiology of ALS; however, the role of mitochondrial DNA (mtDNA) variants in ALS is poorly understood. We aimed to elucidate the role of mtDNA variants in the pathogenesis of ALS.
Methods
The mitochondrial haplogroups of 585 ALS patients and 371 healthy controls were determined; 38 ALS patients and 42 controls underwent long-range polymerase chain reaction combined with next-generation sequencing technology to analyze whole mitochondrial genome variants.
Results
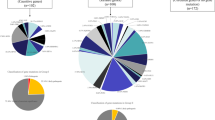
A higher percentage of variants accumulated in ALS patients than in controls. Analysis of coding region variations that were further stratified by mtDNA genes revealed that nonsynonymous variants were more vulnerable in ALS patients than in controls, particularly in the ND4L, ND5, and ATP8 genes. Moreover, pathogenic nonsynonymous variants tended to over-represent in ALS patients. Unsurprisingly, nonsynonymous variants were not related to the phenotype. Haplogroup analysis did not found evidence of association between haplogroups with the risk of ALS, however, patients belonging to haplogroup Y and M7c were prone to develop later onset of ALS.
Conclusions
This is the first study to profile mtDNA variants in ALS patients from mainland China. Our results suggest that an increase in the number of nonsynonymous variants is linked to the pathogenesis of ALS. Moreover, haplogroup Y and M7c may modulate the clinical expression of ALS. Our findings provide independent, albeit limited, evidence for the role of mtDNA in the pathogenesis of ALS.



Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
van Es MA, Hardiman O, Chio A, Al-Chalabi A, Pasterkamp RJ, Veldink JH, van den Berg LH (2017) Amyotrophic lateral sclerosis. Lancet 390:2084–2098. https://doi.org/10.1016/S0140-6736(17)31287-4
Ghasemi M, Brown RH Jr (2018) Genetics of amyotrophic lateral sclerosis. Cold Spring Harb Perspect Med. https://doi.org/10.1101/cshperspect.a024125
Chia R, Chiò A, Traynor BJ (2018) Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol 17:94–102. https://doi.org/10.1016/s1474-4422(17)30401-5
Cha MY, Kim DK, Mook-Jung I (2015) The role of mitochondrial DNA mutation on neurodegenerative diseases. Exp Mol Med 47:e150. https://doi.org/10.1038/emm.2014.122
Ryan TE, Erickson ML, Verma A, Chavez J, Rivner MH, McCully KK (2014) Skeletal muscle oxidative capacity in amyotrophic lateral sclerosis. Muscle Nerve 50:767–774. https://doi.org/10.1002/mus.24223
Ladd AC, Brohawn DG, Thomas RR, Keeney PM, Berr SS, Khan SM, Portell FR, Shakenov MZ, Antkowiak PF, Kundu B, Tustison N, Bennett JP (2017) RNA-seq analyses reveal that cervical spinal cords and anterior motor neurons from amyotrophic lateral sclerosis subjects show reduced expression of mitochondrial DNA-encoded respiratory genes, and rhTFAM may correct this respiratory deficiency. Brain Res 1667:74–83. https://doi.org/10.1016/j.brainres.2017.05.010
Stoccoro A, Mosca L, Carnicelli V, Cavallari U, Lunetta C, Marocchi A, Migliore L, Coppedè F (2018) Mitochondrial DNA copy number and D-loop region methylation in carriers of amyotrophic lateral sclerosis gene mutations. Epigenomics 10:1431–1443. https://doi.org/10.2217/epi-2018-0072
Tuppen HA, Blakely EL, Turnbull DM, Taylor RW (2010) Mitochondrial DNA mutations and human disease. Biochim Biophys Acta 1797:113–128. https://doi.org/10.1016/j.bbabio.2009.09.005
Zhang W, Tang J, Zhang AM, Peng MS, Xie HB, Tan L, Xu L, Zhang YP, Chen X, Yao YG (2014) A matrilineal genetic legacy from the last glacial maximum confers susceptibility to schizophrenia in Han Chinese. J Genet Genomics 41:397–407. https://doi.org/10.1016/j.jgg.2014.05.004
Lynch D, Wanglund C, Spathis R, Chan CW, Reiff DM, Lum JK, Garruto RM (2008) The contribution of mitochondrial dysfunction to a gene-environment model of Guamanian ALS and PD. Mitochondrion 8:109–116. https://doi.org/10.1016/j.mito.2007.09.002
Reiff DM, Spathis R, Chan CW, Vilar MG, Sankaranarayanan K, Lynch D, Ehrlich E, Kerath S, Chowdhury R, Robinowitz L, Koji Lum J, Garruto RM (2011) Inherited and somatic mitochondrial DNA mutations in Guam amyotrophic lateral sclerosis and parkinsonism-dementia. Neurol Sci 32:883–892. https://doi.org/10.1007/s10072-011-0735-9
Keeney PM, Bennett JP Jr (2010) ALS spinal neurons show varied and reduced mtDNA gene copy numbers and increased mtDNA gene deletions. Mol Neurodegener 5:21. https://doi.org/10.1186/1750-1326-5-21
Artuso L, Zoccolella S, Favia P, Amati A, Capozzo R, Logroscino G, Serlenga L, Simone I, Gasparre G, Petruzzella V (2013) Mitochondrial genome aberrations in skeletal muscle of patients with motor neuron disease. Amyotroph Lateral Scler Frontotemporal Degener 14:261–266. https://doi.org/10.3109/21678421.2012.735239
Mancuso M, Conforti FL, Rocchi A, Tessitore A, Muglia M, Tedeschi G, Panza D, Monsurrò M, Sola P, Mandrioli J, Choub A, DelCorona A, Manca ML, Mazzei R, Sprovieri T, Filosto M, Salviati A, Valentino P, Bono F, Caracciolo M, Simone IL, La Bella V, Majorana G, Siciliano G, Murri L, Quattrone A (2004) Could mitochondrial haplogroups play a role in sporadic amyotrophic lateral sclerosis? Neurosci Lett 371:158–162. https://doi.org/10.1016/j.neulet.2004.08.060
Ingram CJ, Weale ME, Plaster CA, Morrison KE, Goodall EF, Pall HS, Beck M, Jablonka S, Sendtner M, Fisher EM, Bradman N, Kasperavičiūtė D (2012) Analysis of European case-control studies suggests that common inherited variation in mitochondrial DNA is not involved in susceptibility to amyotrophic lateral sclerosis. Amyotroph Lateral Scler 13:341–346. https://doi.org/10.3109/17482968.2012.654394
Wei W, Keogh MJ, Wilson I, Coxhead J, Ryan S, Rollinson S, Griffin H, Kurzawa-Akanbi M, Santibanez-Koref M, Talbot K, Turner MR, McKenzie CA, Troakes C, Attems J, Smith C, Al Sarraj S, Morris CM, Ansorge O, Pickering-Brown S, Ironside JW, Chinnery PF (2017) Mitochondrial DNA point mutations and relative copy number in 1363 disease and control human brains. Acta Neuropathol Commun 5:13. https://doi.org/10.1186/s40478-016-0404-6
Bris C, Goudenege D, Desquiret-Dumas V, Charif M, Colin E, Bonneau D, Amati-Bonneau P, Lenaers G, Reynier P, Procaccio V (2018) Bioinformatics tools and databases to assess the pathogenicity of mitochondrial DNA variants in the field of next generation sequencing. Front Genet 9:632. https://doi.org/10.3389/fgene.2018.00632
Ludolph A, Drory V, Hardiman O, Nakano I, Ravits J, Robberecht W, Shefner J (2015) A revision of the El escorial criteria— 2015. Amyotroph Lateral Scler Frontotemporal Degener 16:291–292. https://doi.org/10.3109/21678421.2015.1049183
Zhang W, Cui H, Wong LJ (2012) Comprehensive one-step molecular analyses of mitochondrial genome by massively parallel sequencing. Clin Chem 58:1322–1331. https://doi.org/10.1373/clinchem.2011.181438
Yuan H, Yang H, Peng L, Peng Y, Chen Z, Wan L, Wang C, Shi Y, Zhang VW, Tang B, Qiu R, Jiang H (2020) Profiling of mitochondrial genomes in SCA3/MJD patients from mainland China. Gene 738:144487. https://doi.org/10.1016/j.gene.2020.144487
Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, Howell N (1999) Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet 23:147. https://doi.org/10.1038/13779
Fan L, Yao YG (2013) An update to MitoTool: using a new scoring system for faster mtDNA haplogroup determination. Mitochondrion 13:360–363. https://doi.org/10.1016/j.mito.2013.04.011
Preste R, Vitale O, Clima R, Gasparre G, Attimonelli M (2019) HmtVar: a new resource for human mitochondrial variations and pathogenicity data. Nucleic Acids Res 47:D1202–D1210. https://doi.org/10.1093/nar/gky1024
Lott MT, Leipzig JN, Derbeneva O, Xie HM, Chalkia D, Sarmady M, Procaccio V, Wallace DC (2013) mtDNA variation and analysis using mitomap and mitomaster. Curr Protoc Bioinf 44(1):1–23. https://doi.org/10.1002/0471250953.bi0123s44
Tang S, Wang J, Zhang VW, Li FY, Landsverk M, Cui H, Truong CK, Wang G, Chen LC, Graham B, Scaglia F, Schmitt ES, Craigen WJ, Wong LJ (2013) Transition to next generation analysis of the whole mitochondrial genome: a summary of molecular defects. Hum Mutat 34:882–893. https://doi.org/10.1002/humu.22307
Kennedy SR, Salk JJ, Schmitt MW, Loeb LA (2013) Ultra-sensitive sequencing reveals an age-related increase in somatic mitochondrial mutations that are inconsistent with oxidative damage. PLoS Genet 9:e1003794. https://doi.org/10.1371/journal.pgen.1003794
Hoekstra JG, Hipp MJ, Montine TJ, Kennedy SR (2016) Mitochondrial DNA mutations increase in early stage Alzheimer disease and are inconsistent with oxidative damage. Ann Neurol 80:301–306. https://doi.org/10.1002/ana.24709
Coxhead J, Kurzawa-Akanbi M, Hussain R, Pyle A, Chinnery P, Hudson G (2016) Somatic mtDNA variation is an important component of Parkinson’s disease. Neurobiol Aging 38:217.e211-217.e216. https://doi.org/10.1016/j.neurobiolaging.2015.10.036
Siddiqui A, Rivera-Sánchez S, Castro Mdel R, Acevedo-Torres K, Rane A, Torres-Ramos CA, Nicholls DG, Andersen JK, Ayala-Torres S (2012) Mitochondrial DNA damage is associated with reduced mitochondrial bioenergetics in Huntington’s disease. Free Radic Biol Med 53:1478–1488. https://doi.org/10.1016/j.freeradbiomed.2012.06.008
Murata T, Ohtsuka C, Terayama Y (2008) Increased mitochondrial oxidative damage and oxidative DNA damage contributes to the neurodegenerative process in sporadic amyotrophic lateral sclerosis. Free Radic Res 42:221–225. https://doi.org/10.1080/10715760701877262
Bi R, Zhang W, Yu D, Li X, Wang HZ, Hu QX, Zhang C, Lu W, Ni J, Fang Y, Li T, Yao YG (2015) Mitochondrial DNA haplogroup B5 confers genetic susceptibility to Alzheimer’s disease in Han Chinese. Neurobiol Aging 36:1604.e1607-1616. https://doi.org/10.1016/j.neurobiolaging.2014.10.009
Chen YF, Chen WJ, Lin XZ, Zhang QJ, Cai JP, Liou CW, Wang N (2015) Mitochondrial DNA haplogroups and the risk of sporadic Parkinson’s disease in Han Chinese. Chin Med J (Engl) 128:1748–1754. https://doi.org/10.4103/0366-6999.159348
Chinnery PF, Mowbray C, Elliot H, Elson JL, Nixon H, Hartley J, Shaw PJ (2007) Mitochondrial DNA haplogroups and amyotrophic lateral sclerosis. Neurogenetics 8:65–67. https://doi.org/10.1007/s10048-006-0066-9
Acknowledgements
We are grateful to the participating patients for their involvement.
Funding
This work was supported by the National Key Research and Development Program of China (#2018YFC1312003); the Program of National Natural Science Foundation of China (#81671120, 81300981); and Natural Science Fund for Distinguished Young Scholars of Hunan province, China (#2020JJ2057); the Project Program of National Clinical Research Center for Geriatric Disorders (Xiangya Hospital) (#2020LNJJ13); and the Degree & Postgraduate Education Reform Project of Central South University (#2020JGB136).
Author information
Authors and Affiliations
Contributions
Formal analysis: JN; Investigation: JN, ZL, YY; Methodology: JN; Writing—original draft: JN; Data curation: JN, ZL, YY, WLi, YH, PL, XH, XZ, XT, ML, SZ; Writing—review & editing: XH, JD, JW; Supervision: JT, HJ, LS, BT, JW; Conceptualization: JW; Funding acquisition: JW.
Corresponding author
Ethics declarations
Conflicts of interest
The authors report no actual or potential conflicts of interest.
Ethics approval
The study was approved by the Ethics Committee of Xiangya Hospital, Central South University.
Consent to participate
Written informed consent was obtained from all subjects.
Consent for publication
Written informed consent for publication was obtained from all participants.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ni, J., Liu, Z., Yuan, Y. et al. Mitochondrial genome variations are associated with amyotrophic lateral sclerosis in patients from mainland China. J Neurol 269, 805–814 (2022). https://doi.org/10.1007/s00415-021-10659-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-021-10659-7