Abstract
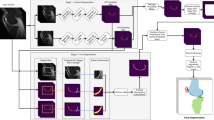
Age assessment is used to estimate the chronological age of an individual who lacks legal documentation. Recent studies indicate that the ossification degree of the growth plates in the knee joint correlates with chronological age of adolescents and young adults. To verify this hypothesis, a high number of datasets need to be analysed. An approach which enables an automated detection and analysis of the bone structures may be necessary to handle large datasets. The purpose of this study was to develop a fully automatic 2D knee segmentation based on 3D MR images using convolutional neural networks. A total of 76 datasets were available and divided into a training set (74%), a validation set (13%) and a test set (13%). Multiple preprocessing steps were applied to correct image intensity values and to reduce the image size. Image augmentation was employed to virtually increase the dataset size for training. The proposed architecture for the segmentation task resembles the encoder-decoder model type used for the U-Net. The trained network achieved a dice similarity coefficient score of 98% compared to the manual segmentations and an intersection over union of 96%. The precision and recall of the model were balanced, and the error was only 1.2%. No overfitting was observed during training. As a proof of concept, the predicted segmentations were used for the age estimation of 145 subjects. Initial results show the potential of this approach attaining a mean absolute error of 0.48 ± 0.32 years for a test set of 14 subjects. The proposed automated segmentation can contribute to faster, reproducible and potentially more reliable age estimation in the future.












Similar content being viewed by others
References
Britting-Reimer E (2015) Altersbestimmung in Deutschland und im Europäischen Vergleich
Geserick G, Schmeling A (2011) Qualitätssicherung der Forensischen Altersdiagnostik bei Lebenden Personen. Rechtsmedizin 21(1):22–25
Kubilay S (2016) Ablauf des deutschen Asylverfahrens, tech. rep. Bundesamt für Migration und Flüchtlinge (BAMF)
European Asylum Support Office (2013) Age assessment practice in Europe
World Health Organization (2016) Ionizing radiation, health effects and protective measures
Dedouit F, Auriol J, Rousseau H, Rougé D, Crubézy E, Telmon N (2012) Age assessment by magnetic resonance imaging of the knee: a preliminary study. Forensic Sci Int 217(1-3):232
Jopp E, Schröder I, Maas R, Adam G, Püschel K (2010) Proximale Tibiaepiphyse im Magnetresonanztomogramm: Neue Möglichkeit zur Altersbestimmung bei Lebenden?. Rechtsmedizin 20:464–468
Jopp E (2013) Die Abschlussphase des menschlichen Wachstums Longitudinale Ganzkörper- und Unterschenkelmessungen (Knemometrie) an jungen Erwachsenen zur Bestimmung des biologischen Alters und für forensische Zwecke. PhD thesis
Krämer JA, Schmidt S, Jürgens KU, Lentschig M, Schmeling A, Vieth V (2014) Forensic age estimation in living individuals using 3.0T MRI of the distal femur. Int J Legal Med 128(3):509–514
Laor T, Chun GFH, Dardzinski BJ, Bean JA, Witte DP (2002) Posterior distal femoral and proximal tibial metaphyseal stripes at mr imaging in children and young adults. Radiology 224:669–674
Ekizoglu O, Hocaoglu E, Inci E, Can IO, Aksoy S, Kazimoglu C (2016) Forensic age estimation via 3-T magnetic resonance imaging of ossification of the proximal tibial and distal femoral epiphyses: Use of a T2-weighted fast spin-echo technique, vol 260
Fan F, Zhang K, Peng Z, Hui Cui J-h, Hu N, Hua Deng Z-h (2016) Forensic age estimation of living persons from the knee: comparison of MRI with radiographs. Forensic Sci Int 268:145–150
Ottow C, Schulz R, Pfeiffer H, Heindel W, Schmeling A, Vieth V (2017) Forensic age estimation by magnetic resonance imaging of the knee: the definite relevance in bony fusion of the distal femoral- and the proximal tibial epiphyses using closest-to-bone T1 TSE sequence, European Radiology, pp 1–8
Jiang J, Trundle P, Ren J (2010) Medical image analysis with artificial neural networks. Comput Med Imaging Graph 34:617–631
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, van der Laak JAWM, van Ginneken B, Sánchez CI (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88
Setiono R, Liu H (1997) Neural-network feature selector. IEEE Trans Neural Netw 8:654–662
Dodin P, Martel-Pelletier J, Pelletier J-PP, Abram F (2011) A fully automated human knee 3D MRI bone segmentation using the ray casting technique. Med Biol Eng Comput 49(12):1413–1424
Dam EB, Lillholm M, Marques J, Nielsen M (2015) Automatic segmentation of high- and low-field knee MRIs using knee image quantification with data from the osteoarthritis initiative, vol 2
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation, vol 9351
Stern D, Ebner T, Bischof H, Grassegger S, Ehammer T, Urschler M (2014) Fully automatic bone age estimation from left hand MR images. Medical Image Computing and Computer-Assisted Intervention - MICCAI 2014 117(2):220–227
Stern D, Urschler M (2016) From individual hand bone age estimates to fully automated age estimation via learning-based information fusion
Stern D, Kainz P, Payer C, Urschler M (2017) Multi-factorial age estimation from skeletal and dental MRI volumes. In: Wang Q, Shi Y, Suk H-I, Suzuki K (eds) Machine learning in medical imaging. Springer International Publishing, Cham, pp 61–69
Auf der Mauer M, Säring D, Stanczus B, Herrmann J, Groth M, Jopp-van Well E (2018) A 2-year follow-up MRI study for the evaluation of an age estimation method based on knee bone development. International Journal of Legal Medicine
Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, Gee JC (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29:1310–1320
NVIDIA GPU vs CPU? What is GPU Computing?
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet Classification with Deep Convolutional Neural Networks. Advances in Neural Information Processing Systems 25 (NIPS 2012)
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: The IEEE conference on computer vision and pattern recognition (CVPR), pp 770–778
Iandola FN, Han S, Moskewicz MW, Ashraf K, Dally WJ, Keutzer K (2016) SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and < 0.5MB model size. pp 1–13
Chen L-C, Papandreou G, Schroff F, Adam H (2017) Rethinking atrous convolution for semantic image segmentation
Zhao H, Shi J, Qi X, Wang X, Jia J (2016) Pyramid scene parsing network
Lin G, Milan A, Shen C, Reid I (2016) RefineNet: multi-path refinement networks for high-resolution semantic segmentation
Chen L-C, Papandreou G, Kokkinos I, Murphy K, Yuille AL (2016) DeepLab: Semantic Image Segmentation with Deep Convolutional Nets, Atrous Convolution, and Fully Connected CRFs. pp 1–14
Nekrasov V, Ju J, Choi J (2016) Global Deconvolutional Networks for Semantic segmentation
Milletari F, Navab N, Ahmadi S-A (2016) V-Net: fully convolutional neural networks for volumetric medical image segmentation. In: 2016 Fourth International conference on 3D vision (3DV), IEEE, pp 565–571
Çiçek Ö, Abdulkadir A, Lienkamp SS, Brox T, Ronneberger O (2016) 3D U-Net: learning dense volumetric segmentation from sparse annotation. In: Medical image computing and computer-assisted intervention – MICCAI 2016, vol 9901, pp 424–432
Badrinarayanan V, Kendall A, Cipolla R (2015) SegNet: A deep convolutional encoder-decoder architecture for image segmentation. pp 1–14
Nair V, Hinton GE (2010) Rectified linear units improve restricted boltzmann machines. In: Proceedings of the 27th international conference on machine learning (ICML-10), pp 807–814
Clevert D-A, Unterthiner T, Hochreiter S (2015) Fast and accurate deep network learning by exponential linear units (ELUs). pp 1–14
Srivastava N, Hinton G, Krizhevsky A, Sutskever I, Salakhutdinov R (2014) Dropout: a simple way to prevent neural networks from overfitting. J Mach Learn Res 15:1929–1958
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. ICLR 2015, pp 1–14
Ioffe S, Szegedy C (2015) Batch normalization: accelerating deep network training by reducing internal covariate shift. In: Proceedings of machine learning research, vol 37, pp 448–456
Kingma DP, Ba J (2015) Adam: a method for stochastic optimization. In: 3rd international conference for learning representations, pp 1–15
Chollet F (2017) Deep Learning With python, vol 1. Manning Publications
Prasoon A, Petersen K, Igel C, Lauze F, Dam E, Nielsen M (2013) Deep feature learning for knee cartilage segmentation using a triplanar convolutional neural network. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2013 8150:246–253
Säring D, Auf der Mauer M, Jopp E (2014) Klassifikation des Verschlussgrades der Epiphyse der proximalen Tibia zur Altersbestimmung. In: Informatik aktuell. Springer, Berlin, pp 60–65
Saint-Martin P, Rérolle C, Dedouit F, Rousseau H, Rougé D, Telmon N (2014) Evaluation of an automatic method for forensic age estimation by magnetic resonance imaging of the distal tibial epiphysis—a preliminary study focusing on the 18-year threshold. Int J Legal Med 128:675–683
Jopp E (2007) Methoden zur Alters- und Geschlechtsbestimmung auf dem Pruefstand - eine rechtsmedizinische empirische Studie. Kovac
Krämer JA, Schmidt S, Jürgens KU, Lentschig M, Schmeling A, Vieth V (2014) The use of magnetic resonance imaging to examine ossification of the proximal tibial epiphysis for forensic age estimation in living individuals. Forensic Sci Med Pathol 10(3):306–313
Saint-Martin P, Rérolle C, Pucheux J, Dedouit F, Telmon N (2015) Contribution of distal femur MRI to the determination of the 18-year limit in forensic age estimation. Int J Legal Med 129(3):619–620
Pelt DM, Sethian JA (2018) A mixed-scale dense convolutional neural network for image analysis. Proc Natl Acad Sci 115(2):254–259
Funding
This project is funded by the German Research Foundation (DFG), Project (SA 2530/6-1) and (JO 1198/2-1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
An ethical approval for this study was granted by the Ethics Committee of the Medical Association Hamburg (PV4527). The director of this study had full control of the data and the material submitted for publication.
Conflict of interests
The authors declare that they have no conflict of interest.
Consent for Publication
Written informed consent was obtained from all subjects in this study.
Rights and permissions
About this article
Cite this article
Pröve, PL., Jopp-van Well, E., Stanczus, B. et al. Automated segmentation of the knee for age assessment in 3D MR images using convolutional neural networks. Int J Legal Med 133, 1191–1205 (2019). https://doi.org/10.1007/s00414-018-1953-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-018-1953-y