Abstract
Background
There has been growing interest in using artificial intelligence/deep learning (DL) to help diagnose prevalent diseases earlier. In this study we sought to survey the landscape of externally validated DL-based computer-aided diagnostic (CADx) models, and assess their diagnostic performance for predicting the risk of malignancy in computed tomography (CT)-detected pulmonary nodules.
Methods
An electronic search was performed in four databases (from inception to 10 August 2023). Studies were eligible if they were peer-reviewed experimental or observational articles comparing the diagnostic performance of externally validated DL-based CADx models with models widely used in clinical practice to predict the risk of malignancy. A bivariate random-effect approach for the meta-analysis on the included studies was used.
Results
Seventeen studies were included, comprising 8553 participants and 9884 nodules. Pooled analyses showed DL-based CADx models were 11.6% more sensitive than physician judgement alone, and 14.5% more than clinical risk models alone. They had a similar pooled specificity to physician judgement alone [0.77 (95% CI 0.68–0.84) v 0.81 (95% CI 0.71–0.88)], and were 7.4% more specific than clinical risk models alone. They had superior pooled areas under the receiver operating curve (AUC), with relative pooled AUCs of 1.03 (95% CI 1.00–1.07) and 1.10 (95% CI 1.07–1.13) versus physician judgement and clinical risk models alone, respectively.
Conclusion
DL-based models are already used in clinical practice in certain settings for nodule management. Our results show their diagnostic performance potentially justifies wider, more routine deployment alongside experienced physician readers to help inform multidisciplinary team decision-making.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Five-year US survival rates for lung cancer fall from 73% at stage I to 9–13% at stages IIIB and IV. [1] Hence, early diagnosis is critical to reducing mortality. Pulmonary nodules are often the first sign. [2] Around 5% of nodules 4–30 mm in size are malignant [3]. Nodules, benign and malignant, are detected in 1.6 million people in the US annually, [3] and the majority are detected in CT scans. [4]
These facts combine to establish two conclusions: (a) early detection and discrimination of pulmonary nodules are crucial to reducing lung cancer mortality, and (b) CT scans are an essential modality for this detection and discrimination.
However, discriminating between malignant and benign nodules is difficult. Assessing the risk of indeterminate nodules poses a particular challenge [5]. Clinical risk models, such as Herder, Mayo, and Brock [6,7,8], that use clinico-demographic or radiological inputs to aid physicians are commonly used.
Recently, image-based computer-aided diagnostic (CADx) models using deep learning (DL) have emerged to assess malignancy. These models are easy and fast to use versus clinical risk models. Therefore, image-based DL models have the potential to fulfil an unmet clinical-management need, [9] providing they produce comparable diagnostic performance.
The objective of this study was to assess the diagnostic performance of DL-based models for predicting the risk of malignancy in CT-detected pulmonary nodules. This is the first systematic review to provide a pooled analysis of studies that externally validate DL-based models and directly compare them with methods routinely used in clinical practice.
Methods
Search Strategy and Screening
An electronic search was performed in MEDLINE (PubMed), EMBASE, Science Citation Index, and Cochrane Library databases (from inception to 10 August 2023). Studies were deemed eligible if they were peer-reviewed experimental or observational articles that assessed the diagnostic performance of externally validated DL-based CADx models to predict the risk of malignancy in solid or part-solid nodules. The full set of keyword search terms (eTable 1) and selection criteria (eTable 2) are found in the Supplementary Material. References of key studies and domain-related systematic reviews were also investigated. This study followed PRISMA and Meta-analysis of Observational Studies in Epidemiology (MOOSE) guidelines. [10, 11]
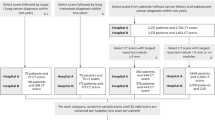
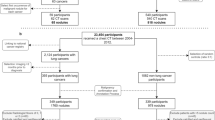
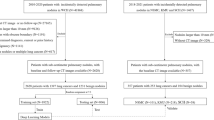
After removing duplicates, 7116 studies were found (Fig. 1). Screening out ineligible studies by title and abstract left 69 studies for final screening. Two investigators independently reviewed each text.
Data Extraction and Quality Assessment
Information from included studies were extracted independently by two investigators (eTable 3). The data were subsequently checked against selection criteria (eTable 2). Risk of bias and applicability were independently assessed using the Quality Assessment of Diagnostic Studies 2 (QUADAS-2) tool [12].
Statistical Analysis and Quantitative Synthesis
A meta-analysis of all included studies was conducted. For each index test type (DL-based models; physician judgement alone; clinical risk models alone; Lung-RADS-based models alone), pooled estimates of area under the curve (AUC), sensitivity, and specificity were calculated using a bivariate, random-effects approach. Deeks’ funnels were plotted to identify publication bias. To assess heterogeneity and inconsistency among the studies, τ2 statistic and I2 index values were calculated.
Potential sources of heterogeneity were investigated by conducting sub-group analyses, stratified by prevalence, route of detection, and geography.
Review Manager (RevMan) version 5.4, R statistical software version 4.3.1 (Beagle Scouts), and R packages ‘mada’ version 0.5.11 and ‘metafor’ version 4.2-0, were used to conduct the statistical analyses [13, 14].
Results
Study Characteristics
The literature search identified 17 studies for inclusion (Fig. 1), comprising 35 validation datasets, 8553 participants, and 9884 pulmonary nodules. Of these nodules, 1991 were confirmed to be malignant within the follow-up period (on average, 24 months) (Table 1).
All the studies’ datasets save one, were retrospective cohorts, with one study containing a prospective-cohort dataset [15]. Datasets included populations from North America (11 studies), Europe (six studies), and Asia (four studies) (Table 1).
Studies primarily assessed diagnostic performance. Some studies reported clinical utility outcomes, such as diagnostic re-classification [16, 17]. However, due to inconsistency, it was not possible to conduct a meta-analysis on clinical utility. The main outcomes sought were the confusion matrices, sensitivity and specificity, and AUCs (Table 1). Many studies did not report confusion matrix values directly. As such, these were calculated using reported sensitivity, specificity, and prevalence values.
Sixteen DL-based CADx models were identified from the included studies. The commonest learning algorithm used were convolutional neural networks (CNNs). Ten of the 16 models and 11 of the 17 included studies used a CNN algorithm as the basis for their malignancy prediction score.
For the external validation index tests, the commonest comparator was physician readers (13 of 35 datasets, from 11 studies). The majority were radiologists with ≥ 3 years’ experience.
The Brock model was the commonest clinical risk model (12 datasets from eight studies), followed by the Mayo model with eight datasets from three studies. The Mayo model is considered the most externally validated model [18], but the Brock model performs better in screening populations [19, 20].
Most studies considered participants in the 50–75 age bracket. All studies included both female and male participants. The studies spanned the range of nodule sizes [21].
The average prevalence of malignancy across studies was 23%. Most incidentally detected nodules had prevalence ≥ 20%, whereas most screening populations had prevalence < 20%.
Diagnostic Performance
DL-Based Models
For the DL-based models, meta-analysis of 34 datasets that reported AUC values or for which AUC values were able to be derived gave a pooled AUC of 0.86 (95% CI 0.83–0.90). (Fig. 2A). Sensitivity ranged from 0.37 (95% CI 0.25–0.50) for a 0.98 (95% CI 0.95–0.99) specificity [16], to 1.00 (95% CI 0.98–1.00) for a 0.28 (95% CI 0.26–0.31) specificity (Figs. 3A and 4A, respectively) [22]. Meta-analysis of 24 datasets gave a pooled sensitivity of 0.88 (95% CI 0.81–0.93) and specificity of 0.77 (95% CI 0.68–0.84) (Figs. 3A and 4A, respectively).
They had an I2 index of 90% (p < 0.01) for sensitivity and 99% (p = 0) for specificity, corresponding to very high statistical heterogeneity (an I2 value ≥ 75% was indicative of heterogeneity). The Deeks’ funnel plot showed no significant asymmetry, with p = 0.08 (a p < 0.05 result was assumed to be statistically significant), indicating no evidence of publication bias (eFigure 1A).
Physician Readers Alone
Separate pooled analysis for physician readers gave a pooled AUC slightly lower than DL-based models at 0.83 (95% CI 0.79–0.88) (Fig. 2B). They had sensitivity of 0.79 (95% CI 0.69–0.86) and specificity of 0.81 (95% CI 0.71–0.88) (Figs. 3B and 4B, respectively). Their I2 index was 89% (p < 0.01) for sensitivity and 95% (p < 0.01) for specificity, demonstrating high statistical heterogeneity. The Deeks’ funnel plot showed no significant asymmetry, with p = 0.31, indicating no evidence of publication bias (eFigure 1B).
Clinical Risk Models Alone
Pooled analysis for clinical risk models gave pooled AUC of 0.79 (95% CI 0.75–0.83) (Fig. 2C). They had sensitivity of 0.77 (95% CI 0.45–0.93) and specificity of 0.72 (95% CI 0.38–0.91) (Figs. 3C and 4C, respectively). Their I2 index was 94% (p < 0.01) for sensitivity and 99% (p < 0.01) for specificity, demonstrating very high statistical heterogeneity. The Deeks’ funnel plot showed no significant asymmetry, with p = 0.28, indicating no evidence of publication bias (eFigure 1C).
Lung-RADS-Based Models
Lastly, pooled analysis for Lung-RADS-based models gave a pooled sensitivity of 0.52 (95% CI 0.31–0.72) (Fig. 3D), and specificity of 0.61 (95% CI 0.49–0.71) (Fig. 4D). They had an I2 index of 94% (p < 0.01) for sensitivity and 97% (p < 0.01), demonstrating very high statistical heterogeneity. There were insufficient studies for a Deeks’ test.
Sub-group Analyses
Sub-group analyses revealed that DL-based CADx models displayed higher sensitivity on incidentally detected nodules than screening-detected nodules, 0.90 (95% CI 0.77–0.96) versus 0.84 (95% CI 0.76–0.90), respectively. This increased reliability in detecting lung cancer came at the cost of specificity with screening-detected nodules having 0.84 (95% CI 0.78–0.89) compared to incidentally detected nodules at 0.70 (95% CI 0.55–0.81). Accounting for threshold effects, screening populations performed better than incidental populations for all risk prediction methods (eFigure 2A–C), particularly clinical risk models: pooled AUC of 0.75 (95% CI 0.69–0.80) in screening-detected nodules versus 0.60 (95% CI 0.56–0.64) in incidentally detected nodules (eFigure 2C). The difference between ROC curves for DL-based and physician reader methods versus clinical risk models for incidentally detected nodules was particularly pronounced, translating into pooled AUCs of 0.74 (95% CI 0.71–0.77) and 0.77 (95% CI 0.71–0.82) for DL-based models and physician readers, respectively, versus 0.60 (95% CI 0.56–0.64) for clinical risk models (eFigure 2).
Further sub-group analyses were carried out on prevalence and geography. For prevalence, the baseline malignancy in CT-detected nodules (4–30 mm) in the US, ~ 5%, [23] was multiplied by a factor of 4, and used as the threshold for classifying a study’s prevalence as high or normal. Thus, datasets with > 20% prevalence were considered high, and < 20%, normal. Further thresholds at 10% and 30% were explored with similar results. For geography, datasets were classified according to continent: Europe, Asia, and North America. Neither prevalence nor geography were found to be a source of heterogeneity.
The analysis was also re-run excluding nodules assessed in follow-up CT scans [24]. The majority of studies assessed nodules from initial CT scans (Table 1). This reduced the pooled sensitivity and specificity of the Lung-RADS-based models, but did not significantly affect any other results.
Quality Assessment
Overall, a low risk of bias was found in most studies using QUADAS-2 (eTable 4). Selection of participants varied between studies. This may have contributed to biased estimates of sensitivity and specificity as well as inter-study heterogeneity. Therefore, most studies (nine of 17) scored an unclear risk of bias owing to patient selection, but low risk in other categories.
Discussion
Seventeen studies with external validation data were identified, from which pooled analyses found DL-based models had superior AUC of 0.86 (95% CI 0.83–0.90) as compared to other methods of predicting malignancy in pulmonary nodules (0.83 [95% CI 0.79–0.88] for experienced physician readers and 0.79 [95% CI 0.75–0.83] for clinical risk models). This review attempted to exhaustively search the literature for all studies and models relevant to the research question. There were two common reasons for ineligibility. First, studies did not conduct external validation of the DL-based model being analysed (at final screening, 30 studies had no direct validation against non-DL methods or in an external dataset) (Fig. 1). Second, studies were excluded because they concerned detection of pulmonary nodules, not risk assessment (at final screening, 11 studies with ineligible index tests) (Fig. 1).
In order to evaluate performance across different populations, external validation is crucial [37]. The majority of studies conducted validation on datasets that were used for training or testing (internal validation). In terms of validation against other methods, many studies were validated against other DL-based models, and not models currently used in clinical practice.
For the second commonest exclusion, computer-aided solutions for pulmonary nodule management can be broadly categorised into two types: computer-aided detection (CADe) and CADx (diagnosis) [38]. CADe detects suspicious nodules and segments them for further analysis. CADx provides a nodule- or patient-level classification of the risk of malignancy.
Two previous systematic reviews have studied this issue [39, 40]. Neither, however, directly compared DL-based models with methods used in clinical practice. Nor did they restrict the search to studies that externally validated models in populations other than the population on which they were trained. Only Forte et al. [39] performed a meta-analysis. They considered six studies, five of which are included here [22, 27,28,29, 35], and one that was excluded due to no external validation. Pooled sensitivity and specificity in Forte et al. [39] were 0.94 (95% CI 0.86–0.98) and 0.69 (95% CI 0.51–0.83), respectively, both with significant heterogeneity. Pooled AUC was 0.90 (95% CI 0.86–0.92) [39]. No quantitative comparisons against physician reader or clinical risk models were performed. The authors noted DL-based models performed well, and that, as non-invasive methods, they could provide support to clinics in diagnosing lung cancer early.
Limitations
Although these results strongly support the use of externally validated DL-based models, two important limitations were noted. First, only observational studies were found. This was expected given evidentiary requirements for diagnostic tools are not set as high as therapeutic interventions [41].
The second was the high heterogeneity between studies. High heterogeneity is likely given the very different DL-based models under consideration, and the further work required to calibrate some models. Sources of heterogeneity were investigated with sub-group analyses. Neither prevalence nor geography were found to be sources of heterogeneity. Route of detection (screening versus incidental) was found to be a potential source for clinical risk models. However, the strongest source of heterogeneity is likely the threshold or operating cut-off point used by researchers in testing the models [39]. The types of thresholds used varied considerably. They included fixing the specificity of models to 0.90 [36], to setting rule-out (definite benignity) malignancy scores (out of 1.0) at 0.05 or rule-in (definite malignancy) malignancy scores to 0.65 [16, 17].
Sensitivity to threshold effects was not investigated due to this variability. Nevertheless, the inclusion of AUC, which captures performance across all possible threshold values, and its concordance with sensitivity and specificity results helped alleviate this concern.
The low risk of bias found in most studies, and no significant publication bias further demonstrate the robustness of the findings.
Clinical Practice Guidelines
Indeterminate nodules are nodules without obvious signs of benignity (such as calcification) or malignancy (such as spiculation). These nodules are particularly problematic [5, 42]. In order to diagnose cancer or refer high-risk cases for further invasive investigation, clinical risk prediction models are used to aid the physician. At least two pulmonary nodule management guidelines explicitly mention the use of specific clinical risk models. The American College of Chest Physicians (ACCP) recommend using a “validated model” for ≥ 8 mm solid nodules along with or instead of physician judgement [43]. The guideline further notes that the Mayo model is the most validated model for nodules that have been incidentally detected. The British Thoracic Society (BTS) goes further, recommending the Brock model for all nodules ≥ 8 mm in size [44].
Approximately five malignant nodule patients are incidentally detected for every one detected via screening [45]. The evidence for the effectiveness of programme-based management of incidentally detected pulmonary nodules has led to more centres across the US looking to implement them [45]. However, incidental programmes require investment in infrastructure and nodule experts [46]. There is also a concerted drive to increase the uptake of and expand access to low-dose CT screening of at-risk populations for lung cancer [47].
Together these trends raise important challenges. With both early detection programmes detecting more pulmonary nodules, the number of nodules requiring image-based discrimination will cause a surge in workload for healthcare facilities. Clinical risk models, which require manual entry of variables, along with a shortage of nodule experts, mean most health systems are ill-equipped to handle such a surge.[23, 46] DL-based models, however, have the potential to mitigate these challenges by increasing throughput and efficiency, non-invasively [48]. Moreover, their automation can guide and enhance the capabilities of non-experts such as radiographic technologists [49]. By providing reliable, automated analyses of nodules that integrate into radiology workflows, DL-based CADx can assist nodule experts in accurately making faster and more rule-in and rule-out diagnoses [9, 50].
Future Research
More research needs to be undertaken on how the diagnostic performance of DL-based models translates into improved clinical utility and patient outcomes. Such research should be prospective, and consider a range of settings. While several studies have demonstrated clinical utility [9, 50, 51], further work is needed.
Nodule type and size over time are also important areas for future research. Most studies assessed only the risk of malignancy in initial CT scans. Studies over time on follow-up low-dose CT scans are a future area of research. For nodule type, ground-glass nodules (GGNs) were not considered in any of the datasets analysed. Although GGNs are mostly transient and comprise ~ 2% of nodules, persistent cases tend to have higher malignancy rates (~ 34%) than solid nodules [52]. As such, assessing the malignancy risk of GGNs needs further research.
As DL-based models are calibrated further, and become more routinely used in clinical practice, heterogeneity may reduce, as observed with the Mayo and Brock models for clinical risk [6, 7]. With the potential high-throughput advantages conferred by DL-based models, and their superior or comparable diagnostic performance as compared to other methods, routine clinical use will be important.
References
Woodard GA, Jones KD, Jablons DM (2016) Lung cancer staging and prognosis. Cancer Treat Res 170:47–75. https://doi.org/10.1007/978-3-319-40389-2_3
Loverdos K, Fotiadis A, Kontogianni C et al (2019) Lung nodules: a comprehensive review on current approach and management. Ann Thorac Med 14:226–238. https://doi.org/10.4103/atm.ATM_110_19
Gould MK, Tang T, Liu I-LA et al (2015) Recent trends in the identification of incidental pulmonary nodules. Am J Respir Crit Care Med 192:1208–1214. https://doi.org/10.1164/rccm.201505-0990OC
Mahesh M, Ansari AJ, Mettler FA (2023) Patient exposure from radiologic and nuclear medicine procedures in the United States and worldwide: 2009–2018. Radiology. https://doi.org/10.1148/radiol.221263
Paez R, Kammer MN, Massion P (2021) Risk stratification of indeterminate pulmonary nodules. Curr Opin Pulm Med 27:240–248. https://doi.org/10.1097/MCP.0000000000000780
Swensen SJ, Silverstein MD, Ilstrup DM et al (1997) The probability of malignancy in solitary pulmonary nodules. Application to small radiologically indeterminate nodules. Arch Intern Med 157:849–855
McWilliams A, Tammemagi MC, Mayo JR et al (2013) Probability of cancer in pulmonary nodules detected on first screening CT. N Engl J Med 369:910–919. https://doi.org/10.1056/NEJMoa1214726
Herder GJ, van Tinteren H, Golding RP et al (2005) Clinical prediction model to characterize pulmonary nodules. Chest 128:2490–2496. https://doi.org/10.1378/chest.128.4.2490
Tsakok MT, Mashar M, Pickup L et al (2021) The utility of a convolutional neural network (CNN) model score for cancer risk in indeterminate small solid pulmonary nodules, compared to clinical practice according to British Thoracic Society guidelines. Eur J Radiol 137:109553. https://doi.org/10.1016/j.ejrad.2021.109553
Stroup DF, Berlin JA, Morton SC (2000) Meta-analysis of observational studies in epidemiology. JAMA 283:2008. https://doi.org/10.1001/jama.283.15.2008
Page MJ, McKenzie JE, Bossuyt PM et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. https://doi.org/10.1136/bmj.n71
Whiting PF, Rutjes AWS, Westwood ME et al (2011) QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med 155:529–536. https://doi.org/10.7326/0003-4819-155-8-201110180-00009
The Cochrane Collaboration (2020) Review manager (RevMan). https://revman.cochrane.org/
Doebler P, Sousa-Pinto B (2022) Meta-analysis of diagnostic accuracy with mada. R Packages. https://r-forge.r-project.org/projects/mada
Chen K, Nie Y, Park S et al (2021) Development and validation of machine learning-based model for the prediction of malignancy in multiple pulmonary nodules: analysis from multicentric cohorts. Clin Cancer Res 27:2255–2265. https://doi.org/10.1158/1078-0432.CCR-20-4007
Massion PP, Antic S, Ather S et al (2020) Assessing the accuracy of a deep learning method to risk stratify indeterminate pulmonary nodules. Am J Respir Crit Care Med 202:241–249. https://doi.org/10.1164/rccm.201903-0505OC
Kim RY, Oke JL, Pickup LC et al (2022) Artificial intelligence tool for assessment of indeterminate pulmonary nodules detected with CT. Radiology 304:683–691. https://doi.org/10.1148/radiol.212182
Choi HK, Ghobrial M, Mazzone PJ (2018) Models to estimate the probability of malignancy in patients with pulmonary nodules. Ann Am Thorac Soc 15:1117–1126. https://doi.org/10.1513/AnnalsATS.201803-173CME
González Maldonado S, Delorme S, Hüsing A et al (2020) Evaluation of prediction models for identifying malignancy in pulmonary nodules detected via low-dose computed tomography. JAMA Netw Open 3:e1921221. https://doi.org/10.1001/jamanetworkopen.2019.21221
White CS, Dharaiya E, Campbell E, Boroczky L (2017) The vancouver lung cancer risk prediction model: assessment by using a subset of the national lung screening trial cohort. Radiology 283:264–272. https://doi.org/10.1148/radiol.2016152627
Hunter B, Chen M, Ratnakumar P et al (2022) A radiomics-based decision support tool improves lung cancer diagnosis in combination with the Herder score in large lung nodules. EBioMedicine 86:104344. https://doi.org/10.1016/j.ebiom.2022.104344
Baldwin DR, Gustafson J, Pickup L et al (2020) External validation of a convolutional neural network artificial intelligence tool to predict malignancy in pulmonary nodules. Thorax 75:306–312. https://doi.org/10.1136/thoraxjnl-2019-214104
Mazzone PJ, Lam L (2022) Evaluating the patient with a pulmonary nodule. JAMA 327:264. https://doi.org/10.1001/jama.2021.24287
Huang P, Lin CT, Li Y et al (2019) Prediction of lung cancer risk at follow-up screening with low-dose CT: a training and validation study of a deep learning method. Lancet Digit Health 1:e353–e362. https://doi.org/10.1016/S2589-7500(19)30159-1
Adams SJ, Mondal P, Penz E et al (2021) Development and cost analysis of a lung nodule management strategy combining artificial intelligence and Lung-RADS for baseline lung cancer screening. J Am Coll Radiol 18:741–751. https://doi.org/10.1016/j.jacr.2020.11.014
Adams SJ, Madtes DK, Burbridge B et al (2023) Clinical impact and generalizability of a computer-assisted diagnostic tool to risk-stratify lung nodules with CT. J Am Coll Radiol 20:232–242. https://doi.org/10.1016/j.jacr.2022.08.006
Ardila D, Kiraly AP, Bharadwaj S et al (2019) End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat Med 25:954–961. https://doi.org/10.1038/s41591-019-0447-x
Chen Y, Tian X, Fan K et al (2022) The value of artificial intelligence film reading system based on deep learning in the diagnosis of non-small-cell lung cancer and the significance of efficacy monitoring: a retrospective, clinical, nonrandomized Controlled Study. Comput Math Methods Med 2022:2864170. https://doi.org/10.1155/2022/2864170
Gürsoy Çoruh A, Yenigün B, Uzun Ç et al (2021) A comparison of the fusion model of deep learning neural networks with human observation for lung nodule detection and classification. Br J Radiol 94:20210222. https://doi.org/10.1259/bjr.20210222
Gao R, Tang Y, Khan MS et al (2021) Cancer risk estimation combining lung screening CT with clinical data elements. Radiol Artif Intell 3:e210032. https://doi.org/10.1148/ryai.2021210032
Gao R, Li T, Tang Y et al (2022) Reducing uncertainty in cancer risk estimation for patients with indeterminate pulmonary nodules using an integrated deep learning model. Comput Biol Med 150:106113. https://doi.org/10.1016/j.compbiomed.2022.106113
Jacobs C, Setio AAA, Scholten ET et al (2021) Deep learning for lung cancer detection on screening CT scans: results of a large-scale public competition and an observer study with 11 radiologists. Radiol Artif Intell 3:e210027. https://doi.org/10.1148/ryai.2021210027
Liao F, Liang M, Li Z et al (2019) Evaluate the malignancy of pulmonary nodules using the 3-D deep leaky noisy-OR network. IEEE Trans Neural Netw Learn Syst 30:3484–3495. https://doi.org/10.1109/TNNLS.2019.2892409
Liu J, Zhao L, Han X et al (2021) Estimation of malignancy of pulmonary nodules at CT scans: effect of computer-aided diagnosis on diagnostic performance of radiologists. Asia Pac J Clin Oncol 17:216–221. https://doi.org/10.1111/ajco.13362
Trajanovski S, Mavroeidis D, Swisher CL et al (2021) Towards radiologist-level cancer risk assessment in CT lung screening using deep learning. Comput Med Imaging Graph 90:101883. https://doi.org/10.1016/j.compmedimag.2021.101883
Venkadesh KV, Setio AAA, Schreuder A et al (2021) Deep learning for malignancy risk estimation of pulmonary nodules detected at low-dose screening CT. Radiology 300:438–447. https://doi.org/10.1148/radiol.2021204433
Ramspek CL, Jager KJ, Dekker FW et al (2021) External validation of prognostic models: what, why, how, when and where? Clin Kidney J 14:49–58. https://doi.org/10.1093/ckj/sfaa188
Maldonado F, Lentz RJ (2020) Reducing uncertainty to a manageable level: the need for a nuanced and patient-centric approach to lung nodule management in the 21st century. J Thorac Dis 12(6):3242–3244. https://doi.org/10.21037/jtd.2020.03.65
Forte GC, Altmayer S, Silva RF et al (2022) Deep learning algorithms for diagnosis of lung cancer: a systematic review and meta-analysis. Cancers (Basel). https://doi.org/10.3390/cancers14163856
Wu Z, Wang F, Cao W et al (2022) Lung cancer risk prediction models based on pulmonary nodules: a systematic review. Thorac Cancer 13:664–677. https://doi.org/10.1111/1759-7714.14333
Mazumdar M, Zhong X, Ferket B (2021) Diagnostic trials. Principles and practice of clinical trials. Springer, Cham
Peikert T, Bartholmai BJ, Maldonado F (2020) Radiomics-based Management of indeterminate lung nodules? Are we there yet? Am J Respir Crit Care Med 202:165–167. https://doi.org/10.1164/rccm.202004-1279ED
Gould MK, Donington J, Lynch WR et al (2013) Evaluation of individuals with pulmonary nodules: when is it lung cancer? Diagnosis and management of lung cancer: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 143:e93S-e120S. https://doi.org/10.1378/chest.12-2351
Callister MEJ, Baldwin DR, Akram AR et al (2015) British thoracic society guidelines for the investigation and management of pulmonary nodules. Thorax. https://doi.org/10.1136/thoraxjnl-2015-207168
Osarogiagbon RU, Liao W, Faris NR et al (2022) Lung cancer diagnosed through screening, lung nodule, and neither program: a prospective observational study of the detecting early lung cancer (DELUGE) in the Mississippi Delta Cohort. J Clin Oncol 40:2094–2105. https://doi.org/10.1200/JCO.21.02496
Hricak H, Abdel-Wahab M, Atun R et al (2021) Medical imaging and nuclear medicine: a lancet oncology commission. Lancet Oncol 22:e136–e172. https://doi.org/10.1016/S1470-2045(20)30751-8
National Lung Screening Trial Research Team, Aberle DR, Adams AM et al (2011) Reduced lung-cancer mortality with low-dose computed tomographic screening. N Engl J Med 365:395–409. https://doi.org/10.1056/NEJMoa1102873
Paez R, Kammer MN, Tanner NT et al (2023) Update on biomarkers for the stratification of indeterminate pulmonary nodules. Chest. https://doi.org/10.1016/j.chest.2023.05.025
Holland P, Spence H, Clubley A et al (2020) Reporting radiographers and their role in thoracic CT service improvement: managing the pulmonary nodule. BJR|Open 2(1):20190018. https://doi.org/10.1259/bjro.20190018
Paez R, Kammer MN, Balar A et al (2023) Longitudinal lung cancer prediction convolutional neural network model improves the classification of indeterminate pulmonary nodules. Sci Rep 13:6157. https://doi.org/10.1038/s41598-023-33098-y
Landy R, Wang VL, Baldwin DR et al (2023) Recalibration of a deep learning model for low-dose computed tomographic images to inform lung cancer screening intervals. JAMA Netw Open 6:e233273. https://doi.org/10.1001/jamanetworkopen.2023.3273
Henschke CI, Yankelevitz DF, Mirtcheva R et al (2002) CT screening for lung cancer: frequency and significance of part-solid and nonsolid nodules. Am J Roentgenol 178:1053–1057. https://doi.org/10.2214/ajr.178.5.1781053
Funding
This work was funded by Optellum Ltd.
Author information
Authors and Affiliations
Contributions
JW and WW conceived and designed the study. JB and CV carried out the literature search and extracted the data. CV and AA carried out statistical analysis. All authors helped interpret the findings. WW and JB wrote the first draft of the manuscript with input from FC. All authors provided input to subsequent drafts. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
JW is an employee of Optellum Ltd. Optellum Ltd holds some patents in the area of research, and funded the study. No other competing interests were reported.
Ethical Approval
This is a systematic review and meta-analysis. No ethical approval is required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wulaningsih, W., Villamaria, C., Akram, A. et al. Deep Learning Models for Predicting Malignancy Risk in CT-Detected Pulmonary Nodules: A Systematic Review and Meta-analysis. Lung (2024). https://doi.org/10.1007/s00408-024-00706-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00408-024-00706-1