Abstract
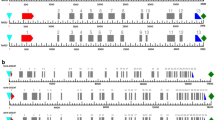
Nitrogen (N) is one of the major limiting elements affecting the growth and productivity of cereal crops including bread wheat. Identification and detailed characterization of the candidate genes involved in the assimilation of N from inorganic to organic form is very important for better N utilization. Glutamine synthetase 2 (GS2) and Ferredoxin-dependent-glutamate synthase (Fd-GOGAT) are the two critical enzymes involved in GS/GOGAT cycle, which are necessary for primary N-assimilation. In the present study, GS2 and Fd-GOGAT genes were cloned from a popular bread wheat variety HD-2967 and their individual homeologues (A, B and D) were characterized under different levels of N-stress. Homeologous cDNAs of GS2 and Fd-GOGAT showed > 93% and > 98% sequence similarity, respectively, in the coding region but were distinguishable at UTRs. The expression patterns and contribution of each homeologue of both the genes in HD-2967 under different N treatments showed downregulation in the shoot and upregulation in the root tissues. However, the contribution of the individual homeologue varied with the tissue, time point and level of N-stress. Further, when the expression pattern and contribution of each homeologue were studied in a diverse set of wheat genotypes, a generalised contribution pattern of each homeologue of wheat was observed. While the transcript ratio of A and D homeologues of both the genes demonstrated to be compensating each other, Fd-GOGAT-B showed the least contribution in all the conditions and at all stages of growth. Substantial variation in cis-regulatory elements was observed among all three homeologues of both the genes. Overall, characterization and spatio-temporal contribution of GS2 and Fd-GOGAT homeologues would provide applied value for further detailed studies as well as their exploration in primary N-assimilation process in wheat.










Similar content being viewed by others
Abbreviations
- GS2:
-
Glutamine synthetase
- Fd-GOGAT:
-
Ferredoxin-dependent-glutamate synthase
- N:
-
Nitrogen
- NO3 − :
-
Nitrate
- CDS:
-
Coding sequence
- mmol/L:
-
Millimolar/litre
- MS:
-
Murashige and Skoog
- H:
-
Hour
- N:P:K:
-
Nitrogen:phosphorus:potassium
- Kg/h:
-
Kilogram/hour
- RNA:
-
Ribonucleic acid
- cDNA:
-
Complementary deoxy- ribonucleic acid
- dNTPs:
-
Deoxynucleoside triphosphates
- PCR:
-
Polymerase chain reaction
- qPCR:
-
Quantitative-polymerase chain reaction
- Tm:
-
Melting temperature
- °C:
-
Degree celsius
- ORF:
-
Open reading frame
- MW:
-
Molecular weight
- pI:
-
Isoelectric point
- NCBI:
-
National Center for Biotechnology Information
- Μl:
-
Microlitre
- Ng:
-
Nanogram
- Bp:
-
Basepair
- UTR:
-
Untranslated region
References
Abrol YP (1990) Pattern of nitrate assimilation and grain nitrogen yield in field grown wheat (Triticum aestivum). In: Van Beusichem ML (ed) Plant nutrition—physiology and application. Kluwer Academic Publishers, Dordrecht, pp 773–778
Aoki N et al (2002) Three sucrose transporter genes are expressed in the developing grain of hexaploid wheat. Plant Mol Biol 50:453–462
Cai H, Lu Y, Xie W, Zhu T, Lian X (2012) Transcriptome response to nitrogen starvation in rice. J Biosci 37:731–747
Ciaffi M, Paolacci AR, Daloisio E, Tanzarella OA, Porceddu E (2006) Cloning and characterization of wheat PDI (protein disulfide isomerase) homeologous genes and promoter sequences. Gene 366:209–218
Crawford NM, Arst HNJ (1993) The molecular genetics of nitrate assimilation in fungi and plants. Annu Rev Genet 27:115–146
Garcia-Oliveira AL, Benito C, Prieto P, de AndradeMenezes R, Rodrigues-Pousada C, Guedes-Pinto H et al (2013) Molecular characterization of TaSTOP1 homeologues and their response to aluminium and proton (H+) toxicity in bread wheat (Triticum aestivum L.). BMC Plant Biol 13:134
Gayatri RM, Mahato AK, Sinha SK, Dalal M, Singh NK, Mandal PK (2018) Homeologue specific gene expression analysis of two vital carbon metabolizing enzymes-citrate synthase and NADP-isocitrate dehydrogenase-from wheat (Triticum aestivum L.) under nitrogen stress. Appl Biochem Biotechnol 188:569–584
Gayatri JK, Sinha SK, Mandal PK (2021) Comparative analysis of GS2 and Fd-GOGAT genes in cultivated wheat and their progenitors under N stress. Plant Mol Biol Rep. https://doi.org/10.1007/s11105-020-01267-2
Gupta A, Singh C, Kumar V, Tyagi B, Tiwari V, Chatrath R, Singh G (2018) Wheat varieties notified in India since 1965. ICAR- Indian Institute of Wheat & Barley Research, Karnal, p 101
He P, Friebe BR, Gill BS et al (2003) Allopolyploidy alters gene expression in the highly stable hexaploid wheat. Plant Mol Biol 52:401–414
Himi E, Noda K (2004) Isolation and location of three homeologous dihydroflavonol-4-reductase (DFR) genes of wheat and their tissue-dependent expression. J Exp Bot 55:365–375
Hirel B, Gadal P (1980) Glutamine synthetase in rice: a comparative study of the enzyme from roots and leaves. Plant Physiol 66:619–623
Hu ZR, Yu Y, Wang R, Yao YY, Peng HR, Ni ZF, Sun QX (2011) Expression divergence of TaMBD2 homeologous genes encoding methyl CpG-binding domain proteins in wheat (Triticum aestivum L.). Gene 471:13–18
Hu ZR, Han ZF, Song N, Chai LL, Yao YY, Peng HR, Ni ZF, Sun QX (2013a) Epigenetic modification contributes to the expression divergence of three TaEXPA1 homeologues in hexaploid wheat (Triticum aestivum). New Phytolog 197:1344–1352
Hu Z, Song N, Xing J, Chen Y, Han Z et al (2013) Overexpression of three TaEXPA1 homeologous genes with distinct expression divergence in hexaploid wheat exhibit functional retention in arabidopsis. PLoS ONE 8(5):e63667
Kraiser T, Gras DE, Gutierrez AG, Gonzalez B, Gutierrez RA (2011) A holistic view of nitrogen acquisition in plants. J Exp Bot 62:1455–1466
Kumagai, E., Araki , T. & Ueno , O. Ammonia emission from leaves of different rice (Oryza sativa L.) cultivars, https://academic.oup.com/aob/article/108/7/1381/135940#83951407 (2011).
Lawlor DW, Kontturi M, Young AT (1989) Photosynthesis by flag leaves of wheat in relation of protein, ribulose bisphosphate carboxylase activity and nitrogen supply. J Exp Bot 40:43–52
Lea PJ (1993) Nitrogen metabolism. In: Lea PJ, Leegood RC (eds) Plant biochemistry and molecular biology. Wiley, New York, pp 155–180
Lescot M, Dehais P, Thijs G et al (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Mattsson M, Johansson E, Lundborg T, Larsson M, Larsson CM (1991) Nitrogen utilization in N-limited barley during vegetative and generative growth. I. Growth and nitrate uptake kinetics in vegetative cultures grown at different relative addition rates of nitrate. J Exp Bot 42:197–205
Mochida K, Yamazaki Y, Ogihara Y (2004) Discrimination of homeologous gene expression in hexaploid wheat by SNP analysis of contigs grouped from a large number of expressed sequence tags. Mol Genet Genom 270:371–377
Nair TVR, Abrol YP (1978) Nitrate assimilation and mobilization of reduced nitrogen from upper three leaf blades to the developing wheat (Triticum aestivum L.) grains. Indian J Plant Physiol 21:41–47
Nomura T, Ishihara A, Yanagita RC, Endo TR, Iwamura H (2005) Three genomes differentially contribute to the biosynthesis of benzoxazinones in hexaploid wheat. Proc Natl Acad Sci USA 102:16490–16495
Pauline CN, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucl Acid Res 31:3812–3814
Poorter H, Fiorani F, Stitt M, Schurr U, Finck A, Gibon Y, Usadel B, Munns R, Atkin OK, Tardieu F, Pons TL (2012) The art of growing plants for experimental purposes: a practical guide for the plant biologist. Funct Plant Biol 39:821–838
Shitsukawa N et al (2007) Genetic and epigenetic alteration among three homeologous genes of a class e MADS box gene in hexaploid wheat. Plant Cell 19:1723–1737
Sinha SK, Kumar A, Tyagi A, Venkatesh K, Paul D, Singh NK, Mandal PK (2020) Root architecture traits variation and nitrate-influx responses in diverse wheat genotypes under different external nitrogen concentrations. Plant Physiol Biochem 148:246–259
Suzuki A, Gadal P (1984) Glutamate synthase: physiochemicaland functional properties of different forms in higher plants and in other organisms. Physiol Veg 22:471–486
Tobin AK, Yamaya T (2001) Cellular compartmentation of ammonium assimilation in rice and barley. J Exp Bot 52:591–604
Tovkach A, Ryan PR, Richardson AE, Lewis DC, Rathjen TM, Ramesh S, Tyerman SD, Delhaize E (2013) Transposon-mediated alteration of TaMATE1B expression in wheat confers constitutive citrate efflux from root apices. Plant Physiol 161:880–892
Vanoni MA, Curti B (1999) Glutamate synthase: a complex iron–sulfur flavoprotein. Cell Mol Life Sci 55:617–638
Wendel JF (2000) Genome evolution in polyploids. Plant Mol Biol 42:225–249
Yadava P, Aggarwal C, Verma R, Kumar K, Singh I (2020) Effect of nitrogen-starvation on growth pattern and expression of nitrogen assimilation genes. Indian J Agric Sci 90:195–200
Acknowledgements
Authors would also like to acknowledge the support and guidance provided by the Project Director, ICAR-National Institute for Plant Biotechnology, New Delhi. The authors also want to thank Puja Mandal for helping in editing the manuscript.
Funding
This research is part of the Indo-UK Virtual Nitrogen Centre project, financially supported by the Department of Biotechnology, Govt. of India (BT/IN/UK-VNC/43/KV/2015-16) under the Indo-UK Centre for Improvement of Nitrogen Use Efficiency in Wheat (INEW) and NIPB institutional in-house project on Biochemical and Molecular Basis of Nitrogen Utilization in Wheat.
Author information
Authors and Affiliations
Contributions
Conceptualization, PKM and G; Data curation, PKM; Formal analysis, G; Funding acquisition, PKM; Investigation, PKM; Methodology, G; Project administration, PKM; Resources, PKM, KV (9 wheat genotypes); Software, G; Supervision, PKM; Validation, PKM; Visualization, PKM; Writing-original draft, G; Writing-review and editing PKM, KV, SKS, PR. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Ranjit Riar.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gayatri, Venkatesh, K., Sinha, S.K. et al. Molecular Characterization of GS2 and Fd-GOGAT Homeologues and Their Biased Response to Nitrogen Stress in Bread Wheat (Triticum aestivum L.). J Plant Growth Regul 41, 2555–2569 (2022). https://doi.org/10.1007/s00344-021-10433-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-021-10433-z